Daniel L. Rubin
Plexus Convolutional Neural Network (PlexusNet): A novel neural network architecture for histologic image analysis
Aug 24, 2019
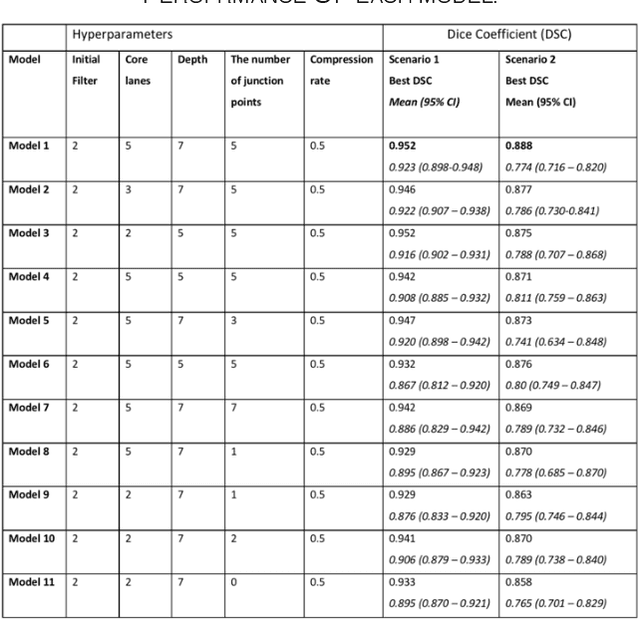
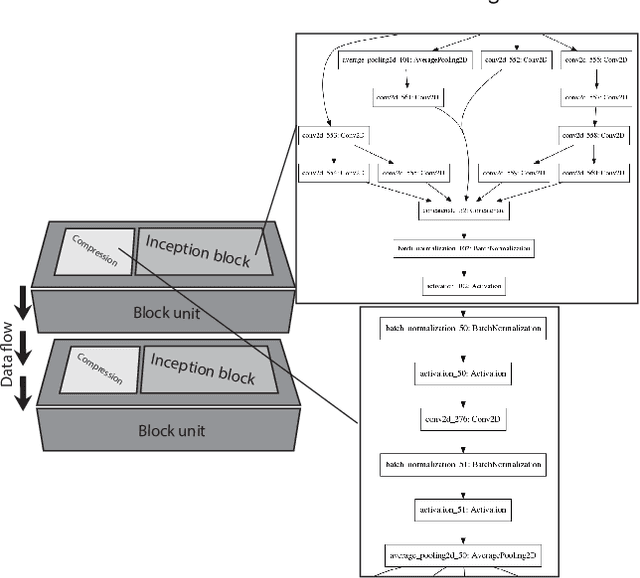
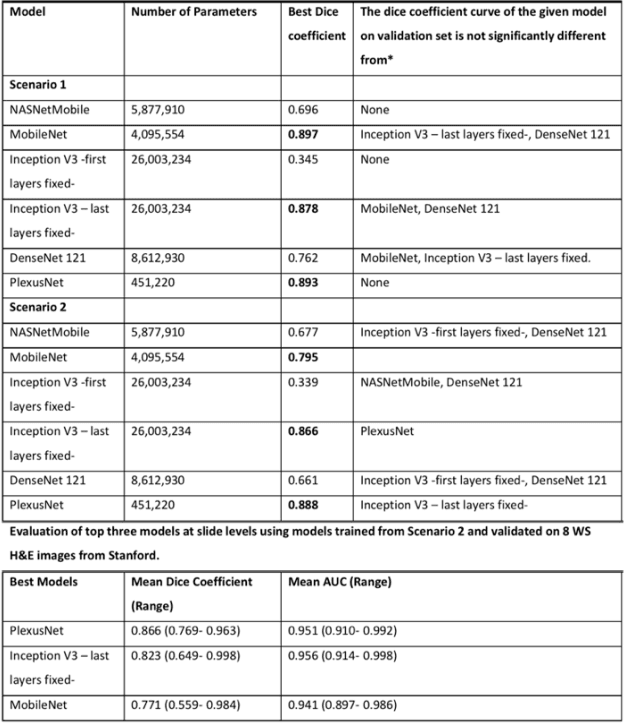
Abstract:Different convolutional neural network (CNN) models have been tested for their application in histologic imaging analyses. However, these models are prone to overfitting due to their large parameter capacity, requiring more data and expensive computational resources for model training. Given these limitations, we developed and tested PlexusNet for histologic evaluation using a single GPU by a batch dimension of 16x512x512x3. We utilized 62 Hematoxylin and eosin stain (H&E) annotated histological images of radical prostatectomy cases from TCGA-PRAD and Stanford University, and 24 H&E whole-slide images with hepatocellular carcinoma from TCGA-LIHC diagnostic histology images. Base models were DenseNet, Inception V3, and MobileNet and compared with PlexusNet. The dice coefficient (DSC) was evaluated for each model. PlexusNet delivered comparable classification performance (DSC at patch level: 0.89) for H&E whole-slice images in distinguishing prostate cancer from normal tissues. The parameter capacity of PlexusNet is 9 times smaller than MobileNet or 58 times smaller than Inception V3, respectively. Similar findings were observed in distinguishing hepatocellular carcinoma from non-cancerous liver histologies (DSC at patch level: 0.85). As conclusion, PlexusNet represents a novel model architecture for histological image analysis that achieves classification performance comparable to the base models while providing orders-of-magnitude memory savings.
Self-Attention Capsule Networks for Image Classification
Apr 29, 2019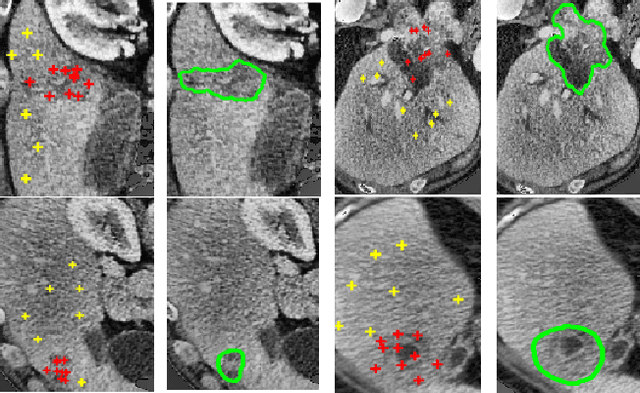
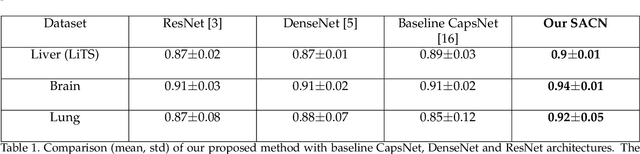
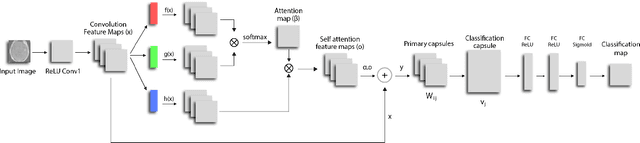
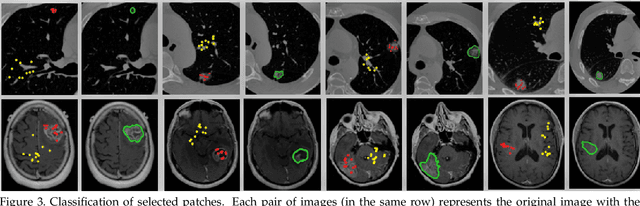
Abstract:We propose a novel architecture for image classification, called Self-Attention Capsule Networks (SACN). SACN is the first model that incorporates the Self-Attention mechanism as an integral layer within the Capsule Network (CapsNet). While the Self-Attention mechanism selects the more dominant image regions to focus on, the CapsNet analyzes the relevant features and their spatial correlations inside these regions only. The features are extracted in the convolutional layer. Then, the Self-Attention layer learns to suppress irrelevant regions based on features analysis, and highlights salient features useful for a specific task. The attention map is then fed into the CapsNet primary layer that is followed by a classification layer. The SACN proposed model was designed to use a relatively shallow CapsNet architecture to reduce computational load, and compensates for the absence of a deeper network by using the Self-Attention module to significantly improve the results. The proposed Self-Attention CapsNet architecture was extensively evaluated on five different datasets, mainly on three different medical sets, in addition to the natural MNIST and SVHN. The model was able to classify images and their patches with diverse and complex backgrounds better than the baseline CapsNet. As a result, the proposed Self-Attention CapsNet significantly improved classification performance within and across different datasets and outperformed the baseline CapsNet not only in classification accuracy but also in robustness.
Deep Learning Enables Automatic Detection and Segmentation of Brain Metastases on Multi-Sequence MRI
Mar 18, 2019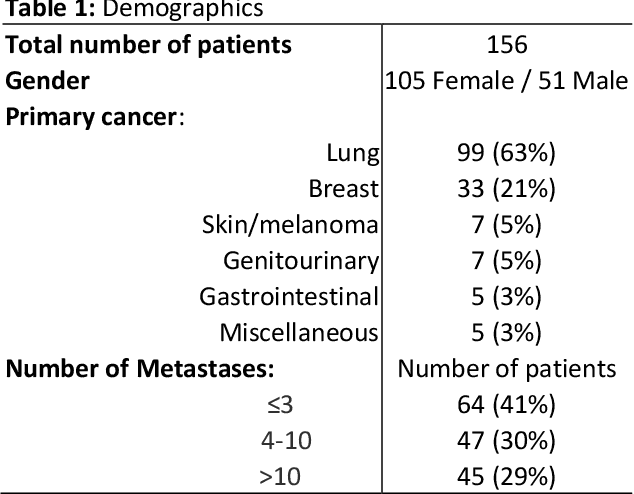
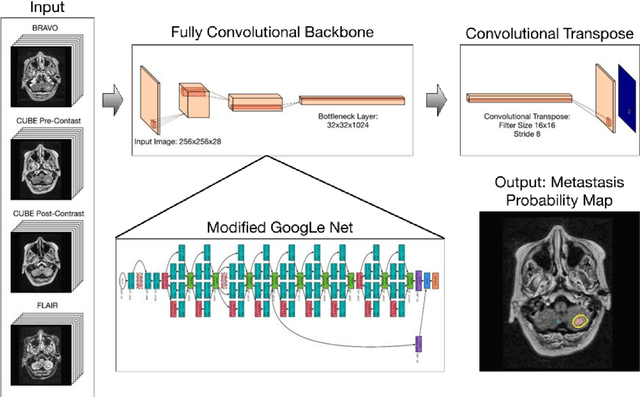
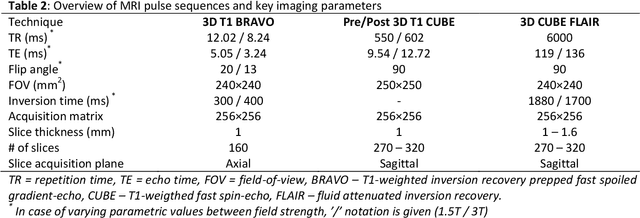
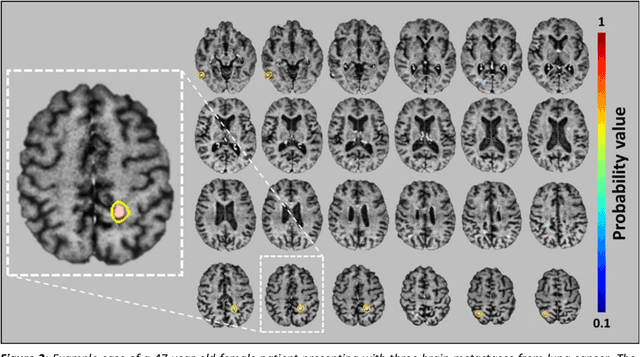
Abstract:Detecting and segmenting brain metastases is a tedious and time-consuming task for many radiologists, particularly with the growing use of multi-sequence 3D imaging. This study demonstrates automated detection and segmentation of brain metastases on multi-sequence MRI using a deep learning approach based on a fully convolution neural network (CNN). In this retrospective study, a total of 156 patients with brain metastases from several primary cancers were included. Pre-therapy MR images (1.5T and 3T) included pre- and post-gadolinium T1-weighted 3D fast spin echo, post-gadolinium T1-weighted 3D axial IR-prepped FSPGR, and 3D fluid attenuated inversion recovery. The ground truth was established by manual delineation by two experienced neuroradiologists. CNN training/development was performed using 100 and 5 patients, respectively, with a 2.5D network based on a GoogLeNet architecture. The results were evaluated in 51 patients, equally separated into those with few (1-3), multiple (4-10), and many (>10) lesions. Network performance was evaluated using precision, recall, Dice/F1 score, and ROC-curve statistics. For an optimal probability threshold, detection and segmentation performance was assessed on a per metastasis basis. The area under the ROC-curve (AUC), averaged across all patients, was 0.98. The AUC in the subgroups was 0.99, 0.97, and 0.97 for patients having 1-3, 4-10, and >10 metastases, respectively. Using an average optimal probability threshold determined by the development set, precision, recall, and Dice-score were 0.79, 0.53, and 0.79, respectively. At the same probability threshold, the network showed an average false positive rate of 8.3/patient (no lesion-size limit) and 3.4/patient (10 mm3 lesion size limit). In conclusion, a deep learning approach using multi-sequence MRI can aid in the detection and segmentation of brain metastases.
CT organ segmentation using GPU data augmentation, unsupervised labels and IOU loss
Nov 27, 2018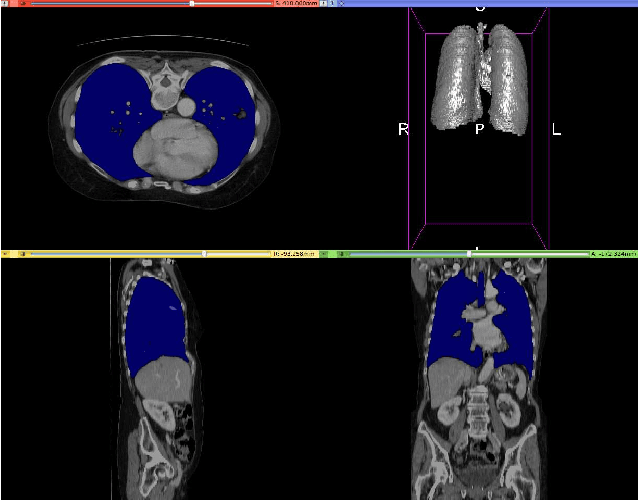
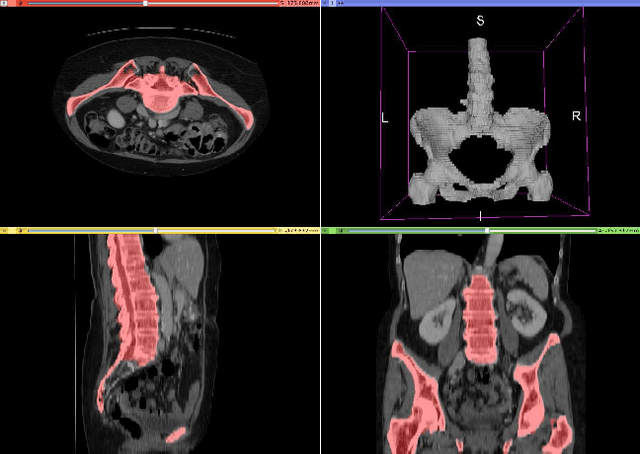
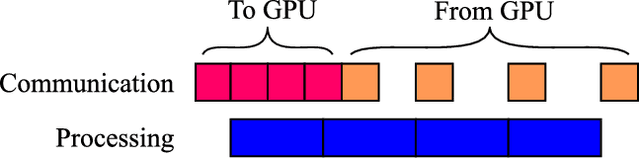
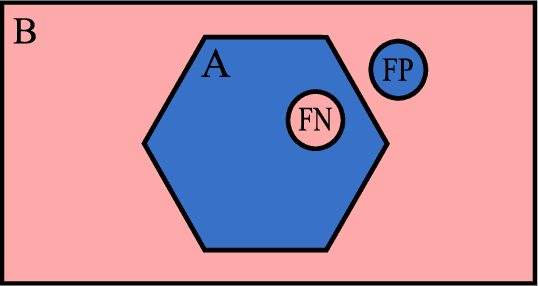
Abstract:Fully-convolutional neural networks have achieved superior performance in a variety of image segmentation tasks. However, their training requires laborious manual annotation of large datasets, as well as acceleration by parallel processors with high-bandwidth memory, such as GPUs. We show that simple models can achieve competitive accuracy for organ segmentation on CT images when trained with extensive data augmentation, which leverages existing graphics hardware to quickly apply geometric and photometric transformations to 3D image data. On 3 mm^3 CT volumes, our GPU implementation is 2.6-8X faster than a widely-used CPU version, including communication overhead. We also show how to automatically generate training labels using rudimentary morphological operations, which are efficiently computed by 3D Fourier transforms. We combined fully-automatic labels for the lungs and bone with semi-automatic ones for the liver, kidneys and bladder, to create a dataset of 130 labeled CT scans. To achieve the best results from data augmentation, our model uses the intersection-over-union (IOU) loss function, a close relative of the Dice loss. We discuss its mathematical properties and explain why it outperforms the usual weighted cross-entropy loss for unbalanced segmentation tasks. We conclude that there is no unique IOU loss function, as the naive one belongs to a broad family of functions with the same essential properties. When combining data augmentation with the IOU loss, our model achieves a Dice score of 78-92% for each organ. The trained model, code and dataset will be made publicly available, to further medical imaging research.
Abstract: Probabilistic Prognostic Estimates of Survival in Metastatic Cancer Patients
Jul 13, 2018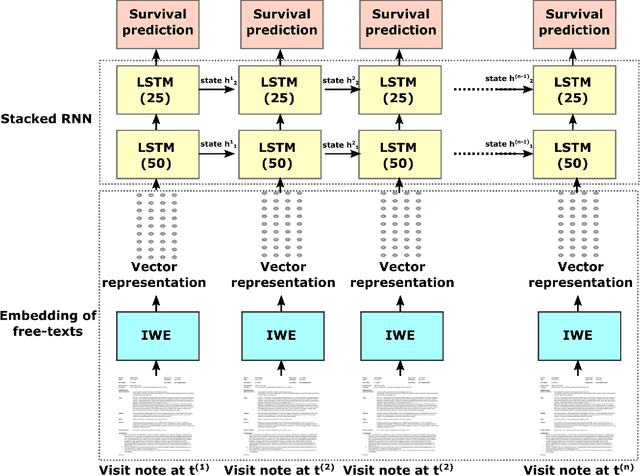
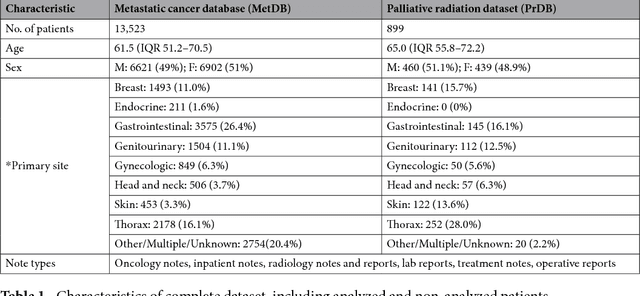
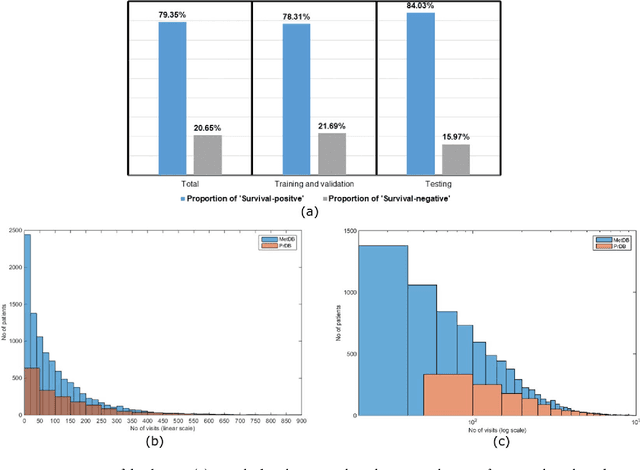
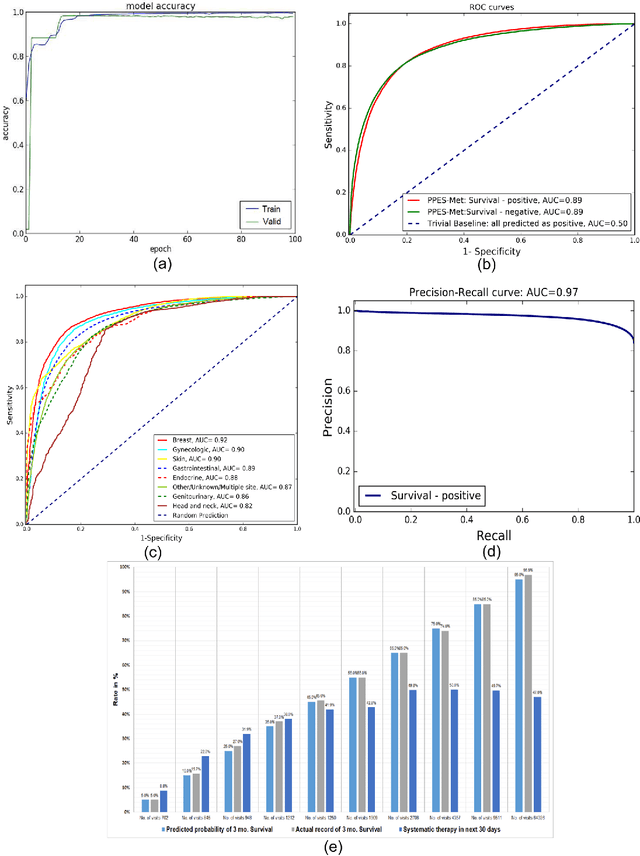
Abstract:We propose a deep learning model - Probabilistic Prognostic Estimates of Survival in Metastatic Cancer Patients (PPES-Met) for estimating short-term life expectancy (3 months) of the patients by analyzing free-text clinical notes in the electronic medical record, while maintaining the temporal visit sequence. In a single framework, we integrated semantic data mapping and neural embedding technique to produce a text processing method that extracts relevant information from heterogeneous types of clinical notes in an unsupervised manner, and we designed a recurrent neural network to model the temporal dependency of the patient visits. The model was trained on a large dataset (10,293 patients) and validated on a separated dataset (1818 patients). Our method achieved an area under the ROC curve (AUC) of 0.89. To provide explain-ability, we developed an interactive graphical tool that may improve physician understanding of the basis for the model's predictions. The high accuracy and explain-ability of the PPES-Met model may enable our model to be used as a decision support tool to personalize metastatic cancer treatment and provide valuable assistance to the physicians.
A Scalable Machine Learning Approach for Inferring Probabilistic US-LI-RADS Categorization
Jun 15, 2018
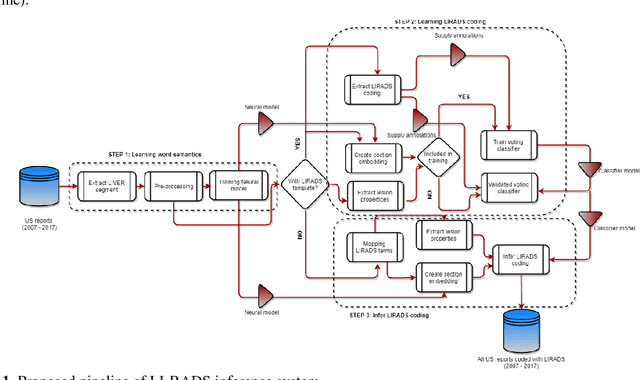
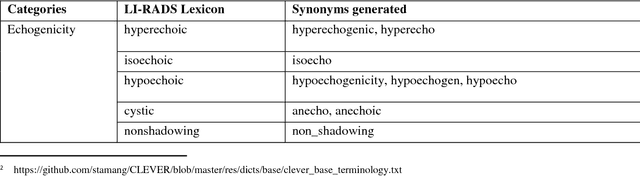
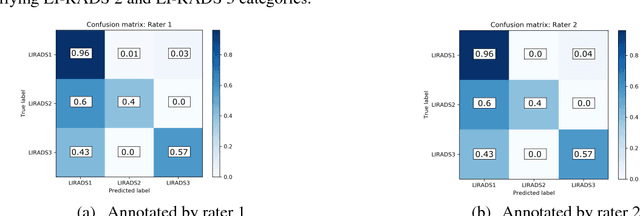
Abstract:We propose a scalable computerized approach for large-scale inference of Liver Imaging Reporting and Data System (LI-RADS) final assessment categories in narrative ultrasound (US) reports. Although our model was trained on reports created using a LI-RADS template, it was also able to infer LI-RADS scoring for unstructured reports that were created before the LI-RADS guidelines were established. No human-labelled data was required in any step of this study; for training, LI-RADS scores were automatically extracted from those reports that contained structured LI-RADS scores, and it translated the derived knowledge to reasoning on unstructured radiology reports. By providing automated LI-RADS categorization, our approach may enable standardizing screening recommendations and treatment planning of patients at risk for hepatocellular carcinoma, and it may facilitate AI-based healthcare research with US images by offering large scale text mining and data gathering opportunities from standard hospital clinical data repositories.
Intelligent Word Embeddings of Free-Text Radiology Reports
Nov 19, 2017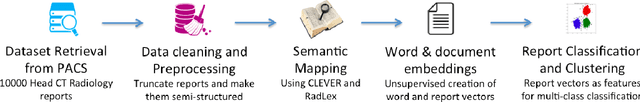
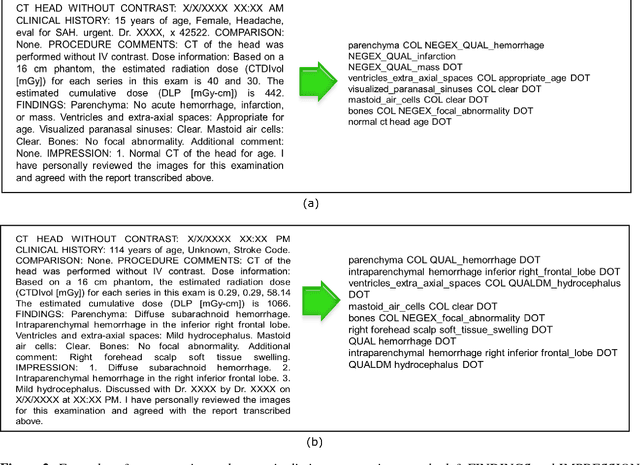



Abstract:Radiology reports are a rich resource for advancing deep learning applications in medicine by leveraging the large volume of data continuously being updated, integrated, and shared. However, there are significant challenges as well, largely due to the ambiguity and subtlety of natural language. We propose a hybrid strategy that combines semantic-dictionary mapping and word2vec modeling for creating dense vector embeddings of free-text radiology reports. Our method leverages the benefits of both semantic-dictionary mapping as well as unsupervised learning. Using the vector representation, we automatically classify the radiology reports into three classes denoting confidence in the diagnosis of intracranial hemorrhage by the interpreting radiologist. We performed experiments with varying hyperparameter settings of the word embeddings and a range of different classifiers. Best performance achieved was a weighted precision of 88% and weighted recall of 90%. Our work offers the potential to leverage unstructured electronic health record data by allowing direct analysis of narrative clinical notes.
Inferring Generative Model Structure with Static Analysis
Sep 07, 2017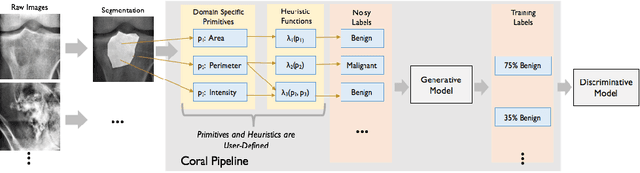

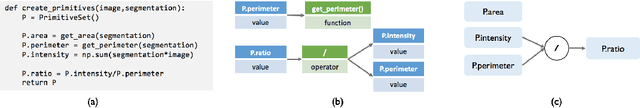
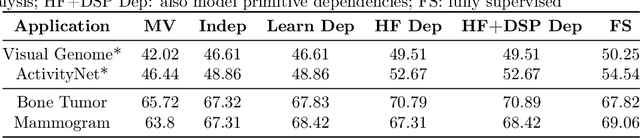
Abstract:Obtaining enough labeled data to robustly train complex discriminative models is a major bottleneck in the machine learning pipeline. A popular solution is combining multiple sources of weak supervision using generative models. The structure of these models affects training label quality, but is difficult to learn without any ground truth labels. We instead rely on these weak supervision sources having some structure by virtue of being encoded programmatically. We present Coral, a paradigm that infers generative model structure by statically analyzing the code for these heuristics, thus reducing the data required to learn structure significantly. We prove that Coral's sample complexity scales quasilinearly with the number of heuristics and number of relations found, improving over the standard sample complexity, which is exponential in $n$ for identifying $n^{\textrm{th}}$ degree relations. Experimentally, Coral matches or outperforms traditional structure learning approaches by up to 3.81 F1 points. Using Coral to model dependencies instead of assuming independence results in better performance than a fully supervised model by 3.07 accuracy points when heuristics are used to label radiology data without ground truth labels.
A Fully-Automated Pipeline for Detection and Segmentation of Liver Lesions and Pathological Lymph Nodes
Mar 19, 2017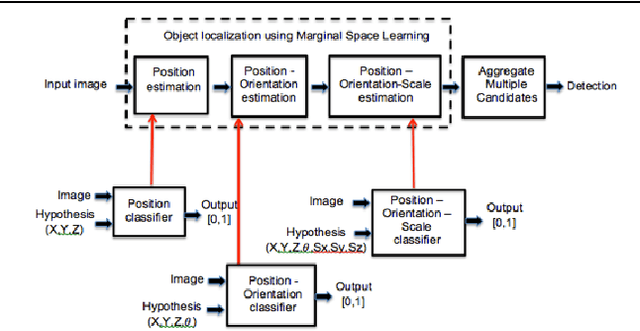

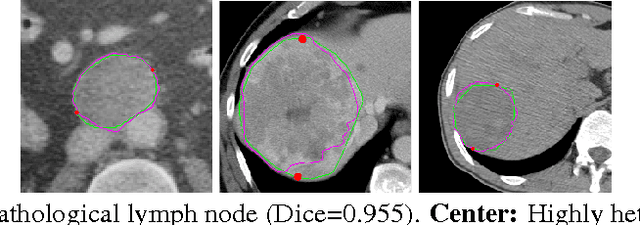
Abstract:We propose a fully-automated method for accurate and robust detection and segmentation of potentially cancerous lesions found in the liver and in lymph nodes. The process is performed in three steps, including organ detection, lesion detection and lesion segmentation. Our method applies machine learning techniques such as marginal space learning and convolutional neural networks, as well as active contour models. The method proves to be robust in its handling of extremely high lesion diversity. We tested our method on volumetric computed tomography (CT) images, including 42 volumes containing liver lesions and 86 volumes containing 595 pathological lymph nodes. Preliminary results under 10-fold cross validation show that for both the liver lesions and the lymph nodes, a total detection sensitivity of 0.53 and average Dice score of $0.71 \pm 0.15$ for segmentation were obtained.
Computerized Multiparametric MR image Analysis for Prostate Cancer Aggressiveness-Assessment
Dec 01, 2016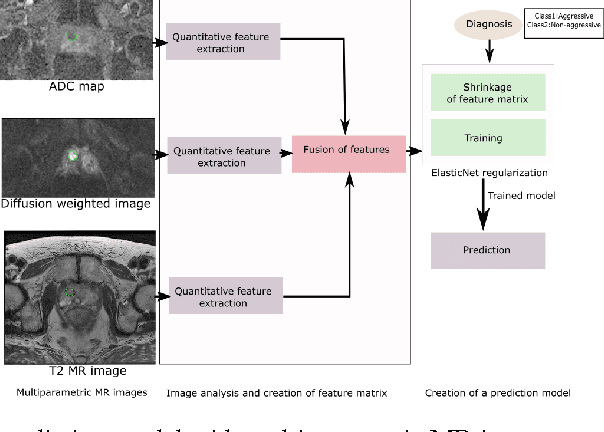
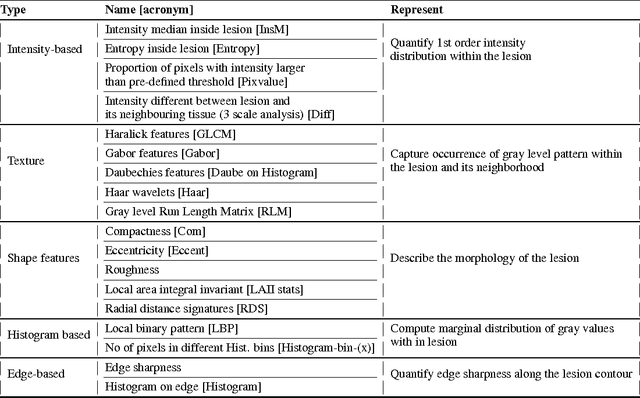
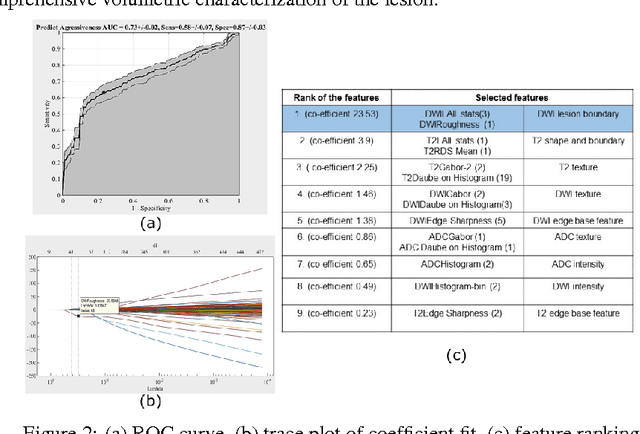
Abstract:We propose an automated method for detecting aggressive prostate cancer(CaP) (Gleason score >=7) based on a comprehensive analysis of the lesion and the surrounding normal prostate tissue which has been simultaneously captured in T2-weighted MR images, diffusion-weighted images (DWI) and apparent diffusion coefficient maps (ADC). The proposed methodology was tested on a dataset of 79 patients (40 aggressive, 39 non-aggressive). We evaluated the performance of a wide range of popular quantitative imaging features on the characterization of aggressive versus non-aggressive CaP. We found that a group of 44 discriminative predictors among 1464 quantitative imaging features can be used to produce an area under the ROC curve of 0.73.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge