Alistair Johnson
EHRCon: Dataset for Checking Consistency between Unstructured Notes and Structured Tables in Electronic Health Records
Jun 24, 2024



Abstract:Electronic Health Records (EHRs) are integral for storing comprehensive patient medical records, combining structured data (e.g., medications) with detailed clinical notes (e.g., physician notes). These elements are essential for straightforward data retrieval and provide deep, contextual insights into patient care. However, they often suffer from discrepancies due to unintuitive EHR system designs and human errors, posing serious risks to patient safety. To address this, we developed EHRCon, a new dataset and task specifically designed to ensure data consistency between structured tables and unstructured notes in EHRs. EHRCon was crafted in collaboration with healthcare professionals using the MIMIC-III EHR dataset, and includes manual annotations of 3,943 entities across 105 clinical notes checked against database entries for consistency. EHRCon has two versions, one using the original MIMIC-III schema, and another using the OMOP CDM schema, in order to increase its applicability and generalizability. Furthermore, leveraging the capabilities of large language models, we introduce CheckEHR, a novel framework for verifying the consistency between clinical notes and database tables. CheckEHR utilizes an eight-stage process and shows promising results in both few-shot and zero-shot settings. The code is available at https://github.com/dustn1259/EHRCon.
Pyclipse, a library for deidentification of free-text clinical notes
Nov 05, 2023



Abstract:Automated deidentification of clinical text data is crucial due to the high cost of manual deidentification, which has been a barrier to sharing clinical text and the advancement of clinical natural language processing. However, creating effective automated deidentification tools faces several challenges, including issues in reproducibility due to differences in text processing, evaluation methods, and a lack of consistency across clinical domains and institutions. To address these challenges, we propose the pyclipse framework, a unified and configurable evaluation procedure to streamline the comparison of deidentification algorithms. Pyclipse serves as a single interface for running open-source deidentification algorithms on local clinical data, allowing for context-specific evaluation. To demonstrate the utility of pyclipse, we compare six deidentification algorithms across four public and two private clinical text datasets. We find that algorithm performance consistently falls short of the results reported in the original papers, even when evaluated on the same benchmark dataset. These discrepancies highlight the complexity of accurately assessing and comparing deidentification algorithms, emphasizing the need for a reproducible, adjustable, and extensible framework like pyclipse. Our framework lays the foundation for a unified approach to evaluate and improve deidentification tools, ultimately enhancing patient protection in clinical natural language processing.
Do We Still Need Clinical Language Models?
Feb 16, 2023



Abstract:Although recent advances in scaling large language models (LLMs) have resulted in improvements on many NLP tasks, it remains unclear whether these models trained primarily with general web text are the right tool in highly specialized, safety critical domains such as clinical text. Recent results have suggested that LLMs encode a surprising amount of medical knowledge. This raises an important question regarding the utility of smaller domain-specific language models. With the success of general-domain LLMs, is there still a need for specialized clinical models? To investigate this question, we conduct an extensive empirical analysis of 12 language models, ranging from 220M to 175B parameters, measuring their performance on 3 different clinical tasks that test their ability to parse and reason over electronic health records. As part of our experiments, we train T5-Base and T5-Large models from scratch on clinical notes from MIMIC III and IV to directly investigate the efficiency of clinical tokens. We show that relatively small specialized clinical models substantially outperform all in-context learning approaches, even when finetuned on limited annotated data. Further, we find that pretraining on clinical tokens allows for smaller, more parameter-efficient models that either match or outperform much larger language models trained on general text. We release the code and the models used under the PhysioNet Credentialed Health Data license and data use agreement.
Clinical Intervention Prediction and Understanding using Deep Networks
May 23, 2017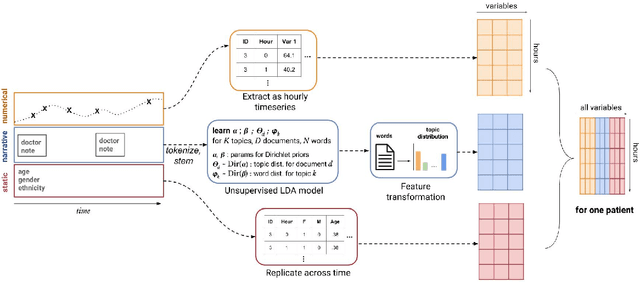
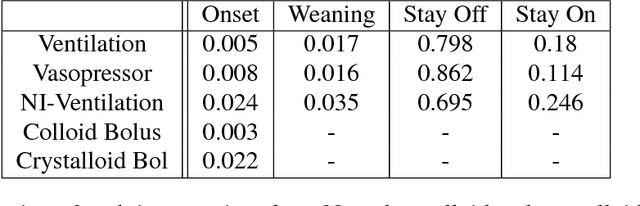
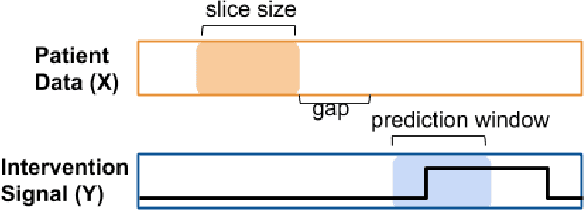
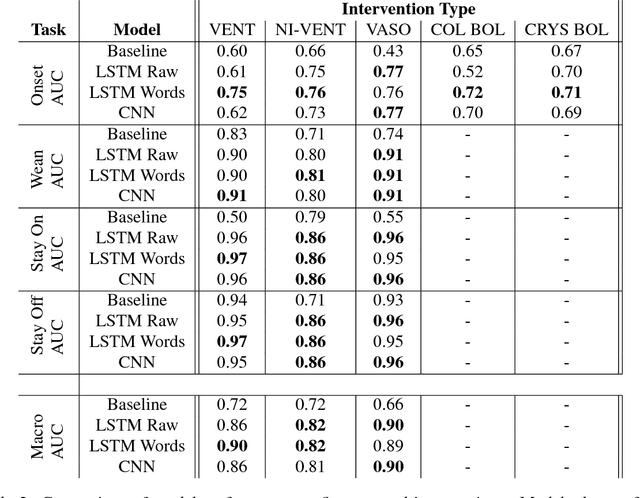
Abstract:Real-time prediction of clinical interventions remains a challenge within intensive care units (ICUs). This task is complicated by data sources that are noisy, sparse, heterogeneous and outcomes that are imbalanced. In this paper, we integrate data from all available ICU sources (vitals, labs, notes, demographics) and focus on learning rich representations of this data to predict onset and weaning of multiple invasive interventions. In particular, we compare both long short-term memory networks (LSTM) and convolutional neural networks (CNN) for prediction of five intervention tasks: invasive ventilation, non-invasive ventilation, vasopressors, colloid boluses, and crystalloid boluses. Our predictions are done in a forward-facing manner to enable "real-time" performance, and predictions are made with a six hour gap time to support clinically actionable planning. We achieve state-of-the-art results on our predictive tasks using deep architectures. We explore the use of feature occlusion to interpret LSTM models, and compare this to the interpretability gained from examining inputs that maximally activate CNN outputs. We show that our models are able to significantly outperform baselines in intervention prediction, and provide insight into model learning, which is crucial for the adoption of such models in practice.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge