Dhritiman Das
Improving the Precision of CNNs for Magnetic Resonance Spectral Modeling
Sep 10, 2024


Abstract:Magnetic resonance spectroscopic imaging is a widely available imaging modality that can non-invasively provide a metabolic profile of the tissue of interest, yet is challenging to integrate clinically. One major reason is the expensive, expert data processing and analysis that is required. Using machine learning to predict MRS-related quantities offers avenues around this problem, but deep learning models bring their own challenges, especially model trust. Current research trends focus primarily on mean error metrics, but comprehensive precision metrics are also needed, e.g. standard deviations, confidence intervals, etc.. This work highlights why more comprehensive error characterization is important and how to improve the precision of CNNs for spectral modeling, a quantitative task. The results highlight advantages and trade-offs of these techniques that should be considered when addressing such regression tasks with CNNs. Detailed insights into the underlying mechanisms of each technique, and how they interact with other techniques, are discussed in depth.
An Optical Flow-Based Approach for Minimally-Divergent Velocimetry Data Interpolation
Dec 20, 2018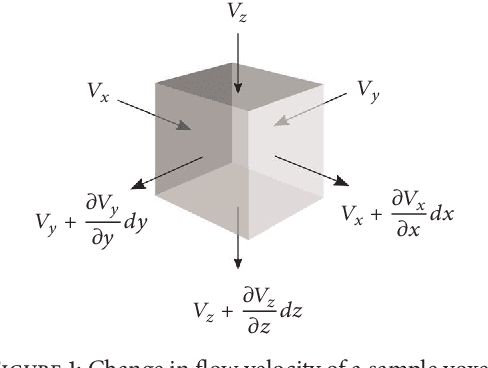
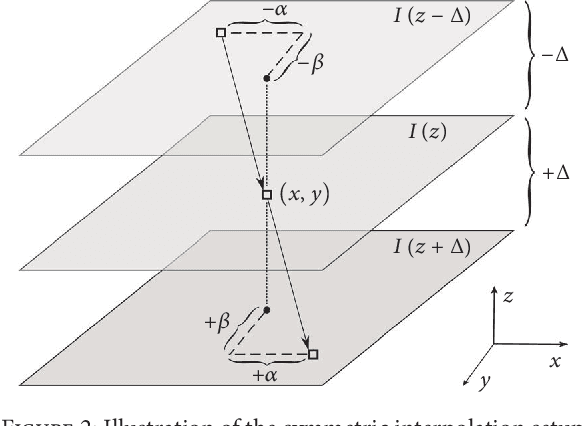
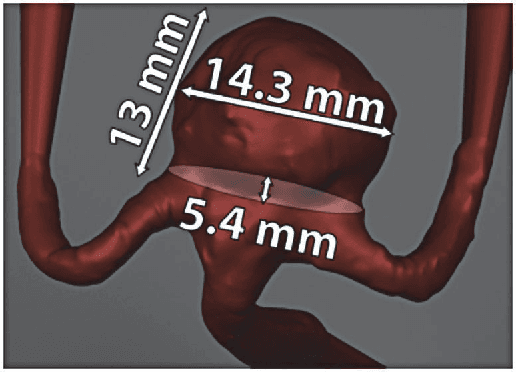
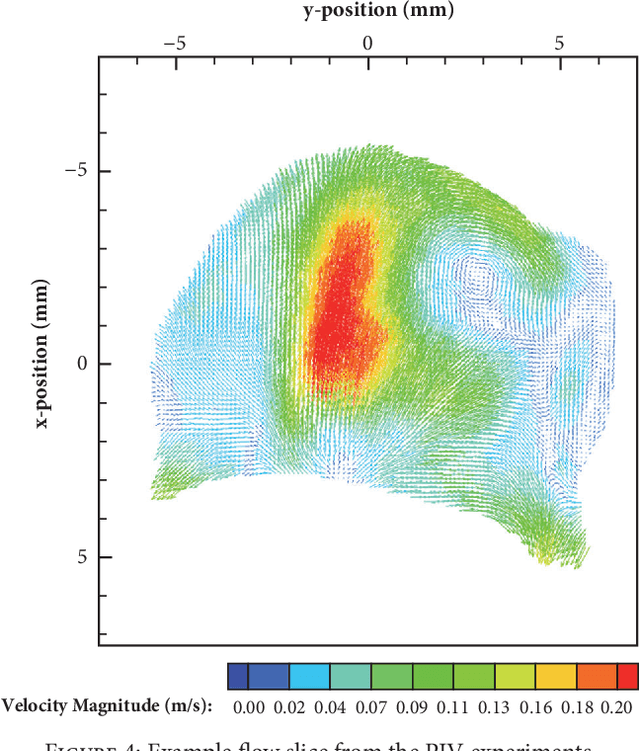
Abstract:Three-dimensional (3D) biomedical image sets are often acquired with in-plane pixel spacings that are far less than the out-of-plane spacings between images. The resultant anisotropy, which can be detrimental in many applications, can be decreased using image interpolation. Optical flow and/or other registration-based interpolators have proven useful in such interpolation roles in the past. When acquired images are comprised of signals that describe the flow velocity of fluids, additional information is available to guide the interpolation process. In this paper, we present an optical-flow based framework for image interpolation that also minimizes resultant divergence in the interpolated data.
Qunatification of Metabolites in MR Spectroscopic Imaging using Machine Learning
May 25, 2018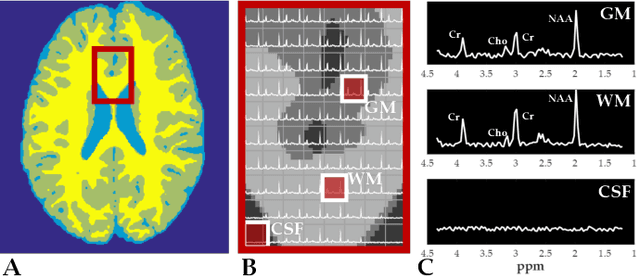
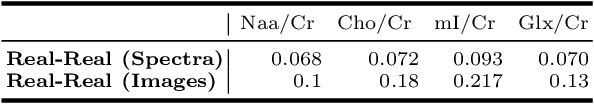
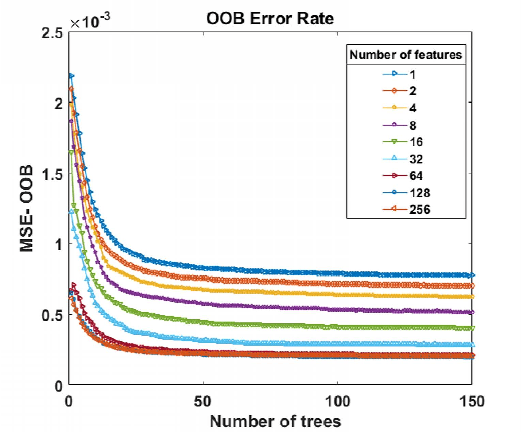
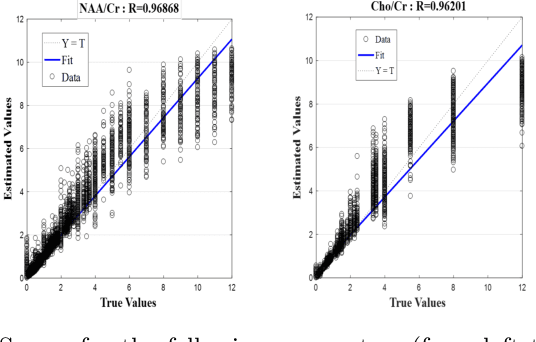
Abstract:Magnetic Resonance Spectroscopic Imaging (MRSI) is a clinical imaging modality for measuring tissue metabolite levels in-vivo. An accurate estimation of spectral parameters allows for better assessment of spectral quality and metabolite concentration levels. The current gold standard quantification method is the LCModel - a commercial fitting tool. However, this fails for spectra having poor signal-to-noise ratio (SNR) or a large number of artifacts. This paper introduces a framework based on random forest regression for accurate estimation of the output parameters of a model based analysis of MR spectroscopy data. The goal of our proposed framework is to learn the spectral features from a training set comprising of different variations of both simulated and in-vivo brain spectra and then use this learning for the subsequent metabolite quantification. Experiments involve training and testing on simulated and in-vivo human brain spectra. We estimate parameters such as concentration of metabolites and compare our results with that from the LCModel.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge