Héctor Carrión
TARDIS STRIDE: A Spatio-Temporal Road Image Dataset for Exploration and Autonomy
Jun 12, 2025Abstract:World models aim to simulate environments and enable effective agent behavior. However, modeling real-world environments presents unique challenges as they dynamically change across both space and, crucially, time. To capture these composed dynamics, we introduce a Spatio-Temporal Road Image Dataset for Exploration (STRIDE) permuting 360-degree panoramic imagery into rich interconnected observation, state and action nodes. Leveraging this structure, we can simultaneously model the relationship between egocentric views, positional coordinates, and movement commands across both space and time. We benchmark this dataset via TARDIS, a transformer-based generative world model that integrates spatial and temporal dynamics through a unified autoregressive framework trained on STRIDE. We demonstrate robust performance across a range of agentic tasks such as controllable photorealistic image synthesis, instruction following, autonomous self-control, and state-of-the-art georeferencing. These results suggest a promising direction towards sophisticated generalist agents--capable of understanding and manipulating the spatial and temporal aspects of their material environments--with enhanced embodied reasoning capabilities. Training code, datasets, and model checkpoints are made available at https://huggingface.co/datasets/Tera-AI/STRIDE.
FEDD -- Fair, Efficient, and Diverse Diffusion-based Lesion Segmentation and Malignancy Classification
Jul 21, 2023



Abstract:Skin diseases affect millions of people worldwide, across all ethnicities. Increasing diagnosis accessibility requires fair and accurate segmentation and classification of dermatology images. However, the scarcity of annotated medical images, especially for rare diseases and underrepresented skin tones, poses a challenge to the development of fair and accurate models. In this study, we introduce a Fair, Efficient, and Diverse Diffusion-based framework for skin lesion segmentation and malignancy classification. FEDD leverages semantically meaningful feature embeddings learned through a denoising diffusion probabilistic backbone and processes them via linear probes to achieve state-of-the-art performance on Diverse Dermatology Images (DDI). We achieve an improvement in intersection over union of 0.18, 0.13, 0.06, and 0.07 while using only 5%, 10%, 15%, and 20% labeled samples, respectively. Additionally, FEDD trained on 10% of DDI demonstrates malignancy classification accuracy of 81%, 14% higher compared to the state-of-the-art. We showcase high efficiency in data-constrained scenarios while providing fair performance for diverse skin tones and rare malignancy conditions. Our newly annotated DDI segmentation masks and training code can be found on https://github.com/hectorcarrion/fedd.
HealNet -- Self-Supervised Acute Wound Heal-Stage Classification
Jun 23, 2022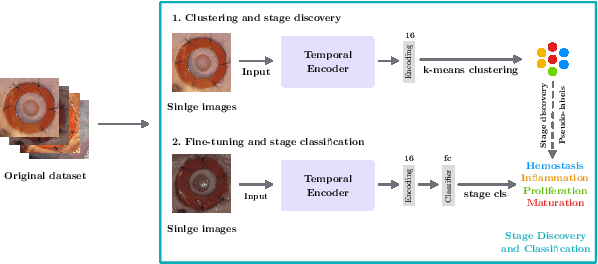
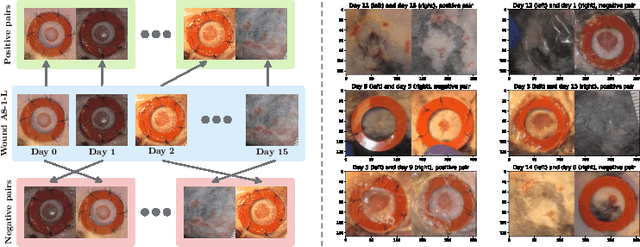
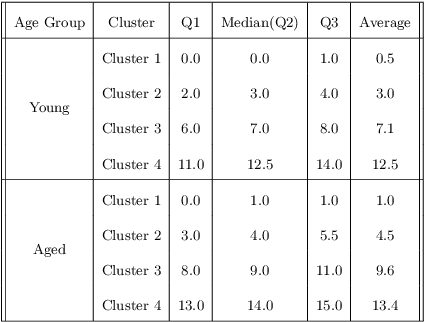

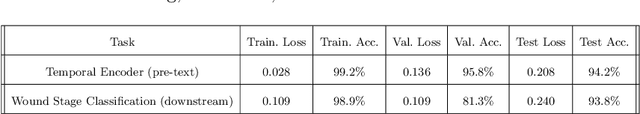
Abstract:Identifying, tracking, and predicting wound heal-stage progression is a fundamental task towards proper diagnosis, effective treatment, facilitating healing, and reducing pain. Traditionally, a medical expert might observe a wound to determine the current healing state and recommend treatment. However, sourcing experts who can produce such a diagnosis solely from visual indicators can be difficult, time-consuming and expensive. In addition, lesions may take several weeks to undergo the healing process, demanding resources to monitor and diagnose continually. Automating this task can be challenging; datasets that follow wound progression from onset to maturation are small, rare, and often collected without computer vision in mind. To tackle these challenges, we introduce a self-supervised learning scheme composed of (a) learning embeddings of wound's temporal dynamics, (b) clustering for automatic stage discovery, and (c) fine-tuned classification. The proposed self-supervised and flexible learning framework is biologically inspired and trained on a small dataset with zero human labeling. The HealNet framework achieved high pre-text and downstream classification accuracy; when evaluated on held-out test data, HealNet achieved 97.7% pre-text accuracy and 90.62% heal-stage classification accuracy.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge