Florian Engert
ZAPBench: A Benchmark for Whole-Brain Activity Prediction in Zebrafish
Mar 04, 2025



Abstract:Data-driven benchmarks have led to significant progress in key scientific modeling domains including weather and structural biology. Here, we introduce the Zebrafish Activity Prediction Benchmark (ZAPBench) to measure progress on the problem of predicting cellular-resolution neural activity throughout an entire vertebrate brain. The benchmark is based on a novel dataset containing 4d light-sheet microscopy recordings of over 70,000 neurons in a larval zebrafish brain, along with motion stabilized and voxel-level cell segmentations of these data that facilitate development of a variety of forecasting methods. Initial results from a selection of time series and volumetric video modeling approaches achieve better performance than naive baseline methods, but also show room for further improvement. The specific brain used in the activity recording is also undergoing synaptic-level anatomical mapping, which will enable future integration of detailed structural information into forecasting methods.
Prospective Learning: Back to the Future
Jan 19, 2022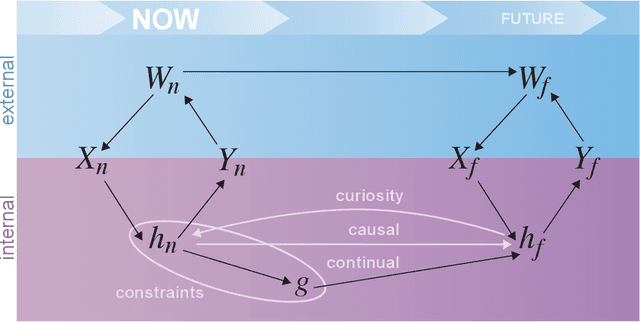
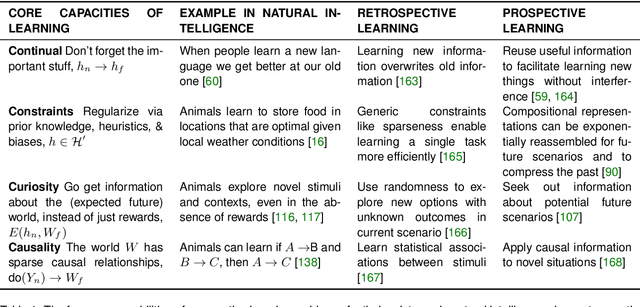

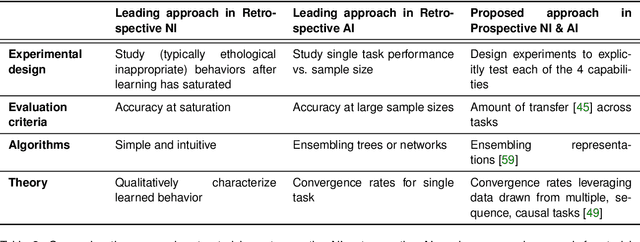
Abstract:Research on both natural intelligence (NI) and artificial intelligence (AI) generally assumes that the future resembles the past: intelligent agents or systems (what we call 'intelligence') observe and act on the world, then use this experience to act on future experiences of the same kind. We call this 'retrospective learning'. For example, an intelligence may see a set of pictures of objects, along with their names, and learn to name them. A retrospective learning intelligence would merely be able to name more pictures of the same objects. We argue that this is not what true intelligence is about. In many real world problems, both NIs and AIs will have to learn for an uncertain future. Both must update their internal models to be useful for future tasks, such as naming fundamentally new objects and using these objects effectively in a new context or to achieve previously unencountered goals. This ability to learn for the future we call 'prospective learning'. We articulate four relevant factors that jointly define prospective learning. Continual learning enables intelligences to remember those aspects of the past which it believes will be most useful in the future. Prospective constraints (including biases and priors) facilitate the intelligence finding general solutions that will be applicable to future problems. Curiosity motivates taking actions that inform future decision making, including in previously unmet situations. Causal estimation enables learning the structure of relations that guide choosing actions for specific outcomes, even when the specific action-outcome contingencies have never been observed before. We argue that a paradigm shift from retrospective to prospective learning will enable the communities that study intelligence to unite and overcome existing bottlenecks to more effectively explain, augment, and engineer intelligences.
NucMM Dataset: 3D Neuronal Nuclei Instance Segmentation at Sub-Cubic Millimeter Scale
Jul 13, 2021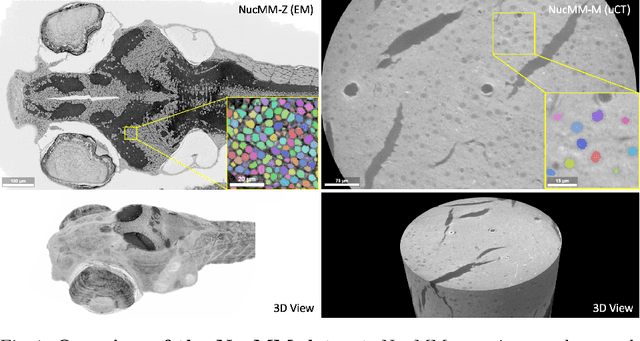

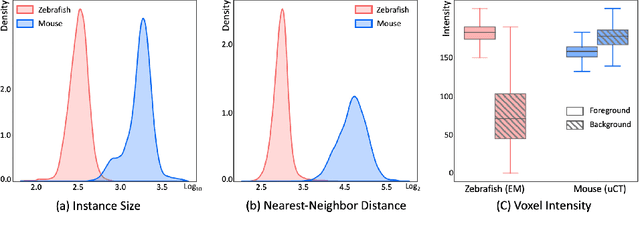
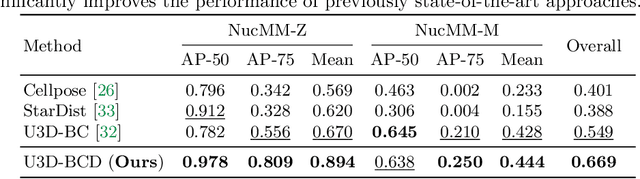
Abstract:Segmenting 3D cell nuclei from microscopy image volumes is critical for biological and clinical analysis, enabling the study of cellular expression patterns and cell lineages. However, current datasets for neuronal nuclei usually contain volumes smaller than $10^{\text{-}3}\ mm^3$ with fewer than 500 instances per volume, unable to reveal the complexity in large brain regions and restrict the investigation of neuronal structures. In this paper, we have pushed the task forward to the sub-cubic millimeter scale and curated the NucMM dataset with two fully annotated volumes: one $0.1\ mm^3$ electron microscopy (EM) volume containing nearly the entire zebrafish brain with around 170,000 nuclei; and one $0.25\ mm^3$ micro-CT (uCT) volume containing part of a mouse visual cortex with about 7,000 nuclei. With two imaging modalities and significantly increased volume size and instance numbers, we discover a great diversity of neuronal nuclei in appearance and density, introducing new challenges to the field. We also perform a statistical analysis to illustrate those challenges quantitatively. To tackle the challenges, we propose a novel hybrid-representation learning model that combines the merits of foreground mask, contour map, and signed distance transform to produce high-quality 3D masks. The benchmark comparisons on the NucMM dataset show that our proposed method significantly outperforms state-of-the-art nuclei segmentation approaches. Code and data are available at https://connectomics-bazaar.github.io/proj/nucMM/index.html.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge