Harini Suresh
A Framework for Understanding Unintended Consequences of Machine Learning
Jan 28, 2019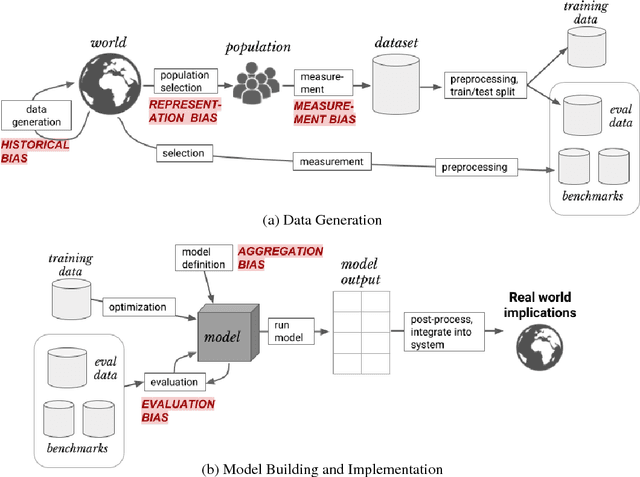
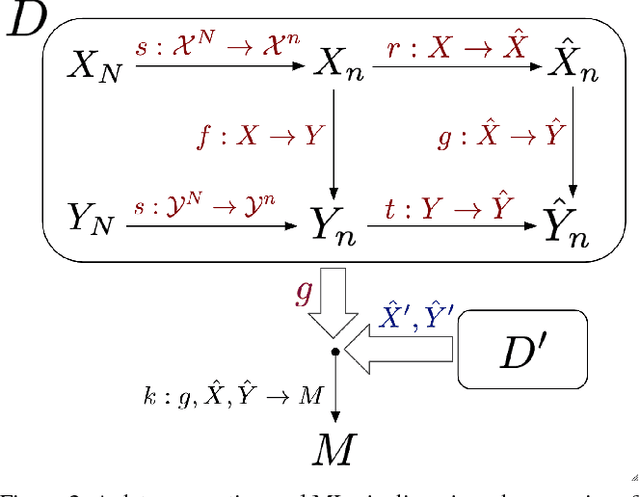
Abstract:As machine learning increasingly affects people and society, it is important that we strive for a comprehensive and unified understanding of how and why unwanted consequences arise. For instance, downstream harms to particular groups are often blamed on "biased data," but this concept encompass too many issues to be useful in developing solutions. In this paper, we provide a framework that partitions sources of downstream harm in machine learning into five distinct categories spanning the data generation and machine learning pipeline. We describe how these issues arise, how they are relevant to particular applications, and how they motivate different solutions. In doing so, we aim to facilitate the development of solutions that stem from an understanding of application-specific populations and data generation processes, rather than relying on general claims about what may or may not be "fair."
Modeling Mistrust in End-of-Life Care
Jun 30, 2018
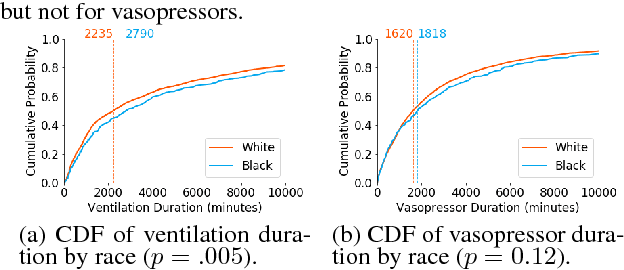
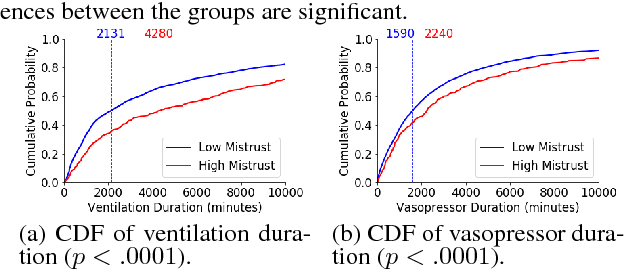
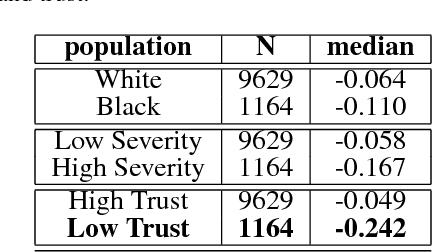
Abstract:In this work, we characterize the doctor-patient relationship using a machine learning-derived trust score. We show that this score has statistically significant racial associations, and that by modeling trust directly we find stronger disparities in care than by stratifying on race. We further demonstrate that mistrust is indicative of worse outcomes, but is only weakly associated with physiologically-created severity scores. Finally, we describe sentiment analysis experiments indicating patients with higher levels of mistrust have worse experiences and interactions with their caregivers. This work is a step towards measuring fairer machine learning in the healthcare domain.
Learning Tasks for Multitask Learning: Heterogenous Patient Populations in the ICU
Jun 07, 2018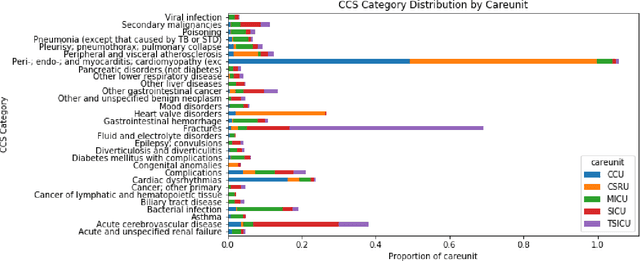
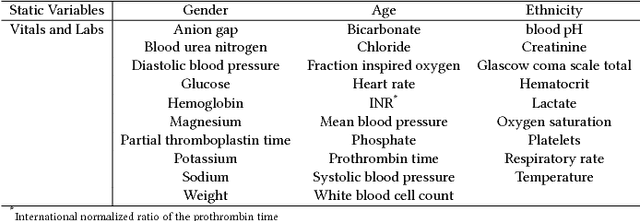
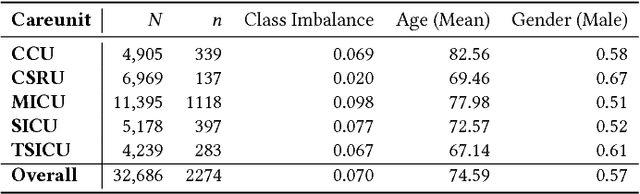


Abstract:Machine learning approaches have been effective in predicting adverse outcomes in different clinical settings. These models are often developed and evaluated on datasets with heterogeneous patient populations. However, good predictive performance on the aggregate population does not imply good performance for specific groups. In this work, we present a two-step framework to 1) learn relevant patient subgroups, and 2) predict an outcome for separate patient populations in a multi-task framework, where each population is a separate task. We demonstrate how to discover relevant groups in an unsupervised way with a sequence-to-sequence autoencoder. We show that using these groups in a multi-task framework leads to better predictive performance of in-hospital mortality both across groups and overall. We also highlight the need for more granular evaluation of performance when dealing with heterogeneous populations.
Clinical Intervention Prediction and Understanding using Deep Networks
May 23, 2017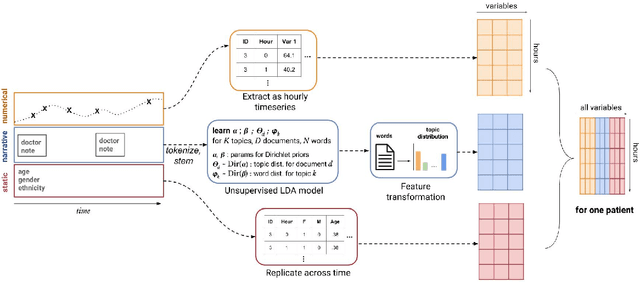
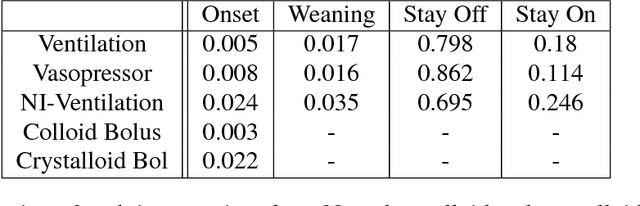
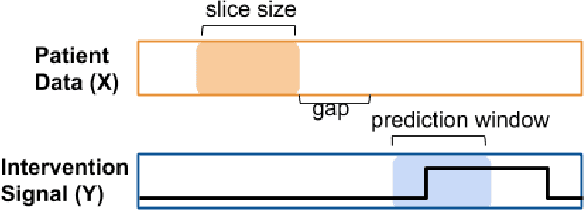
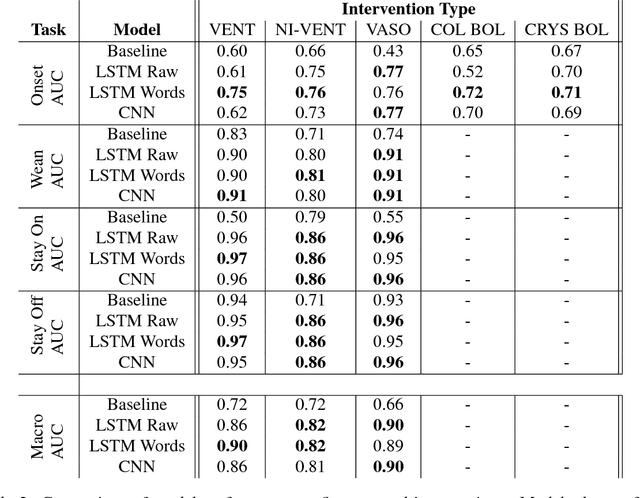
Abstract:Real-time prediction of clinical interventions remains a challenge within intensive care units (ICUs). This task is complicated by data sources that are noisy, sparse, heterogeneous and outcomes that are imbalanced. In this paper, we integrate data from all available ICU sources (vitals, labs, notes, demographics) and focus on learning rich representations of this data to predict onset and weaning of multiple invasive interventions. In particular, we compare both long short-term memory networks (LSTM) and convolutional neural networks (CNN) for prediction of five intervention tasks: invasive ventilation, non-invasive ventilation, vasopressors, colloid boluses, and crystalloid boluses. Our predictions are done in a forward-facing manner to enable "real-time" performance, and predictions are made with a six hour gap time to support clinically actionable planning. We achieve state-of-the-art results on our predictive tasks using deep architectures. We explore the use of feature occlusion to interpret LSTM models, and compare this to the interpretability gained from examining inputs that maximally activate CNN outputs. We show that our models are able to significantly outperform baselines in intervention prediction, and provide insight into model learning, which is crucial for the adoption of such models in practice.
The Use of Autoencoders for Discovering Patient Phenotypes
Mar 20, 2017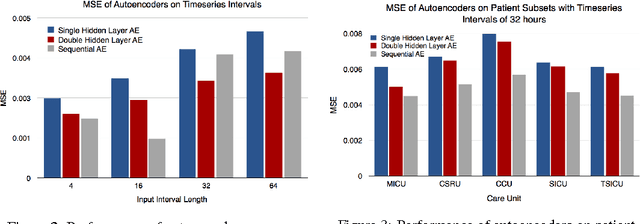
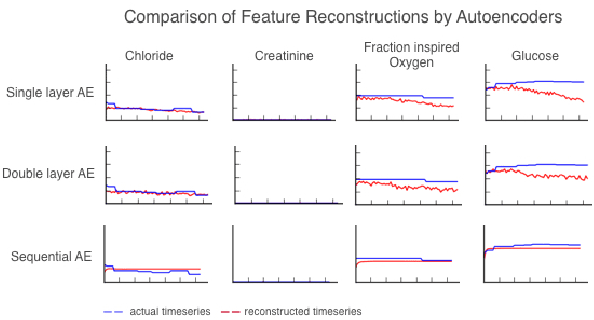
Abstract:We use autoencoders to create low-dimensional embeddings of underlying patient phenotypes that we hypothesize are a governing factor in determining how different patients will react to different interventions. We compare the performance of autoencoders that take fixed length sequences of concatenated timesteps as input with a recurrent sequence-to-sequence autoencoder. We evaluate our methods on around 35,500 patients from the latest MIMIC III dataset from Beth Israel Deaconess Hospital.
Feature Representation for ICU Mortality
Feb 07, 2016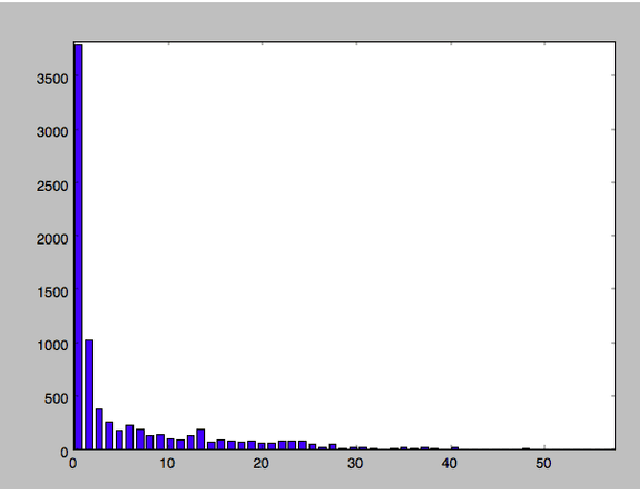
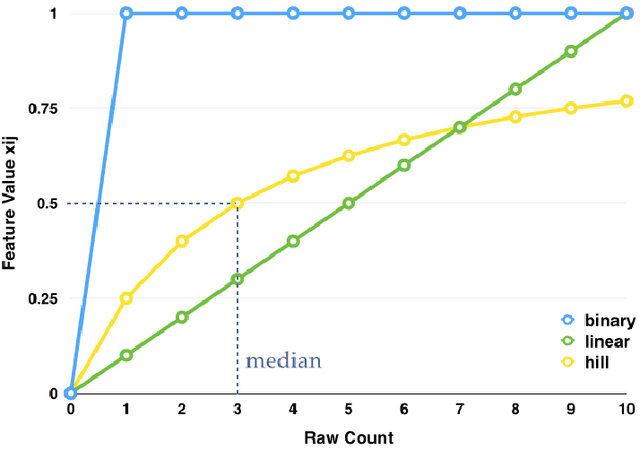
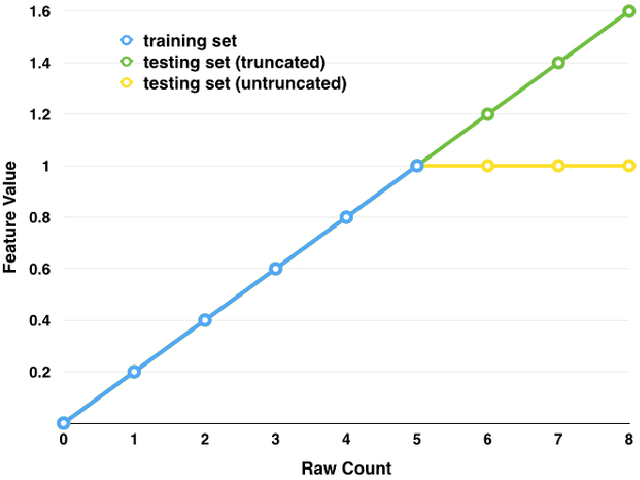
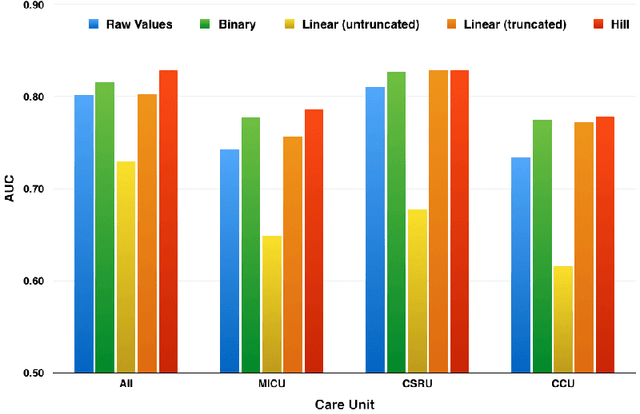
Abstract:Good predictors of ICU Mortality have the potential to identify high-risk patients earlier, improve ICU resource allocation, or create more accurate population-level risk models. Machine learning practitioners typically make choices about how to represent features in a particular model, but these choices are seldom evaluated quantitatively. This study compares the performance of different representations of clinical event data from MIMIC II in a logistic regression model to predict 36-hour ICU mortality. The most common representations are linear (normalized counts) and binary (yes/no). These, along with a new representation termed "hill", are compared using both L1 and L2 regularization. Results indicate that the introduced "hill" representation outperforms both the binary and linear representations, the hill representation thus has the potential to improve existing models of ICU mortality.
Autodetection and Classification of Hidden Cultural City Districts from Yelp Reviews
Jan 12, 2015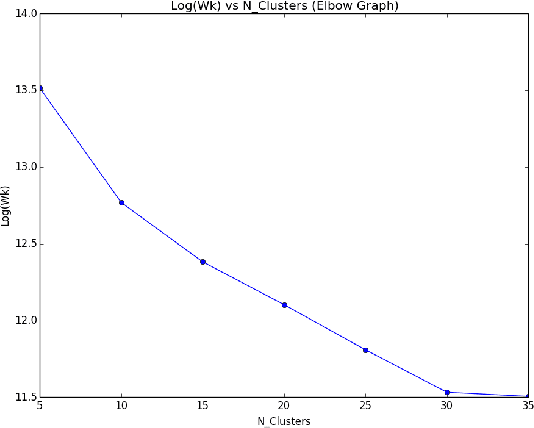
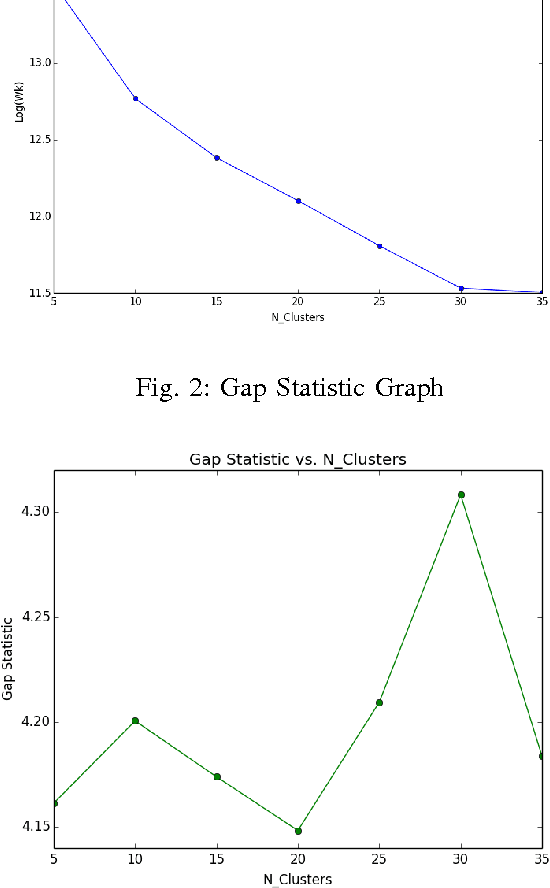
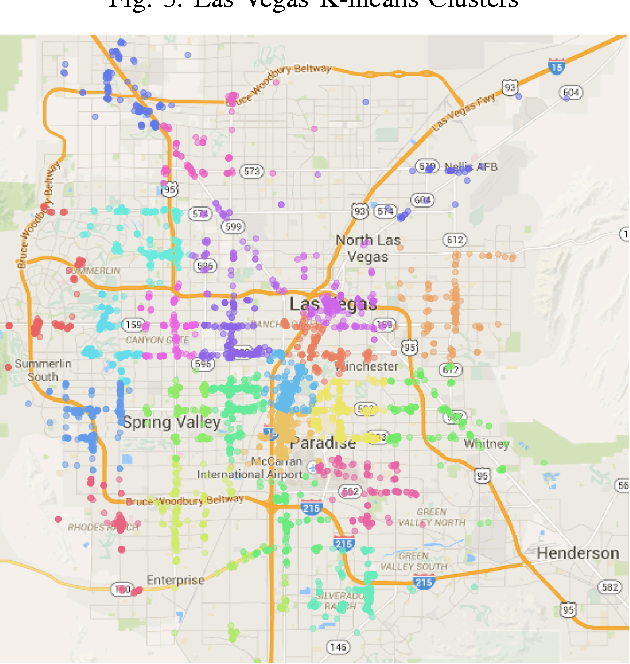
Abstract:Topic models are a way to discover underlying themes in an otherwise unstructured collection of documents. In this study, we specifically used the Latent Dirichlet Allocation (LDA) topic model on a dataset of Yelp reviews to classify restaurants based off of their reviews. Furthermore, we hypothesize that within a city, restaurants can be grouped into similar "clusters" based on both location and similarity. We used several different clustering methods, including K-means Clustering and a Probabilistic Mixture Model, in order to uncover and classify districts, both well-known and hidden (i.e. cultural areas like Chinatown or hearsay like "the best street for Italian restaurants") within a city. We use these models to display and label different clusters on a map. We also introduce a topic similarity heatmap that displays the similarity distribution in a city to a new restaurant.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge