David A. Clausi
Puck localization and multi-task event recognition in broadcast hockey videos
May 21, 2021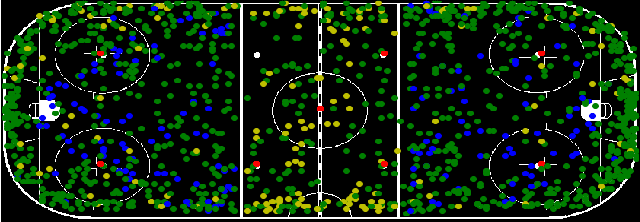
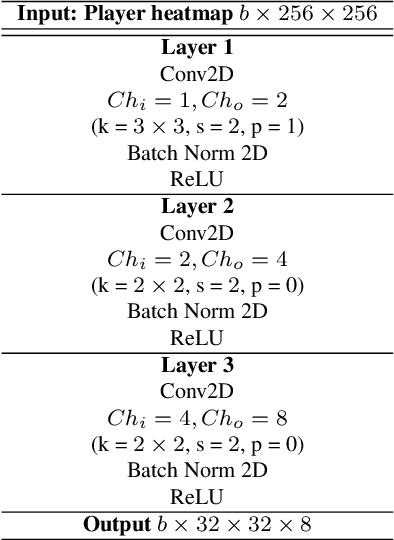
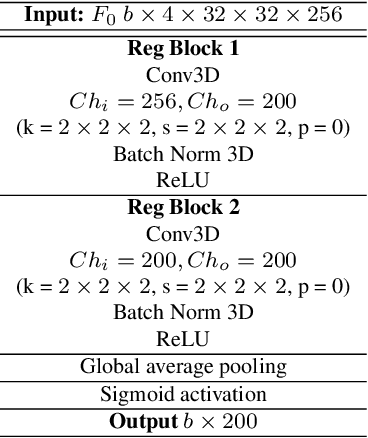

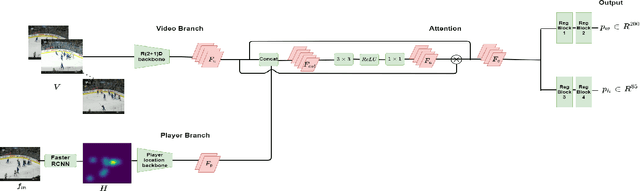

Abstract:Puck localization is an important problem in ice hockey video analytics useful for analyzing the game, determining play location, and assessing puck possession. The problem is challenging due to the small size of the puck, excessive motion blur due to high puck velocity and occlusions due to players and boards. In this paper, we introduce and implement a network for puck localization in broadcast hockey video. The network leverages expert NHL play-by-play annotations and uses temporal context to locate the puck. Player locations are incorporated into the network through an attention mechanism by encoding player positions with a Gaussian-based spatial heatmap drawn at player positions. Since event occurrence on the rink and puck location are related, we also perform event recognition by augmenting the puck localization network with an event recognition head and training the network through multi-task learning. Experimental results demonstrate that the network is able to localize the puck with an AUC of $73.1 \%$ on the test set. The puck location can be inferred in 720p broadcast videos at $5$ frames per second. It is also demonstrated that multi-task learning with puck location improves event recognition accuracy.
Localization of Ice-Rink for Broadcast Hockey Videos
Apr 22, 2021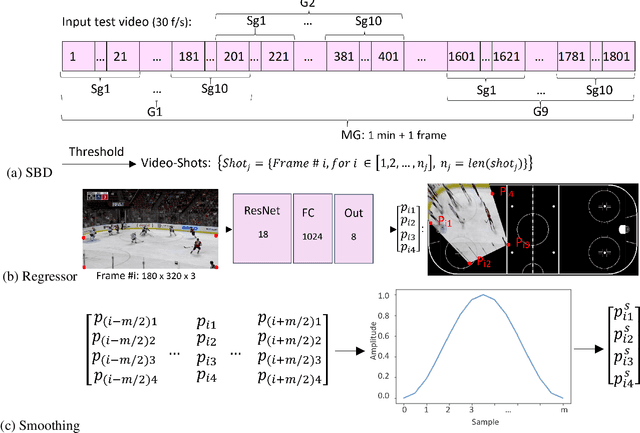
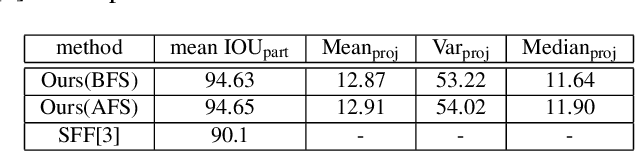
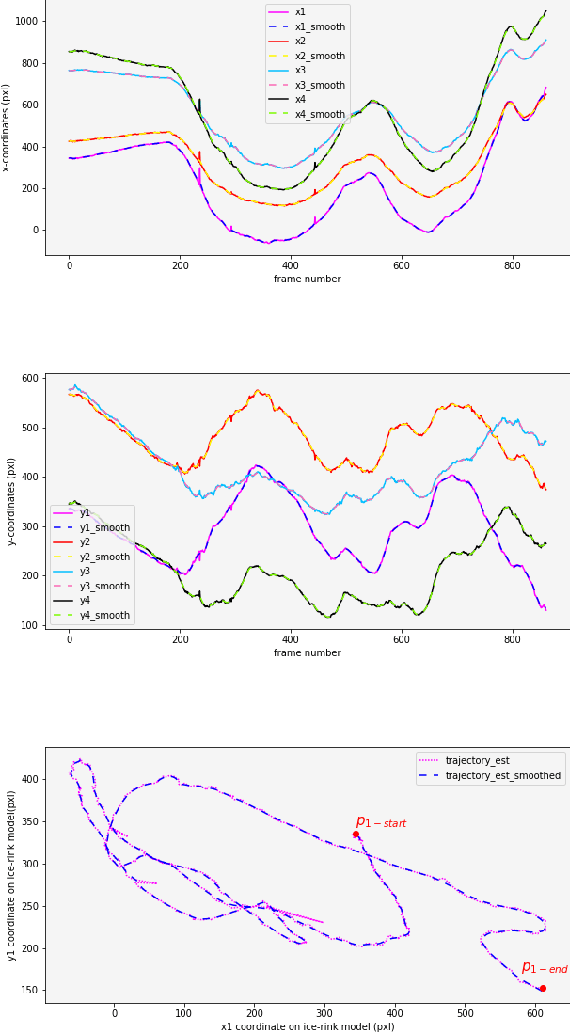
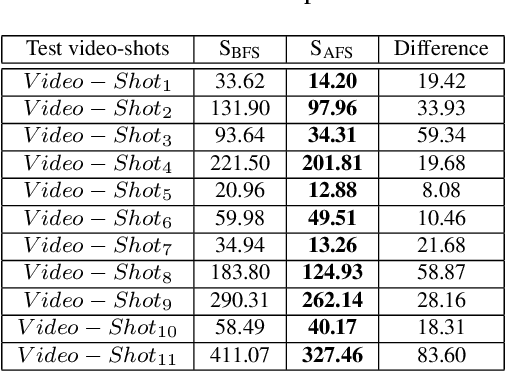
Abstract:In this work, an automatic and simple framework for hockey ice-rink localization from broadcast videos is introduced. First, video is broken into video-shots by a hierarchical partitioning of the video frames, and thresholding based on their histograms. To localize the frames on the ice-rink model, a ResNet18-based regressor is implemented and trained, which regresses to four control points on the model in a frame-by-frame fashion. This leads to the projection jittering problem in the video. To overcome this, in the inference phase, the trajectory of the control points on the ice-rink model are smoothed, for all the consecutive frames of a given video-shot, by convolving a Hann window with the achieved coordinates. Finally, the smoothed homography matrix is computed by using the direct linear transform on the four pairs of corresponding points. A hockey dataset for training and testing the regressor is gathered. The results show success of this simple and comprehensive procedure for localizing the hockey ice-rink and addressing the problem of jittering without affecting the accuracy of homography estimation.
Event detection in coarsely annotated sports videos via parallel multi receptive field 1D convolutions
Apr 13, 2020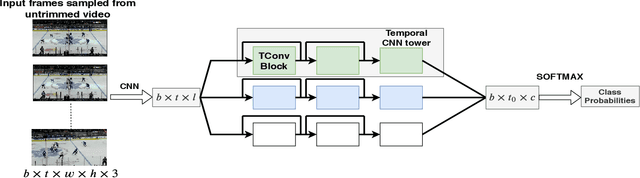

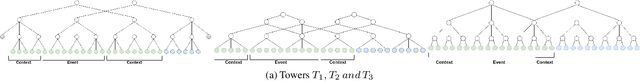
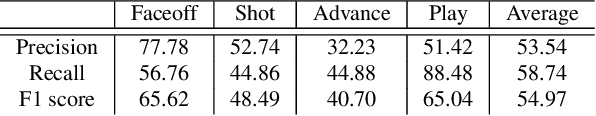
Abstract:In problems such as sports video analytics, it is difficult to obtain accurate frame level annotations and exact event duration because of the lengthy videos and sheer volume of video data. This issue is even more pronounced in fast-paced sports such as ice hockey. Obtaining annotations on a coarse scale can be much more practical and time efficient. We propose the task of event detection in coarsely annotated videos. We introduce a multi-tower temporal convolutional network architecture for the proposed task. The network, with the help of multiple receptive fields, processes information at various temporal scales to account for the uncertainty with regard to the exact event location and duration. We demonstrate the effectiveness of the multi-receptive field architecture through appropriate ablation studies. The method is evaluated on two tasks - event detection in coarsely annotated hockey videos in the NHL dataset and event spotting in soccer on the SoccerNet dataset. The two datasets lack frame-level annotations and have very distinct event frequencies. Experimental results demonstrate the effectiveness of the network by obtaining a 55% average F1 score on the NHL dataset and by achieving competitive performance compared to the state of the art on the SoccerNet dataset. We believe our approach will help develop more practical pipelines for event detection in sports video.
PuckNet: Estimating hockey puck location from broadcast video
Dec 11, 2019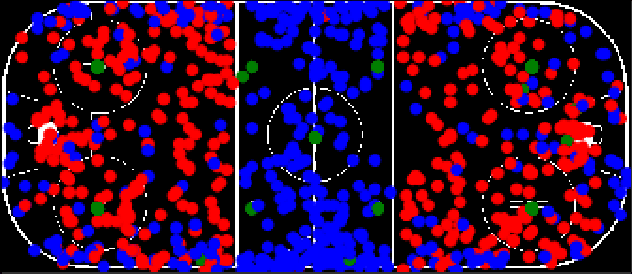
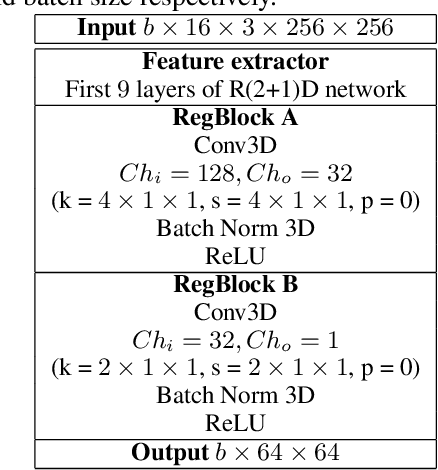


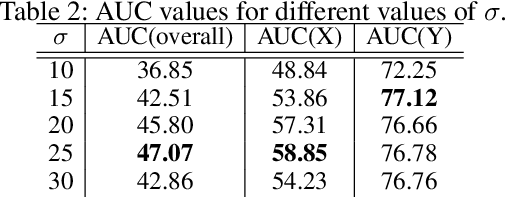
Abstract:Puck location in ice hockey is essential for hockey analysts for determining the location of play and analyzing game events. However, because of the difficulty involved in obtaining accurate annotations due to the extremely low visibility and commonly occurring occlusions of the puck, the problem is very challenging. The problem becomes even more challenging in broadcast videos with changing camera angles. We introduce a novel methodology for determining puck location from approximate puck location annotations in broadcast video. Our method uniquely leverages the existing puck location information that is publicly available in existing hockey event data and uses the corresponding one-second broadcast video clips as input to the network. The rationale behind using video as input instead of static images is that with video, the temporal information can be utilized to handle puck occlusions. The network outputs a heatmap representing the probability of the puck location using a 3D CNN based architecture. The network is able to regress the puck location from broadcast hockey video clips with varying camera angles. Experimental results demonstrate the capability of the method, achieving 47.07% AUC on the test dataset. The network is also able to estimate the puck location in defensive/offensive zones with an accuracy of greater than 80%.
Generative Adversarial Networks and Conditional Random Fields for Hyperspectral Image Classification
May 12, 2019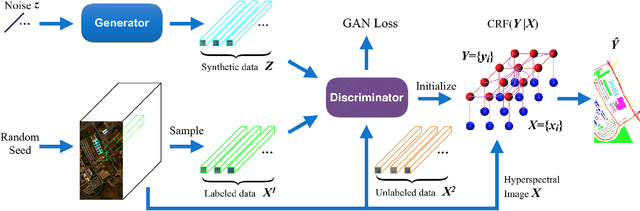
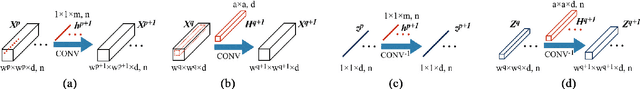
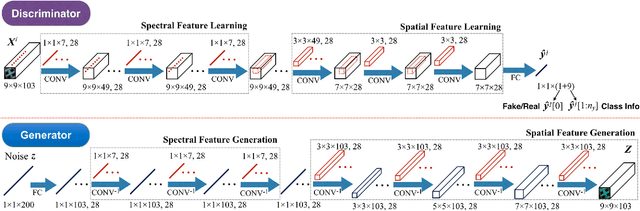
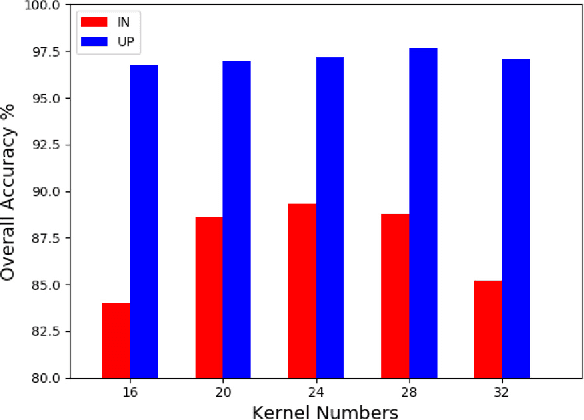
Abstract:In this paper, we address the hyperspectral image (HSI) classification task with a generative adversarial network and conditional random field (GAN-CRF) -based framework, which integrates a semi-supervised deep learning and a probabilistic graphical model, and make three contributions. First, we design four types of convolutional and transposed convolutional layers that consider the characteristics of HSIs to help with extracting discriminative features from limited numbers of labeled HSI samples. Second, we construct semi-supervised GANs to alleviate the shortage of training samples by adding labels to them and implicitly reconstructing real HSI data distribution through adversarial training. Third, we build dense conditional random fields (CRFs) on top of the random variables that are initialized to the softmax predictions of the trained GANs and are conditioned on HSIs to refine classification maps. This semi-supervised framework leverages the merits of discriminative and generative models through a game-theoretical approach. Moreover, even though we used very small numbers of labeled training HSI samples from the two most challenging and extensively studied datasets, the experimental results demonstrated that spectral-spatial GAN-CRF (SS-GAN-CRF) models achieved top-ranking accuracy for semi-supervised HSI classification.
KPTransfer: improved performance and faster convergence from keypoint subset-wise domain transfer in human pose estimation
Mar 24, 2019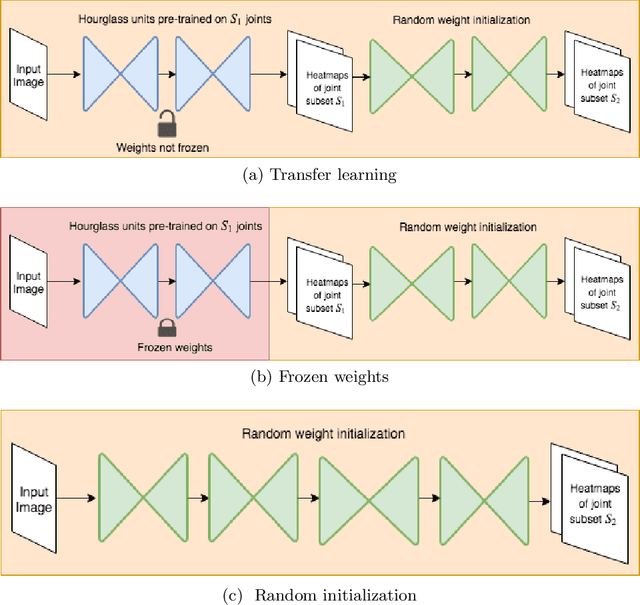

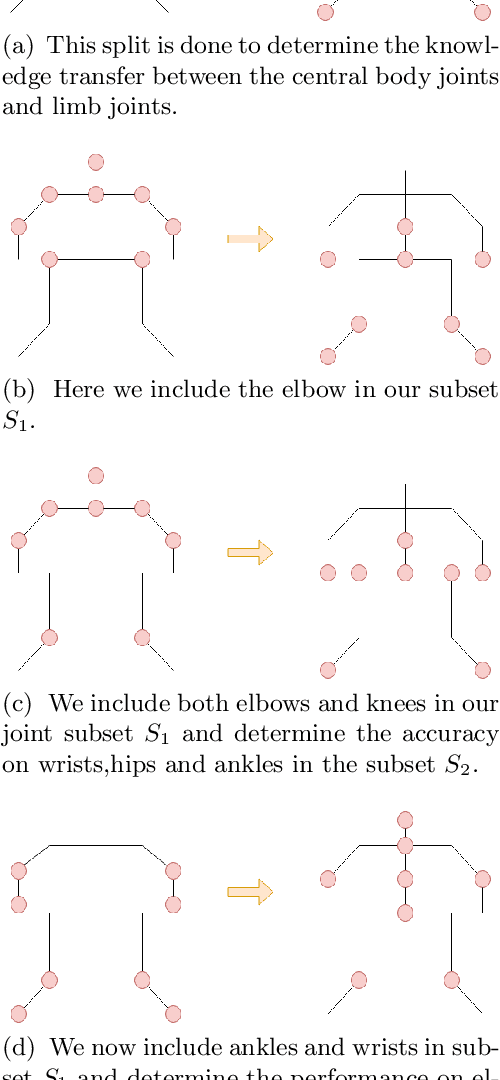
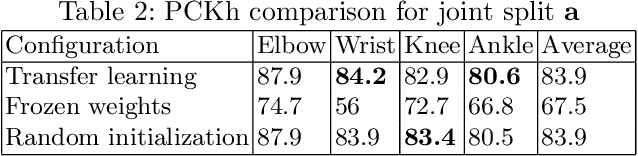
Abstract:In this paper, we present a novel approach called KPTransfer for improving modeling performance for keypoint detection deep neural networks via domain transfer between different keypoint subsets. This approach is motivated by the notion that rich contextual knowledge can be transferred between different keypoint subsets representing separate domains. In particular, the proposed method takes into account various keypoint subsets/domains by sequentially adding and removing keypoints. Contextual knowledge is transferred between two separate domains via domain transfer. Experiments to demonstrate the efficacy of the proposed KPTransfer approach were performed for the task of human pose estimation on the MPII dataset, with comparisons against random initialization and frozen weight extraction configurations. Experimental results demonstrate the efficacy of performing domain transfer between two different joint subsets resulting in a PCKh improvement of up to 1.1 over random initialization on joints such as wrists and knee in certain joint splits with an overall PCKh improvement of 0.5. Domain transfer from a different set of joints not only results in improved accuracy but also results in faster convergence because of mutual co-adaptations of weights resulting from the contextual knowledge of the pose from a different set of joints.
SISC: End-to-end Interpretable Discovery Radiomics-Driven Lung Cancer Prediction via Stacked Interpretable Sequencing Cells
Jan 15, 2019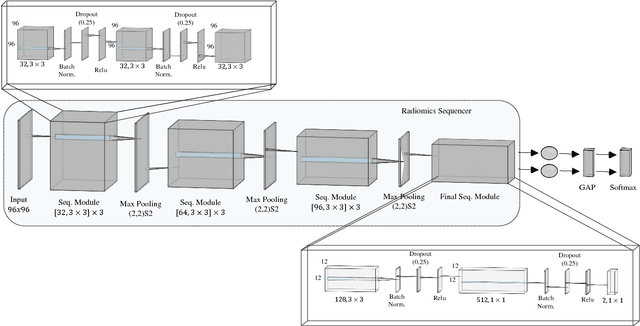
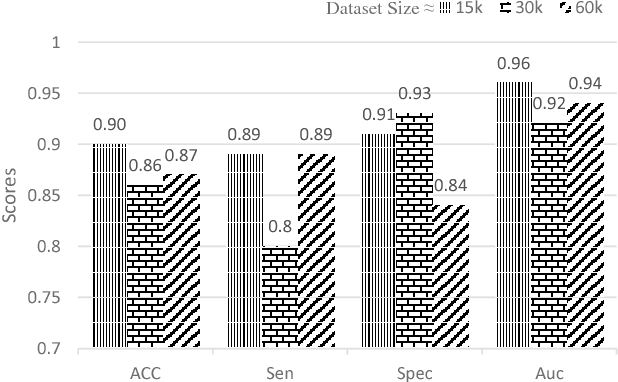
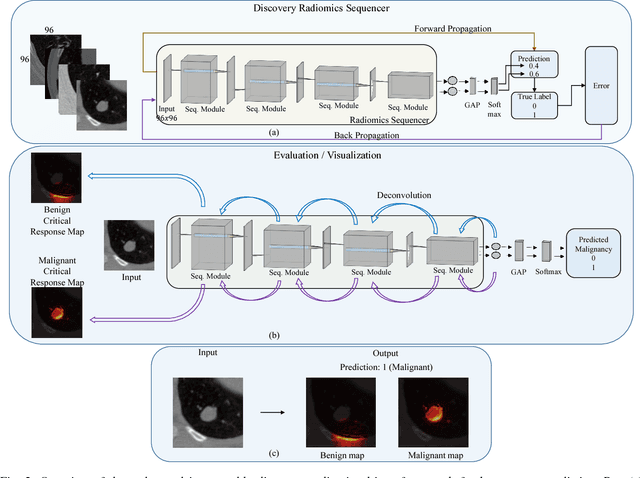
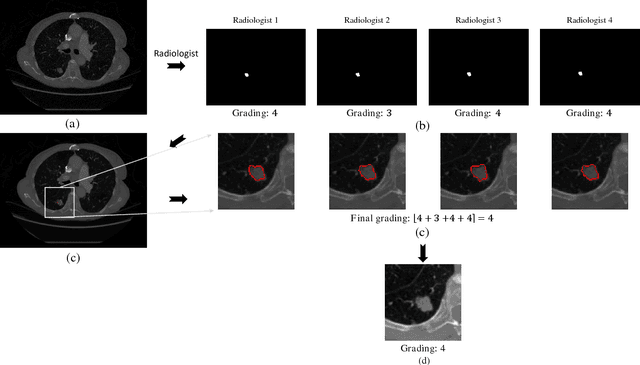
Abstract:Objective: Lung cancer is the leading cause of cancer-related death worldwide. Computer-aided diagnosis (CAD) systems have shown significant promise in recent years for facilitating the effective detection and classification of abnormal lung nodules in computed tomography (CT) scans. While hand-engineered radiomic features have been traditionally used for lung cancer prediction, there have been significant recent successes achieving state-of-the-art results in the area of discovery radiomics. Here, radiomic sequencers comprising of highly discriminative radiomic features are discovered directly from archival medical data. However, the interpretation of predictions made using such radiomic sequencers remains a challenge. Method: A novel end-to-end interpretable discovery radiomics-driven lung cancer prediction pipeline has been designed, build, and tested. The radiomic sequencer being discovered possesses a deep architecture comprised of stacked interpretable sequencing cells (SISC). Results: The SISC architecture is shown to outperform previous approaches while providing more insight in to its decision making process. Conclusion: The SISC radiomic sequencer is able to achieve state-of-the-art results in lung cancer prediction, and also offers prediction interpretability in the form of critical response maps. Significance: The critical response maps are useful for not only validating the predictions of the proposed SISC radiomic sequencer, but also provide improved radiologist-machine collaboration for effective diagnosis.
Numerical Demultiplexing of Color Image Sensor Measurements via Non-linear Random Forest Modeling
Dec 17, 2015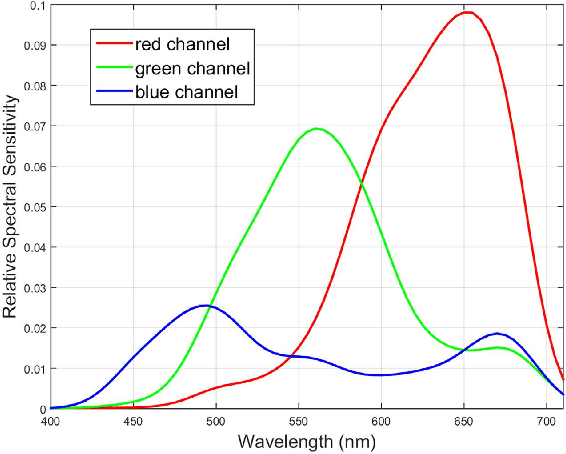
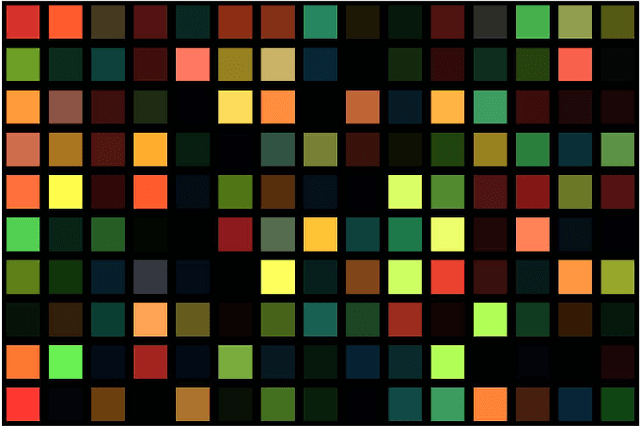
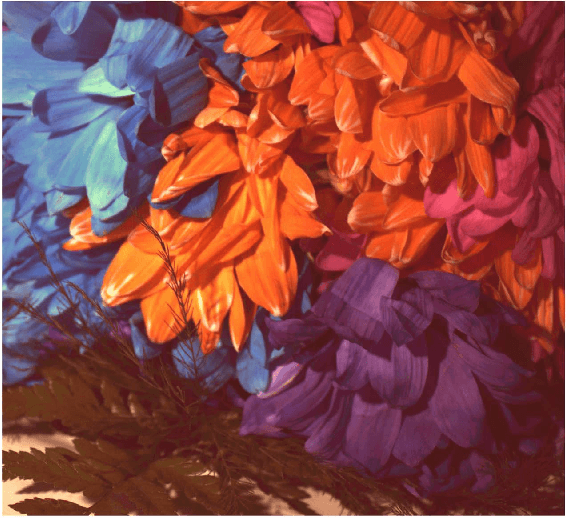
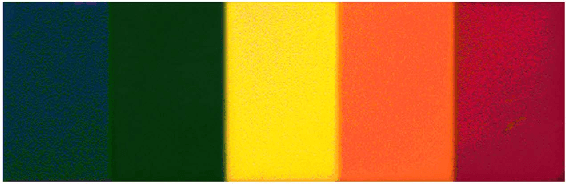



Abstract:The simultaneous capture of imaging data at multiple wavelengths across the electromagnetic spectrum is highly challenging, requiring complex and costly multispectral image sensors. In this study, we introduce a comprehensive framework for performing simultaneous multispectral imaging using conventional image sensors with color filter arrays via numerical demultiplexing of the color image sensor measurements. A numerical forward model characterizing the formation of sensor measurements from light spectra hitting the sensor is constructed based on a comprehensive spectral characterization of the sensor. A numerical demultiplexer is then learned via non-linear random forest modeling based on the forward model. Given the learned numerical demultiplexer, one can then demultiplex simultaneously-acquired measurements made by the image sensor into reflectance intensities at discrete selectable wavelengths, resulting in a higher resolution reflectance spectrum. Simulation and real-world experimental results demonstrate the efficacy of such a method for simultaneous multispectral imaging.
Non-contact transmittance photoplethysmographic imaging (PPGI) for long-distance cardiovascular monitoring
Mar 23, 2015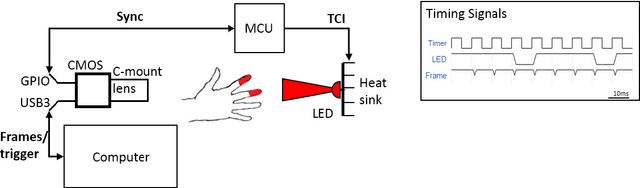
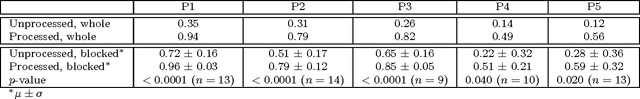

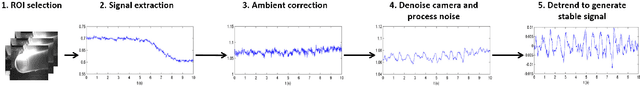
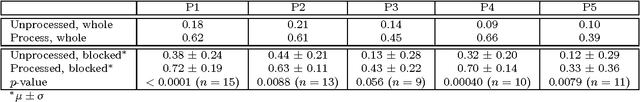
Abstract:Photoplethysmography (PPG) devices are widely used for monitoring cardiovascular function. However, these devices require skin contact, which restrict their use to at-rest short-term monitoring using single-point measurements. Photoplethysmographic imaging (PPGI) has been recently proposed as a non-contact monitoring alternative by measuring blood pulse signals across a spatial region of interest. Existing systems operate in reflectance mode, of which many are limited to short-distance monitoring and are prone to temporal changes in ambient illumination. This paper is the first study to investigate the feasibility of long-distance non-contact cardiovascular monitoring at the supermeter level using transmittance PPGI. For this purpose, a novel PPGI system was designed at the hardware and software level using ambient correction via temporally coded illumination (TCI) and signal processing for PPGI signal extraction. Experimental results show that the processing steps yield a substantially more pulsatile PPGI signal than the raw acquired signal, resulting in statistically significant increases in correlation to ground-truth PPG in both short- ($p \in [<0.0001, 0.040]$) and long-distance ($p \in [<0.0001, 0.056]$) monitoring. The results support the hypothesis that long-distance heart rate monitoring is feasible using transmittance PPGI, allowing for new possibilities of monitoring cardiovascular function in a non-contact manner.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge