Hannah Thompson
Prediction of Rectal Cancer Regrowth from Longitudinal Endoscopy
May 13, 2026Abstract:Clinical trial studies indicate benefit of watch-and-wait (WW) surveillance for patients with rectal cancer showing a complete or near clinical response (CR) directly after treatment (restaging). However, there are no objectively accurate methods to early detect local tumor regrowth (LR) in patients undergoing WW from follow-up exams. Hence, we developed Temporal Rectal Endoscopy Cross-attention (TREX), a longitudinal deep learning approach that combines pairs of images acquired at restaging and follow-up to distinguish CR from LR. TREX uses pretrained Swin Transformers in a siamese setting to extract features from longitudinal images and dual cross-attention to combine the features without spatial co-registration between image pairs. TREX and Swin-based baselines were trained under two settings: (a) detecting LR or CR at the last available follow-up and (b) early detection of LR at 3--6, 6--12, and 12--24 months before clinical confirmation. TREX achieved the highest accuracy in detecting LR with a high sensitivity of 97% $\pm$ 6% and a balanced accuracy of 90% $\pm$ 3%, and outperformed all baselines in early detection at both 3--6 (74% $\pm$ 1%) and 6--12 months (62% $\pm$ 4%) prior to clinical detection. Clinical validation via a surgeon survey showed that TREX matched attending-level overall accuracy (TREX: 86.21% vs.\ Clinicians: 87.84% $\pm$ 1.28%). Finally, we explored TREX's ability to predict treatment response by combining pre-treatment (pre-TNT) and restaging endoscopies, achieving a balanced accuracy of 73% $\pm$ 12%. These results show that longitudinal deep learning analysis of endoscopy may improve surveillance and enable earlier identification of rectal cancer regrowth.
Deep learning classifier of locally advanced rectal cancer treatment response from endoscopy images
May 06, 2024



Abstract:We developed a deep learning classifier of rectal cancer response (tumor vs. no-tumor) to total neoadjuvant treatment (TNT) from endoscopic images acquired before, during, and following TNT. We further evaluated the network's ability in a near out-of-distribution (OOD) problem to identify local regrowth (LR) from follow-up endoscopy images acquired several months to years after completing TNT. We addressed endoscopic image variability by using optimal mass transport-based image harmonization. We evaluated multiple training regularization schemes to study the ResNet-50 network's in-distribution and near-OOD generalization ability. Test time augmentation resulted in the most considerable accuracy improvement. Image harmonization resulted in slight accuracy improvement for the near-OOD cases. Our results suggest that off-the-shelf deep learning classifiers can detect rectal cancer from endoscopic images at various stages of therapy for surveillance.
A Framework for Implementing Machine Learning on Omics Data
Nov 26, 2018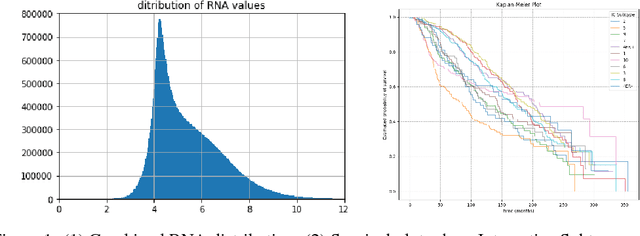
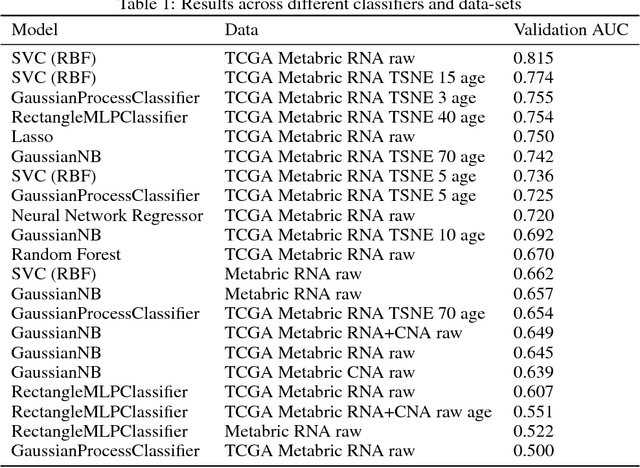
Abstract:The potential benefits of applying machine learning methods to -omics data are becoming increasingly apparent, especially in clinical settings. However, the unique characteristics of these data are not always well suited to machine learning techniques. These data are often generated across different technologies in different labs, and frequently with high dimensionality. In this paper we present a framework for combining -omics data sets, and for handling high dimensional data, making -omics research more accessible to machine learning applications. We demonstrate the success of this framework through integration and analysis of multi-analyte data for a set of 3,533 breast cancers. We then use this data-set to predict breast cancer patient survival for individuals at risk of an impending event, with higher accuracy and lower variance than methods trained on individual data-sets. We hope that our pipelines for data-set generation and transformation will open up -omics data to machine learning researchers. We have made these freely available for noncommercial use at www.ccg.ai.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge