Hesham Mostafa
Robust Federated Learning Through Representation Matching and Adaptive Hyper-parameters
Dec 30, 2019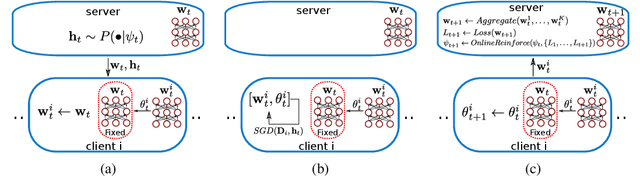

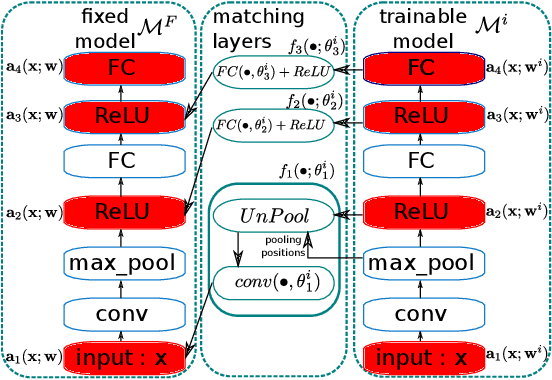
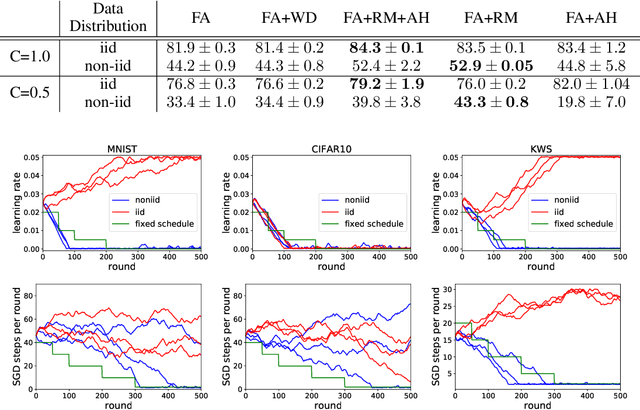
Abstract:Federated learning is a distributed, privacy-aware learning scenario which trains a single model on data belonging to several clients. Each client trains a local model on its data and the local models are then aggregated by a central party. Current federated learning methods struggle in cases with heterogeneous client-side data distributions which can quickly lead to divergent local models and a collapse in performance. Careful hyper-parameter tuning is particularly important in these cases but traditional automated hyper-parameter tuning methods would require several training trials which is often impractical in a federated learning setting. We describe a two-pronged solution to the issues of robustness and hyper-parameter tuning in federated learning settings. We propose a novel representation matching scheme that reduces the divergence of local models by ensuring the feature representations in the global (aggregate) model can be derived from the locally learned representations. We also propose an online hyper-parameter tuning scheme which uses an online version of the REINFORCE algorithm to find a hyper-parameter distribution that maximizes the expected improvements in training loss. We show on several benchmarks that our two-part scheme of local representation matching and global adaptive hyper-parameters significantly improves performance and training robustness.
Single-bit-per-weight deep convolutional neural networks without batch-normalization layers for embedded systems
Jul 22, 2019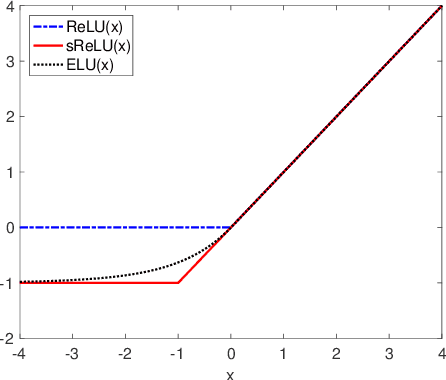
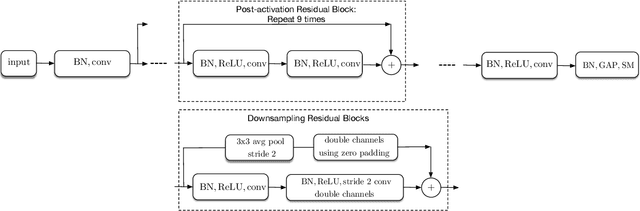
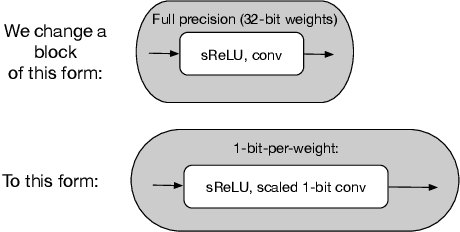
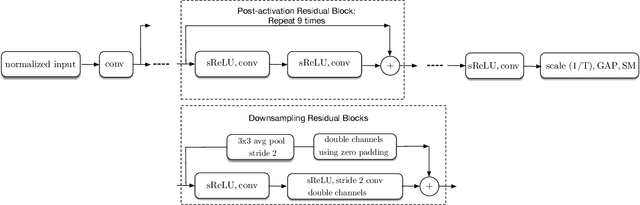
Abstract:Batch-normalization (BN) layers are thought to be an integrally important layer type in today's state-of-the-art deep convolutional neural networks for computer vision tasks such as classification and detection. However, BN layers introduce complexity and computational overheads that are highly undesirable for training and/or inference on low-power custom hardware implementations of real-time embedded vision systems such as UAVs, robots and Internet of Things (IoT) devices. They are also problematic when batch sizes need to be very small during training, and innovations such as residual connections introduced more recently than BN layers could potentially have lessened their impact. In this paper we aim to quantify the benefits BN layers offer in image classification networks, in comparison with alternative choices. In particular, we study networks that use shifted-ReLU layers instead of BN layers. We found, following experiments with wide residual networks applied to the ImageNet, CIFAR 10 and CIFAR 100 image classification datasets, that BN layers do not consistently offer a significant advantage. We found that the accuracy margin offered by BN layers depends on the data set, the network size, and the bit-depth of weights. We conclude that in situations where BN layers are undesirable due to speed, memory or complexity costs, that using shifted-ReLU layers instead should be considered; we found they can offer advantages in all these areas, and often do not impose a significant accuracy cost.
Parameter Efficient Training of Deep Convolutional Neural Networks by Dynamic Sparse Reparameterization
Mar 19, 2019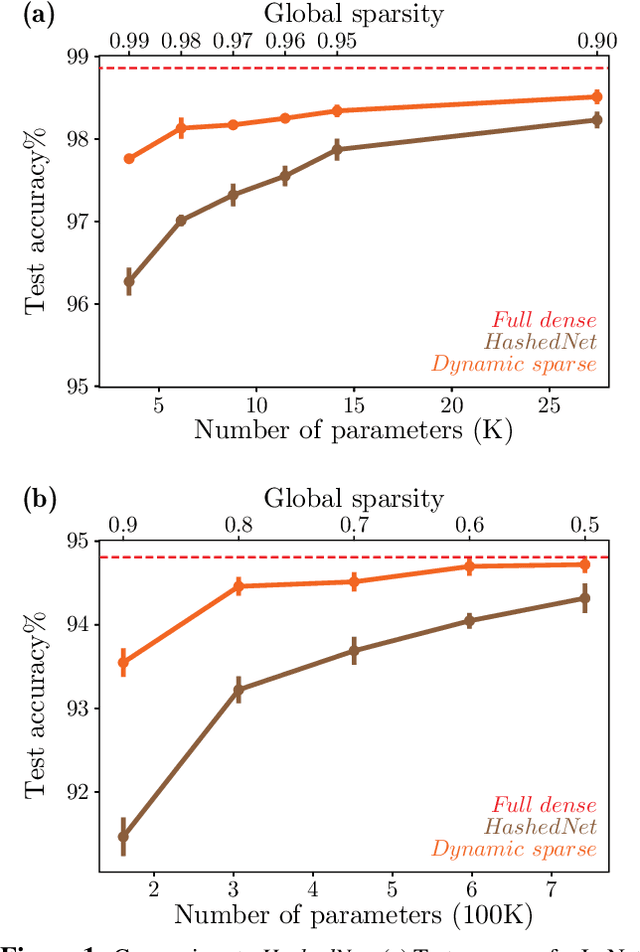
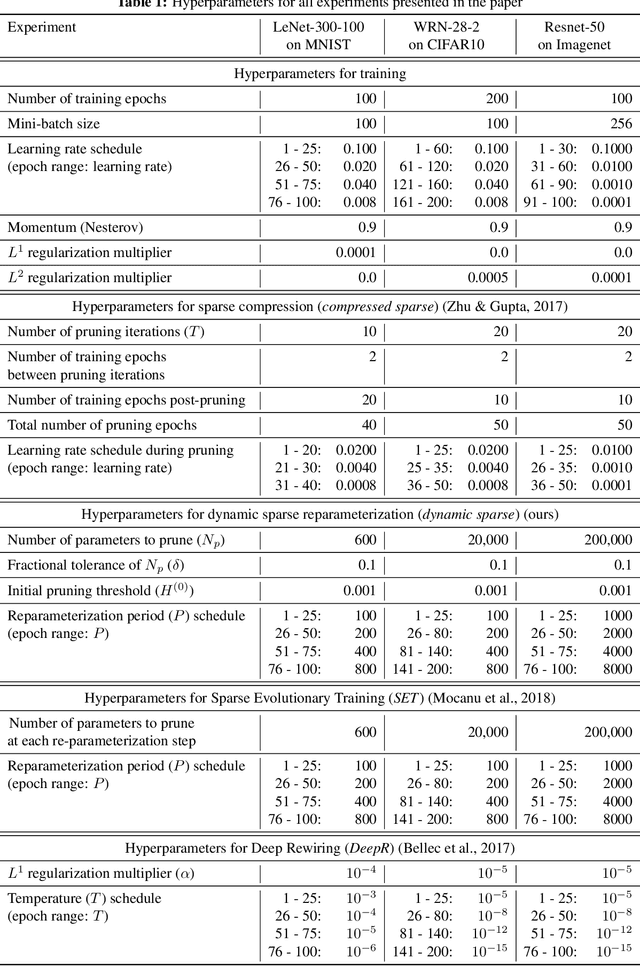
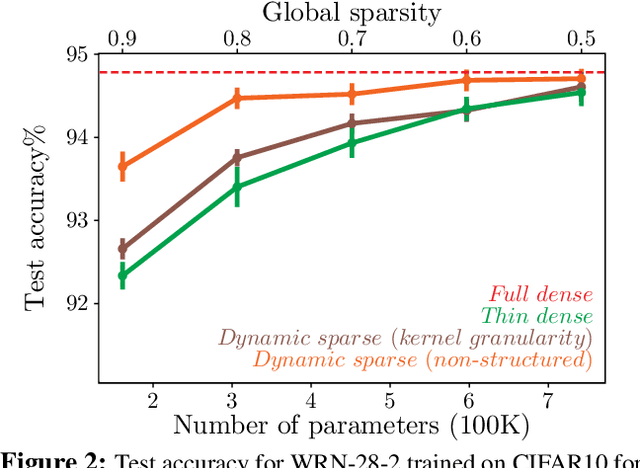

Abstract:Deep neural networks are typically highly over-parameterized with pruning techniques able to remove a significant fraction of network parameters with little loss in accuracy. Recently, techniques based on dynamic re-allocation of non-zero parameters have emerged for training sparse networks directly without having to train a large dense model beforehand. We present a parameter re-allocation scheme that addresses the limitations of previous methods such as their high computational cost and the fixed number of parameters they allocate to each layer. We investigate the performance of these dynamic re-allocation methods in deep convolutional networks and show that our method outperforms previous static and dynamic parameterization methods, yielding the best accuracy for a given number of training parameters, and performing on par with networks obtained by iteratively pruning a trained dense model. We further investigated the mechanisms underlying the superior performance of the resulting sparse networks. We found that neither the structure, nor the initialization of the sparse networks discovered by our parameter reallocation scheme are sufficient to explain their superior generalization performance. Rather, it is the continuous exploration of different sparse network structures during training that is critical to effective learning. We show that it is more fruitful to explore these structural degrees of freedom than to add extra parameters to the network.
Surrogate Gradient Learning in Spiking Neural Networks
Jan 28, 2019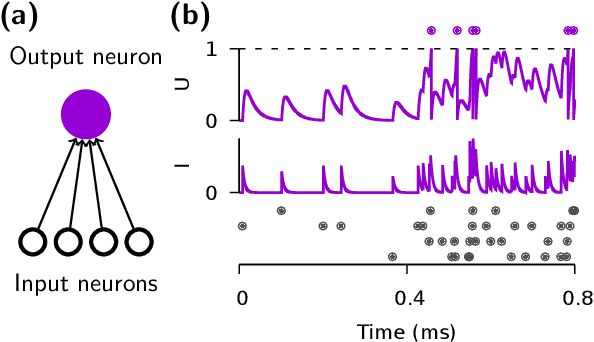
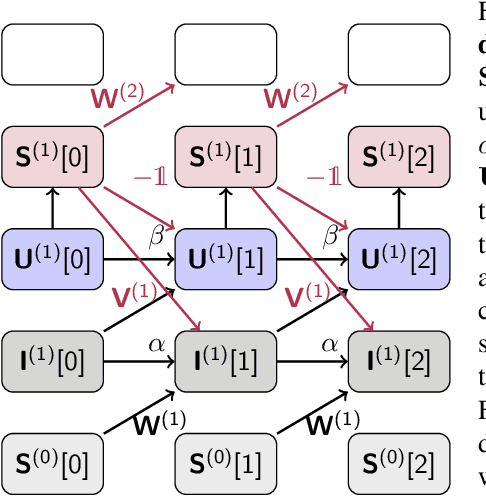
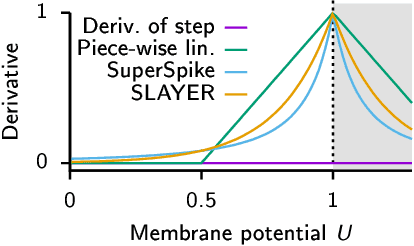
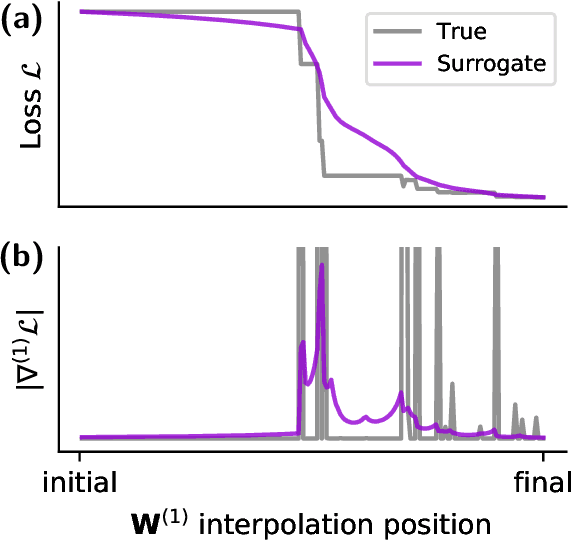
Abstract:A growing number of neuromorphic spiking neural network processors that emulate biological neural networks create an imminent need for methods and tools to enable them to solve real-world signal processing problems. Like conventional neural networks, spiking neural networks are particularly efficient when trained on real, domain specific data. However, their training requires overcoming a number of challenges linked to their binary and dynamical nature. This tutorial elucidates step-by-step the problems typically encountered when training spiking neural networks, and guides the reader through the key concepts of synaptic plasticity and data-driven learning in the spiking setting. To that end, it gives an overview of existing approaches and provides an introduction to surrogate gradient methods, specifically, as a particularly flexible and efficient method to overcome the aforementioned challenges.
Synaptic Plasticity Dynamics for Deep Continuous Local Learning
Nov 27, 2018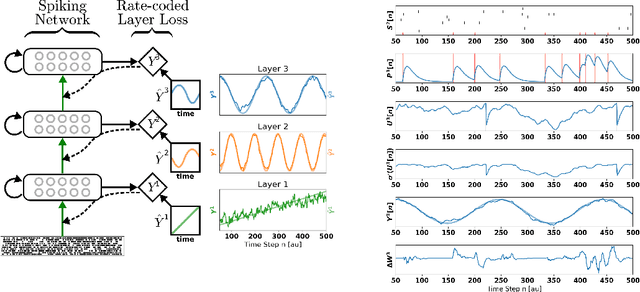
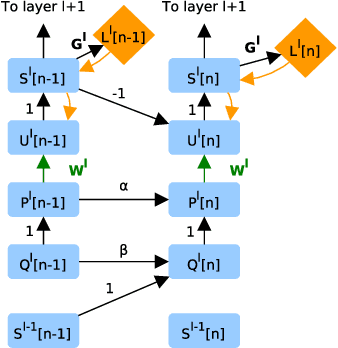
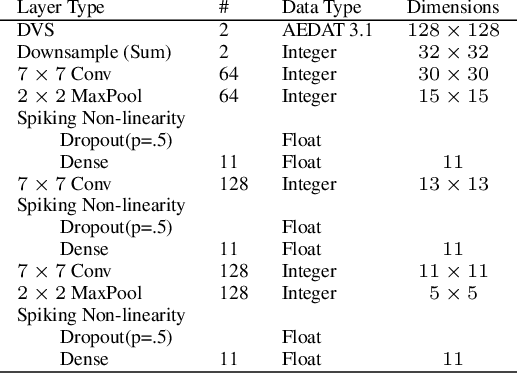
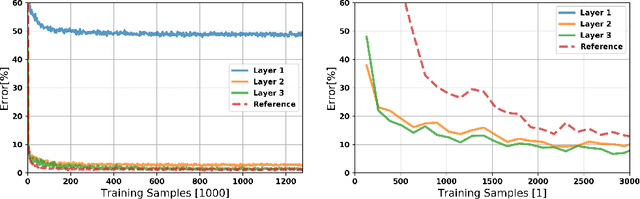
Abstract:A growing body of work underlines striking similarities between spiking neural networks modeling biological networks and recurrent, binary neural networks. A relatively smaller body of work, however, discuss similarities between learning dynamics employed in deep artificial neural networks and synaptic plasticity in spiking neural networks. The challenge preventing this is largely due to the discrepancy between dynamical properties of synaptic plasticity and the requirements for gradient backpropagation. Here, we demonstrate that deep learning algorithms that locally approximate the gradient backpropagation updates using locally synthesized gradients overcome this challenge. Locally synthesized gradients were initially proposed to decouple one or more layers from the rest of the network so as to improve parallelism. Here, we exploit these properties to derive gradient-based learning rules in spiking neural networks. Our approach results in highly efficient spiking neural networks and synaptic plasticity capable of training deep neural networks. Furthermore, our method utilizes existing autodifferentation methods in machine learning frameworks to systematically derive synaptic plasticity rules from task-relevant cost functions and neural dynamics. We benchmark our approach on the MNIST and DVS Gestures dataset, and report state-of-the-art results on the latter. Our results provide continuously learning machines that are not only relevant to biology, but suggestive of a brain-inspired computer architecture that matches the performances of GPUs on target tasks.
NullHop: A Flexible Convolutional Neural Network Accelerator Based on Sparse Representations of Feature Maps
Mar 06, 2018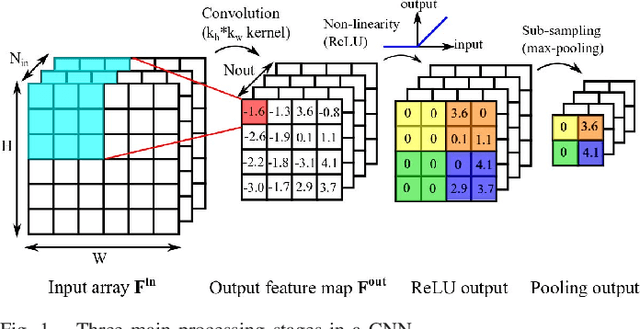
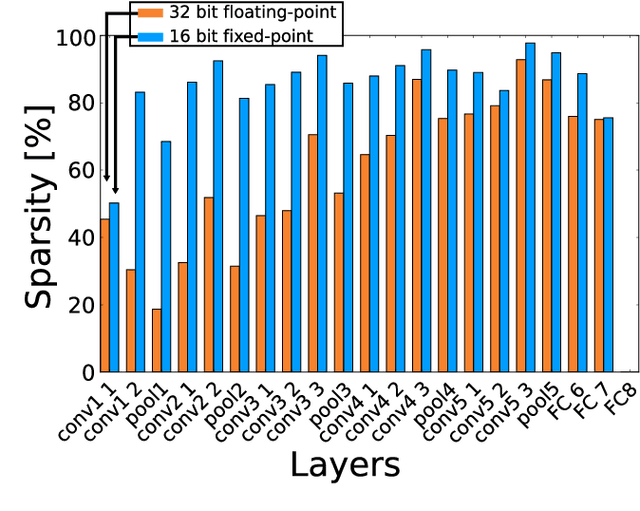
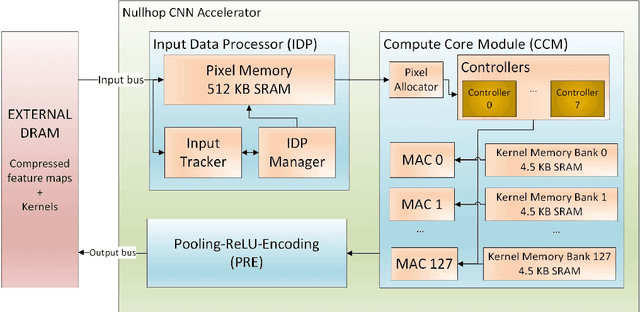
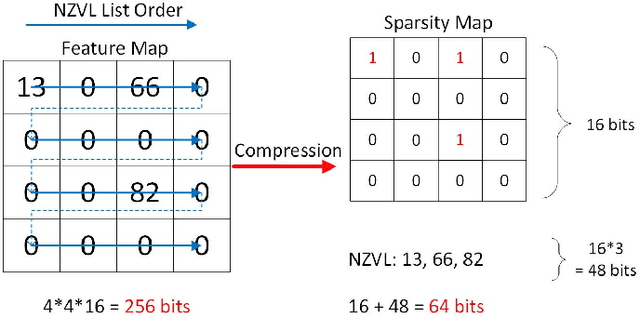
Abstract:Convolutional neural networks (CNNs) have become the dominant neural network architecture for solving many state-of-the-art (SOA) visual processing tasks. Even though Graphical Processing Units (GPUs) are most often used in training and deploying CNNs, their power efficiency is less than 10 GOp/s/W for single-frame runtime inference. We propose a flexible and efficient CNN accelerator architecture called NullHop that implements SOA CNNs useful for low-power and low-latency application scenarios. NullHop exploits the sparsity of neuron activations in CNNs to accelerate the computation and reduce memory requirements. The flexible architecture allows high utilization of available computing resources across kernel sizes ranging from 1x1 to 7x7. NullHop can process up to 128 input and 128 output feature maps per layer in a single pass. We implemented the proposed architecture on a Xilinx Zynq FPGA platform and present results showing how our implementation reduces external memory transfers and compute time in five different CNNs ranging from small ones up to the widely known large VGG16 and VGG19 CNNs. Post-synthesis simulations using Mentor Modelsim in a 28nm process with a clock frequency of 500 MHz show that the VGG19 network achieves over 450 GOp/s. By exploiting sparsity, NullHop achieves an efficiency of 368%, maintains over 98% utilization of the MAC units, and achieves a power efficiency of over 3TOp/s/W in a core area of 6.3mm$^2$. As further proof of NullHop's usability, we interfaced its FPGA implementation with a neuromorphic event camera for real time interactive demonstrations.
A learning framework for winner-take-all networks with stochastic synapses
Feb 05, 2018Abstract:Many recent generative models make use of neural networks to transform the probability distribution of a simple low-dimensional noise process into the complex distribution of the data. This raises the question of whether biological networks operate along similar principles to implement a probabilistic model of the environment through transformations of intrinsic noise processes. The intrinsic neural and synaptic noise processes in biological networks, however, are quite different from the noise processes used in current abstract generative networks. This, together with the discrete nature of spikes and local circuit interactions among the neurons, raises several difficulties when using recent generative modeling frameworks to train biologically motivated models. In this paper, we show that a biologically motivated model based on multi-layer winner-take-all (WTA) circuits and stochastic synapses admits an approximate analytical description. This allows us to use the proposed networks in a variational learning setting where stochastic backpropagation is used to optimize a lower bound on the data log likelihood, thereby learning a generative model of the data. We illustrate the generality of the proposed networks and learning technique by using them in a structured output prediction task, and in a semi-supervised learning task. Our results extend the domain of application of modern stochastic network architectures to networks where synaptic transmission failure is the principal noise mechanism.
Deep supervised learning using local errors
Nov 17, 2017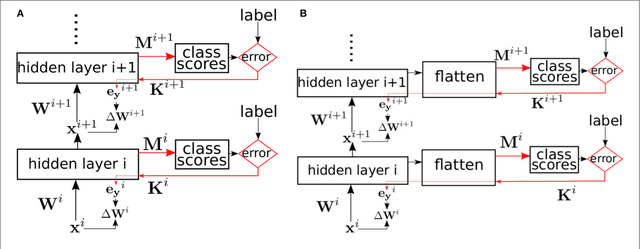

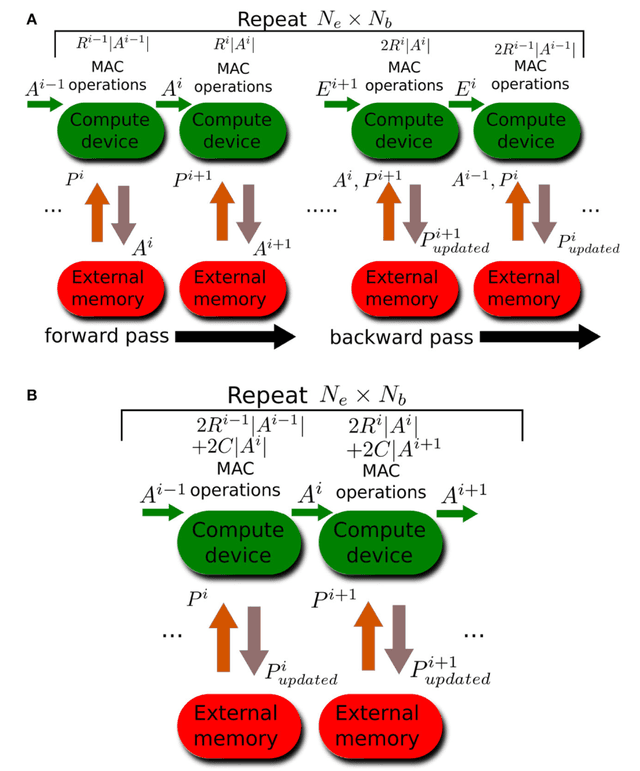
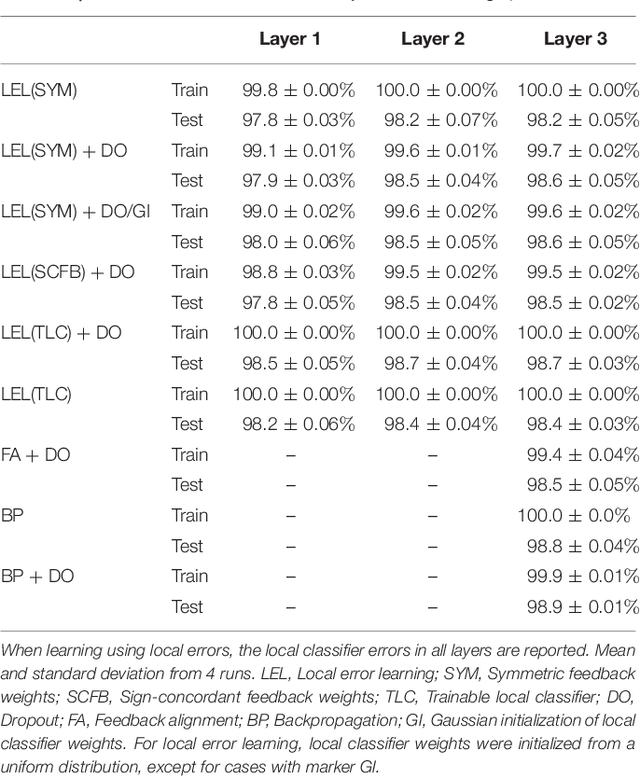
Abstract:Error backpropagation is a highly effective mechanism for learning high-quality hierarchical features in deep networks. Updating the features or weights in one layer, however, requires waiting for the propagation of error signals from higher layers. Learning using delayed and non-local errors makes it hard to reconcile backpropagation with the learning mechanisms observed in biological neural networks as it requires the neurons to maintain a memory of the input long enough until the higher-layer errors arrive. In this paper, we propose an alternative learning mechanism where errors are generated locally in each layer using fixed, random auxiliary classifiers. Lower layers could thus be trained independently of higher layers and training could either proceed layer by layer, or simultaneously in all layers using local error information. We address biological plausibility concerns such as weight symmetry requirements and show that the proposed learning mechanism based on fixed, broad, and random tuning of each neuron to the classification categories outperforms the biologically-motivated feedback alignment learning technique on the MNIST, CIFAR10, and SVHN datasets, approaching the performance of standard backpropagation. Our approach highlights a potential biological mechanism for the supervised, or task-dependent, learning of feature hierarchies. In addition, we show that it is well suited for learning deep networks in custom hardware where it can drastically reduce memory traffic and data communication overheads.
Hardware-efficient on-line learning through pipelined truncated-error backpropagation in binary-state networks
Aug 16, 2017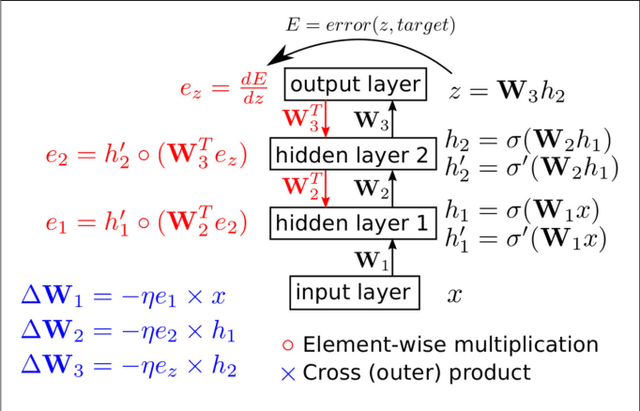
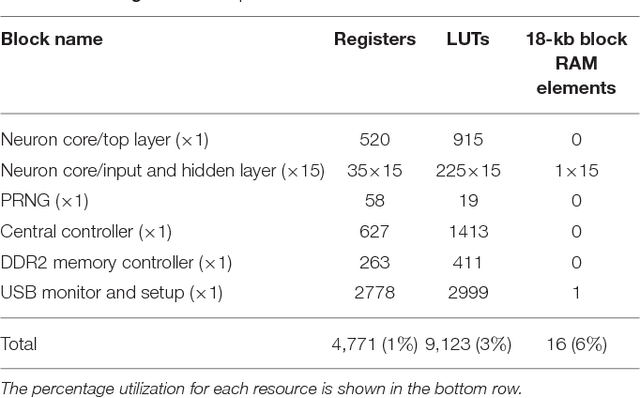
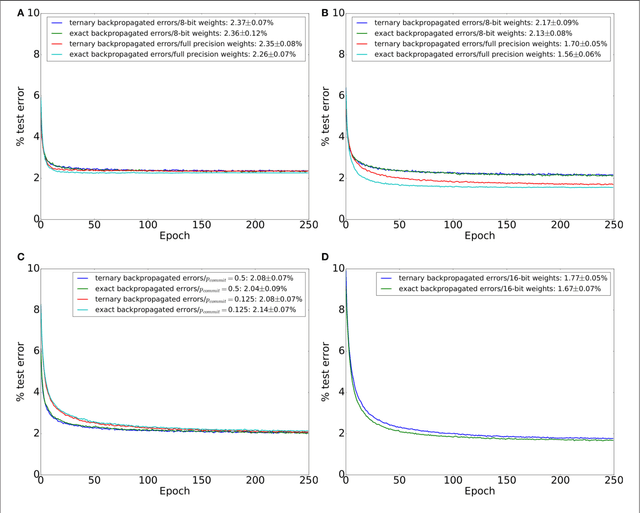
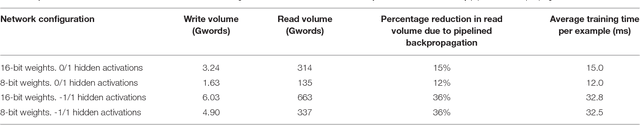
Abstract:Artificial neural networks (ANNs) trained using backpropagation are powerful learning architectures that have achieved state-of-the-art performance in various benchmarks. Significant effort has been devoted to developing custom silicon devices to accelerate inference in ANNs. Accelerating the training phase, however, has attracted relatively little attention. In this paper, we describe a hardware-efficient on-line learning technique for feedforward multi-layer ANNs that is based on pipelined backpropagation. Learning is performed in parallel with inference in the forward pass, removing the need for an explicit backward pass and requiring no extra weight lookup. By using binary state variables in the feedforward network and ternary errors in truncated-error backpropagation, the need for any multiplications in the forward and backward passes is removed, and memory requirements for the pipelining are drastically reduced. Further reduction in addition operations owing to the sparsity in the forward neural and backpropagating error signal paths contributes to highly efficient hardware implementation. For proof-of-concept validation, we demonstrate on-line learning of MNIST handwritten digit classification on a Spartan 6 FPGA interfacing with an external 1Gb DDR2 DRAM, that shows small degradation in test error performance compared to an equivalently sized binary ANN trained off-line using standard back-propagation and exact errors. Our results highlight an attractive synergy between pipelined backpropagation and binary-state networks in substantially reducing computation and memory requirements, making pipelined on-line learning practical in deep networks.
Supervised learning based on temporal coding in spiking neural networks
Aug 16, 2017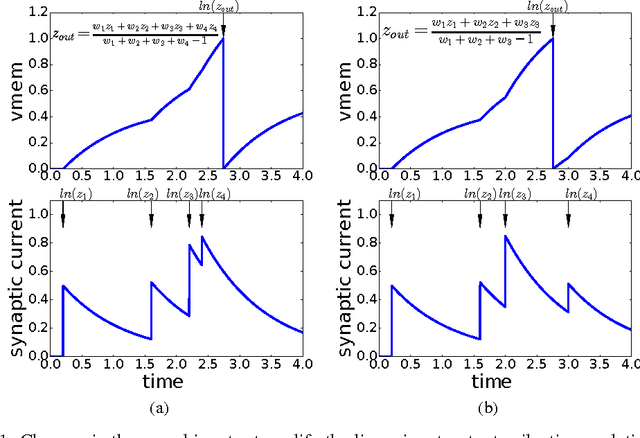
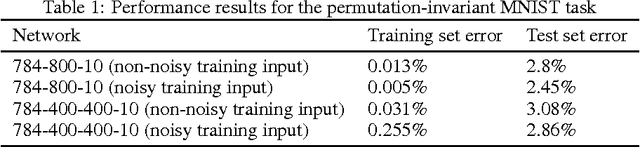
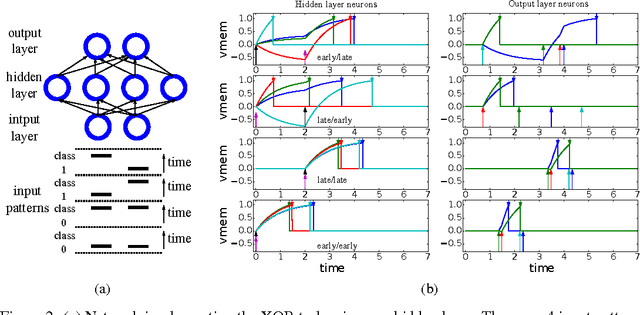
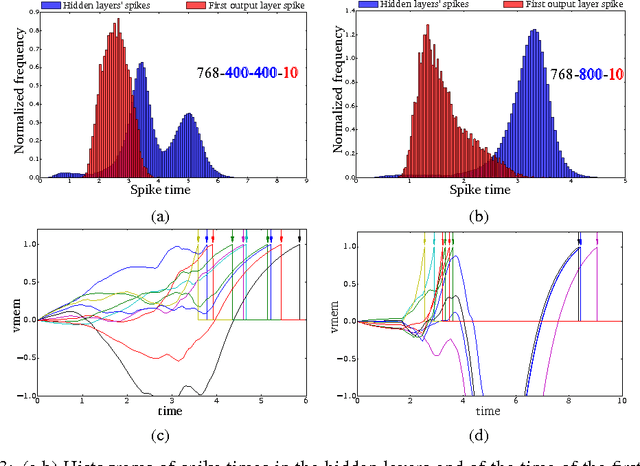
Abstract:Gradient descent training techniques are remarkably successful in training analog-valued artificial neural networks (ANNs). Such training techniques, however, do not transfer easily to spiking networks due to the spike generation hard non-linearity and the discrete nature of spike communication. We show that in a feedforward spiking network that uses a temporal coding scheme where information is encoded in spike times instead of spike rates, the network input-output relation is differentiable almost everywhere. Moreover, this relation is piece-wise linear after a transformation of variables. Methods for training ANNs thus carry directly to the training of such spiking networks as we show when training on the permutation invariant MNIST task. In contrast to rate-based spiking networks that are often used to approximate the behavior of ANNs, the networks we present spike much more sparsely and their behavior can not be directly approximated by conventional ANNs. Our results highlight a new approach for controlling the behavior of spiking networks with realistic temporal dynamics, opening up the potential for using these networks to process spike patterns with complex temporal information.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge