Ganesh Narasimha
Quality-Controlled Active Learning via Gaussian Processes for Robust Structure-Property Learning in Autonomous Microscopy
Mar 31, 2026Abstract:Autonomous experimental systems are increasingly used in materials research to accelerate scientific discovery, but their performance is often limited by low-quality, noisy data. This issue is especially problematic in data-intensive structure-property learning tasks such as Image-to-Spectrum (Im2Spec) and Spectrum-to-Image (Spec2Im) translations, where standard active learning strategies can mistakenly prioritize poor-quality measurements. We introduce a gated active learning framework that combines curiosity-driven sampling with a physics-informed quality control filter based on the Simple Harmonic Oscillator model fits, allowing the system to automatically exclude low-fidelity data during acquisition. Evaluations on a pre-acquired dataset of band-excitation piezoresponse spectroscopy (BEPS) data from PbTiO3 thin films with spatially localized noise show that the proposed method outperforms random sampling, standard active learning, and multitask learning strategies. The gated approach enhances both Im2Spec and Spec2Im by handling noise during training and acquisition, leading to more reliable forward and inverse predictions. In contrast, standard active learners often misinterpret noise as uncertainty and end up acquiring bad samples that hurt performance. Given its promising applicability, we further deployed the framework in real-time experiments on BiFeO3 thin films, demonstrating its effectiveness in real autonomous microscopy experiments. Overall, this work supports a shift toward hybrid autonomy in self-driving labs, where physics-informed quality assessment and active decision-making work hand-in-hand for more reliable discovery.
Mic-hackathon 2024: Hackathon on Machine Learning for Electron and Scanning Probe Microscopy
Jun 10, 2025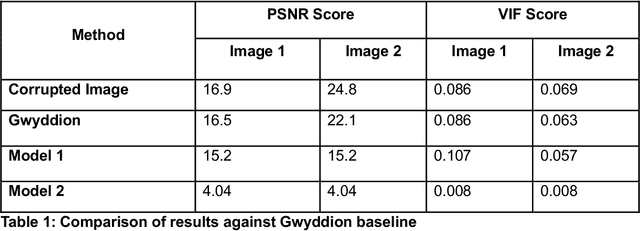
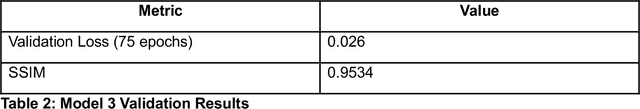
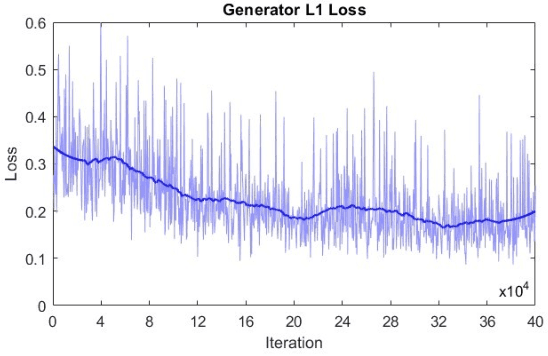
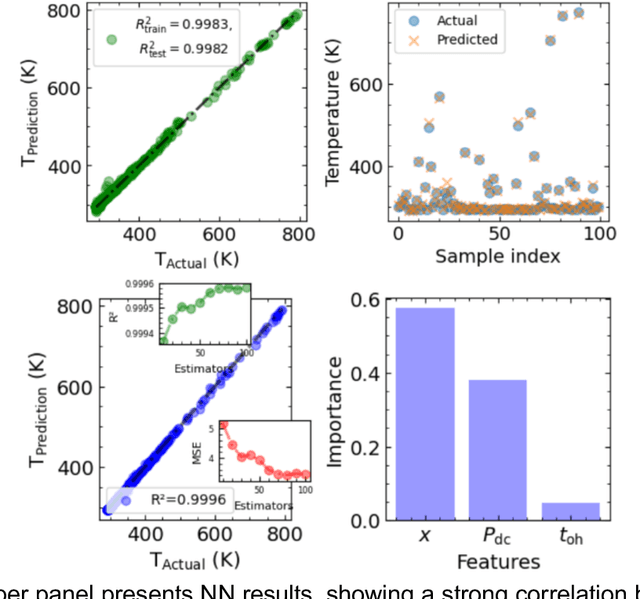
Abstract:Microscopy is a primary source of information on materials structure and functionality at nanometer and atomic scales. The data generated is often well-structured, enriched with metadata and sample histories, though not always consistent in detail or format. The adoption of Data Management Plans (DMPs) by major funding agencies promotes preservation and access. However, deriving insights remains difficult due to the lack of standardized code ecosystems, benchmarks, and integration strategies. As a result, data usage is inefficient and analysis time is extensive. In addition to post-acquisition analysis, new APIs from major microscope manufacturers enable real-time, ML-based analytics for automated decision-making and ML-agent-controlled microscope operation. Yet, a gap remains between the ML and microscopy communities, limiting the impact of these methods on physics, materials discovery, and optimization. Hackathons help bridge this divide by fostering collaboration between ML researchers and microscopy experts. They encourage the development of novel solutions that apply ML to microscopy, while preparing a future workforce for instrumentation, materials science, and applied ML. This hackathon produced benchmark datasets and digital twins of microscopes to support community growth and standardized workflows. All related code is available at GitHub: https://github.com/KalininGroup/Mic-hackathon-2024-codes-publication/tree/1.0.0.1
Curiosity Driven Exploration to Optimize Structure-Property Learning in Microscopy
Apr 28, 2025



Abstract:Rapidly determining structure-property correlations in materials is an important challenge in better understanding fundamental mechanisms and greatly assists in materials design. In microscopy, imaging data provides a direct measurement of the local structure, while spectroscopic measurements provide relevant functional property information. Deep kernel active learning approaches have been utilized to rapidly map local structure to functional properties in microscopy experiments, but are computationally expensive for multi-dimensional and correlated output spaces. Here, we present an alternative lightweight curiosity algorithm which actively samples regions with unexplored structure-property relations, utilizing a deep-learning based surrogate model for error prediction. We show that the algorithm outperforms random sampling for predicting properties from structures, and provides a convenient tool for efficient mapping of structure-property relationships in materials science.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge