Evis Sala
on behalf of the AIX-COVNET collaboration
Machine learning for COVID-19 detection and prognostication using chest radiographs and CT scans: a systematic methodological review
Sep 01, 2020
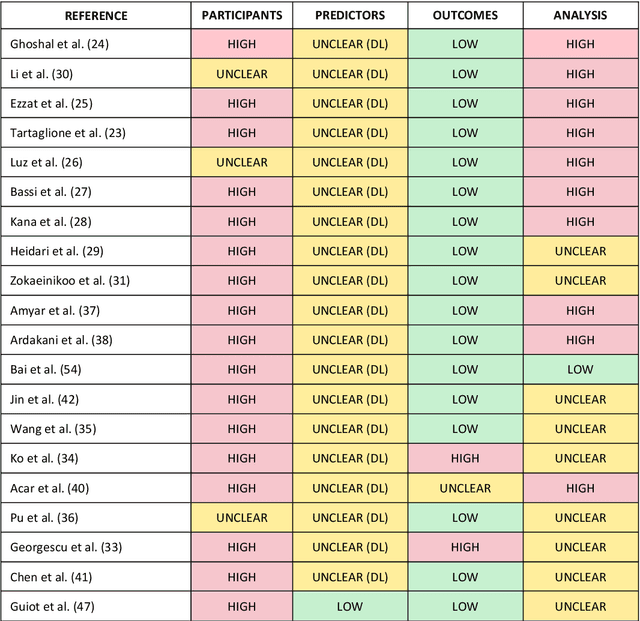
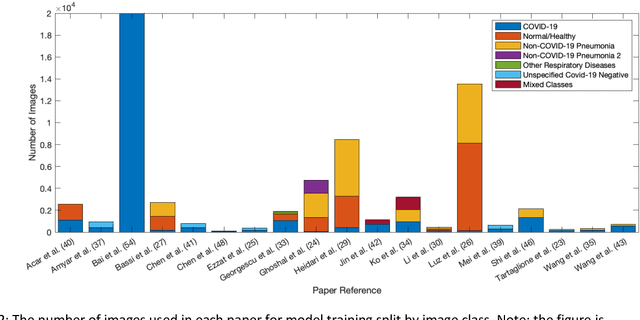
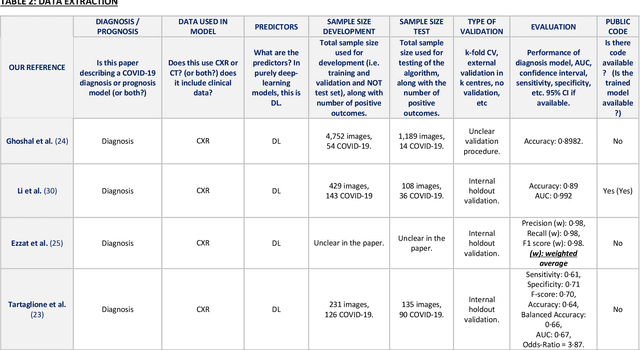
Abstract:Background: Machine learning methods offer great potential for fast and accurate detection and prognostication of COVID-19 from standard-of-care chest radiographs (CXR) and computed tomography (CT) images. In this systematic review we critically evaluate the machine learning methodologies employed in the rapidly growing literature. Methods: In this systematic review we reviewed EMBASE via OVID, MEDLINE via PubMed, bioRxiv, medRxiv and arXiv for published papers and preprints uploaded from Jan 1, 2020 to June 24, 2020. Studies which consider machine learning models for the diagnosis or prognosis of COVID-19 from CXR or CT images were included. A methodology quality review of each paper was performed against established benchmarks to ensure the review focusses only on high-quality reproducible papers. This study is registered with PROSPERO [CRD42020188887]. Interpretation: Our review finds that none of the developed models discussed are of potential clinical use due to methodological flaws and underlying biases. This is a major weakness, given the urgency with which validated COVID-19 models are needed. Typically, we find that the documentation of a model's development is not sufficient to make the results reproducible and therefore of 168 candidate papers only 29 are deemed to be reproducible and subsequently considered in this review. We therefore encourage authors to use established machine learning checklists to ensure sufficient documentation is made available, and to follow the PROBAST (prediction model risk of bias assessment tool) framework to determine the underlying biases in their model development process and to mitigate these where possible. This is key to safe clinical implementation which is urgently needed.
MADGAN: unsupervised Medical Anomaly Detection GAN using multiple adjacent brain MRI slice reconstruction
Jul 24, 2020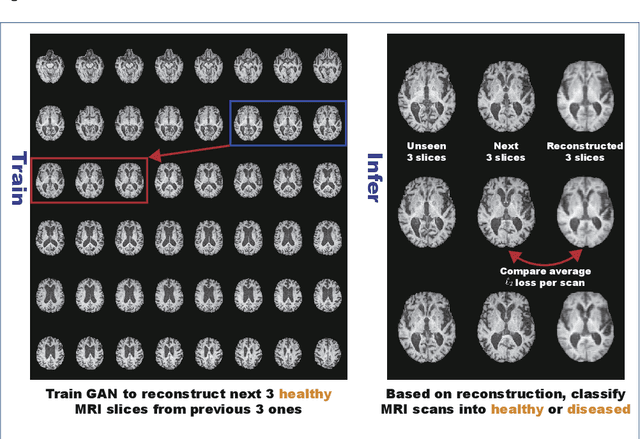
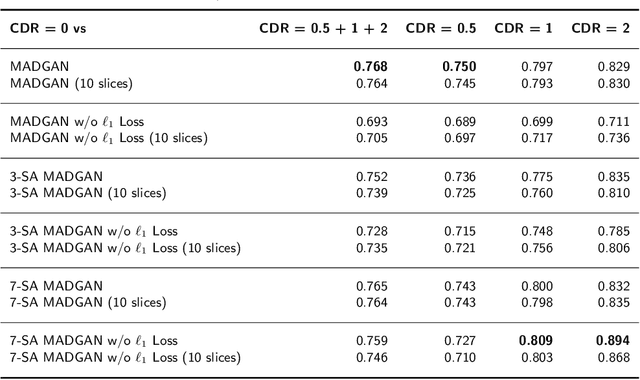
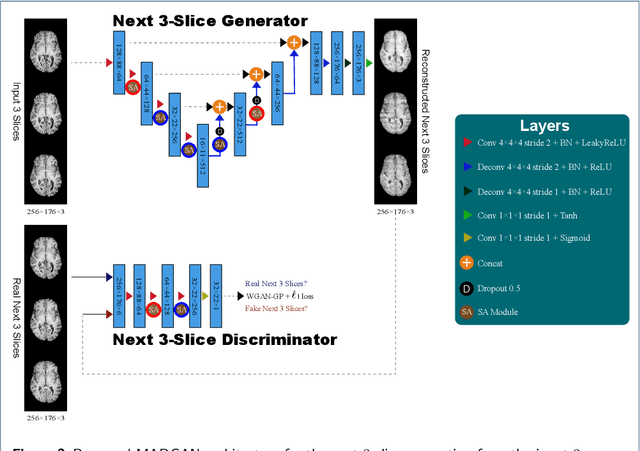
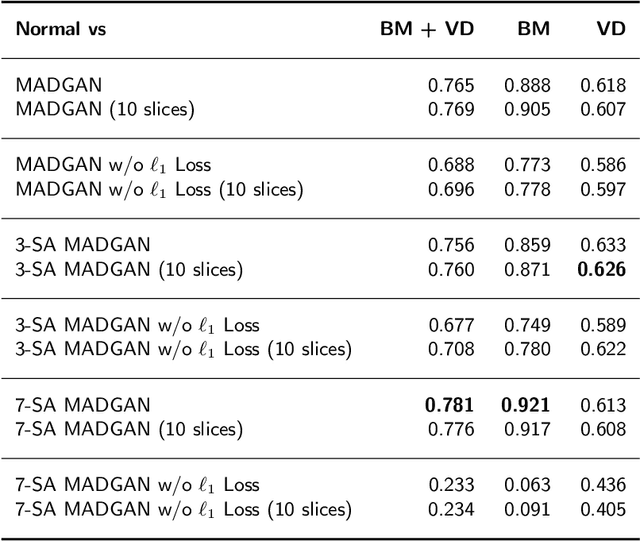
Abstract:Unsupervised learning can discover various unseen diseases, relying on large-scale unannotated medical images of healthy subjects. Towards this, unsupervised methods reconstruct a 2D/3D single medical image to detect outliers either in the learned feature space or from high reconstruction loss. However, without considering continuity between multiple adjacent slices, they cannot directly discriminate diseases composed of the accumulation of subtle anatomical anomalies, such as Alzheimer's Disease (AD). Moreover, no study has shown how unsupervised anomaly detection is associated with either disease stages, various (i.e., more than two types of) diseases, or multi-sequence Magnetic Resonance Imaging (MRI) scans. Therefore, we propose unsupervised Medical Anomaly Detection Generative Adversarial Network (MADGAN), a novel two-step method using GAN-based multiple adjacent brain MRI slice reconstruction to detect various diseases at different stages on multi-sequence structural MRI: (Reconstruction) Wasserstein loss with Gradient Penalty + 100 L1 loss-trained on 3 healthy brain axial MRI slices to reconstruct the next 3 ones-reconstructs unseen healthy/abnormal scans; (Diagnosis) Average L2 loss per scan discriminates them, comparing the ground truth/reconstructed slices. For training, we use 1,133 healthy T1-weighted (T1) and 135 healthy contrast-enhanced T1 (T1c) brain MRI scans. Our Self-Attention MADGAN can detect AD on T1 scans at a very early stage, Mild Cognitive Impairment (MCI), with Area Under the Curve (AUC) 0.727, and AD at a late stage with AUC 0.894, while detecting brain metastases on T1c scans with AUC 0.921.
3D deformable registration of longitudinal abdominopelvic CT images using unsupervised deep learning
May 15, 2020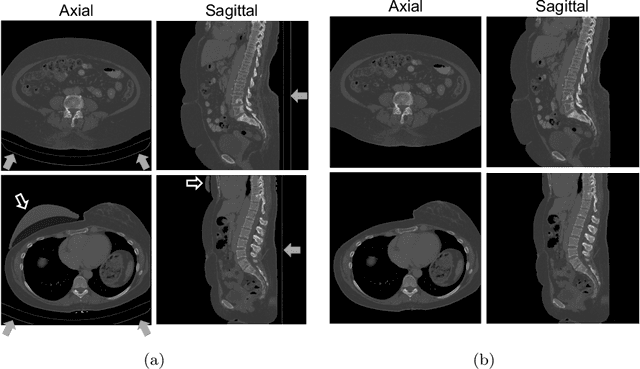
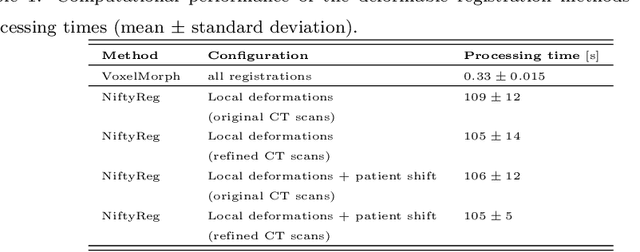
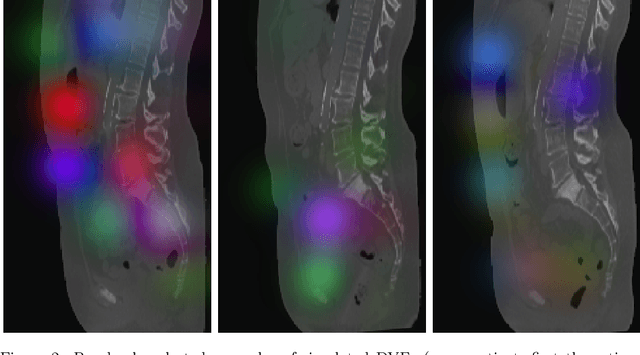
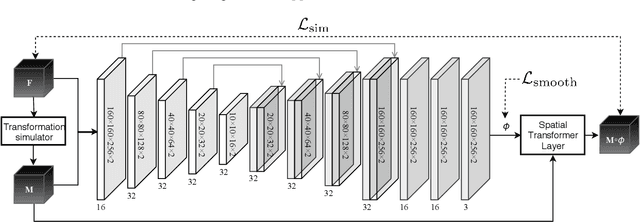
Abstract:This study investigates the use of the unsupervised deep learning framework VoxelMorph for deformable registration of longitudinal abdominopelvic CT images acquired in patients with bone metastases from breast cancer. The CT images were refined prior to registration by automatically removing the CT table and all other extra-corporeal components. To improve the learning capabilities of VoxelMorph when only a limited amount of training data is available, a novel incremental training strategy is proposed based on simulated deformations of consecutive CT images. In a 4-fold cross-validation scheme, the incremental training strategy achieved significantly better registration performance compared to training on a single volume. Although our deformable image registration method did not outperform iterative registration using NiftyReg (considered as a benchmark) in terms of registration quality, the registrations were approximately 300 times faster. This study showed the feasibility of deep learning based deformable registration of longitudinal abdominopelvic CT images via a novel incremental training strategy based on simulated deformations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge