Francis Motta
Topological Machine Learning with Unreduced Persistence Diagrams
Jul 09, 2025Abstract:Supervised machine learning pipelines trained on features derived from persistent homology have been experimentally observed to ignore much of the information contained in a persistence diagram. Computing persistence diagrams is often the most computationally demanding step in such a pipeline, however. To explore this, we introduce several methods to generate topological feature vectors from unreduced boundary matrices. We compared the performance of pipelines trained on vectorizations of unreduced PDs to vectorizations of fully-reduced PDs across several data and task types. Our results indicate that models trained on PDs built from unreduced diagrams can perform on par and even outperform those trained on fully-reduced diagrams on some tasks. This observation suggests that machine learning pipelines which incorporate topology-based features may benefit in terms of computational cost and performance by utilizing information contained in unreduced boundary matrices.
A Pipeline for Data-Driven Learning of Topological Features with Applications to Protein Stability Prediction
Aug 09, 2024Abstract:In this paper, we propose a data-driven method to learn interpretable topological features of biomolecular data and demonstrate the efficacy of parsimonious models trained on topological features in predicting the stability of synthetic mini proteins. We compare models that leverage automatically-learned structural features against models trained on a large set of biophysical features determined by subject-matter experts (SME). Our models, based only on topological features of the protein structures, achieved 92%-99% of the performance of SME-based models in terms of the average precision score. By interrogating model performance and feature importance metrics, we extract numerous insights that uncover high correlations between topological features and SME features. We further showcase how combining topological features and SME features can lead to improved model performance over either feature set used in isolation, suggesting that, in some settings, topological features may provide new discriminating information not captured in existing SME features that are useful for protein stability prediction.
Persistence Images: A Stable Vector Representation of Persistent Homology
Jul 11, 2016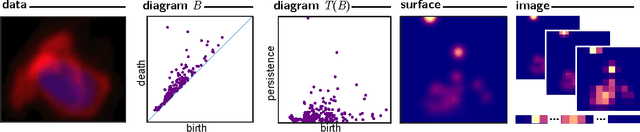
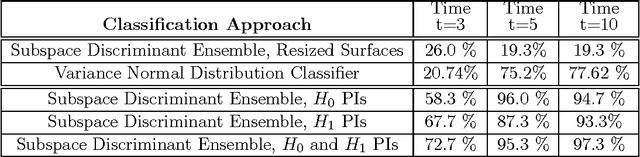
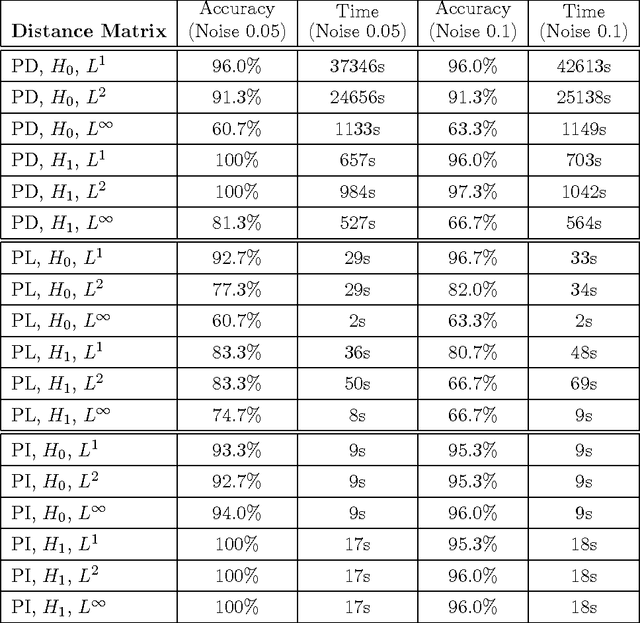
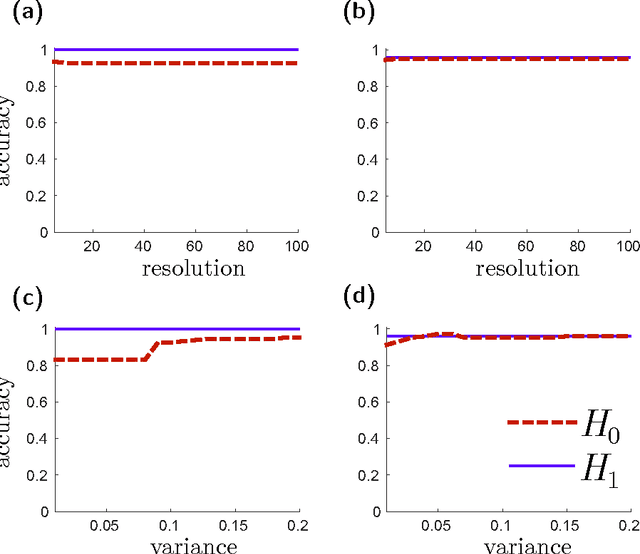

Abstract:Many datasets can be viewed as a noisy sampling of an underlying space, and tools from topological data analysis can characterize this structure for the purpose of knowledge discovery. One such tool is persistent homology, which provides a multiscale description of the homological features within a dataset. A useful representation of this homological information is a persistence diagram (PD). Efforts have been made to map PDs into spaces with additional structure valuable to machine learning tasks. We convert a PD to a finite-dimensional vector representation which we call a persistence image (PI), and prove the stability of this transformation with respect to small perturbations in the inputs. The discriminatory power of PIs is compared against existing methods, showing significant performance gains. We explore the use of PIs with vector-based machine learning tools, such as linear sparse support vector machines, which identify features containing discriminating topological information. Finally, high accuracy inference of parameter values from the dynamic output of a discrete dynamical system (the linked twist map) and a partial differential equation (the anisotropic Kuramoto-Sivashinsky equation) provide a novel application of the discriminatory power of PIs.
* Version 3 contains updated theoretical results supporting methodology; expanded discussion of related works; extended list of references; extended applications section; additional experimental results and new figures
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge