Brandon Carter
Lost in Pruning: The Effects of Pruning Neural Networks beyond Test Accuracy
Mar 04, 2021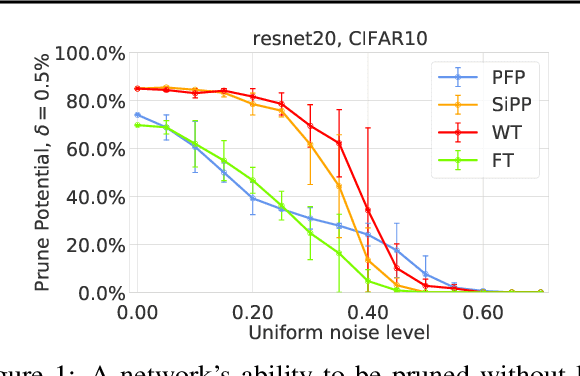

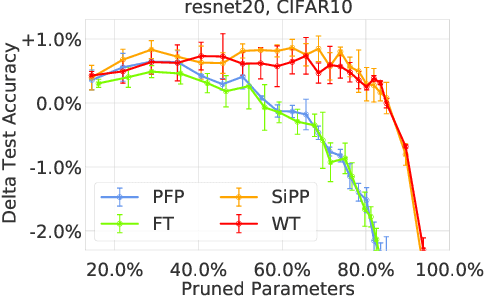
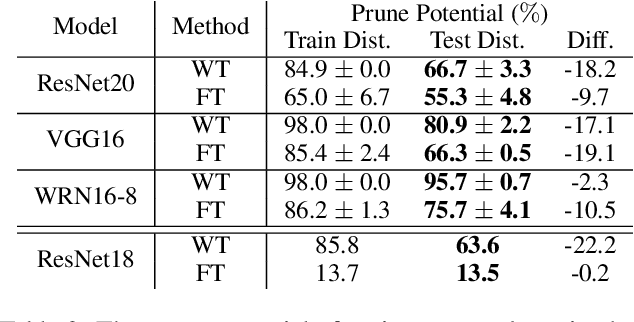
Abstract:Neural network pruning is a popular technique used to reduce the inference costs of modern, potentially overparameterized, networks. Starting from a pre-trained network, the process is as follows: remove redundant parameters, retrain, and repeat while maintaining the same test accuracy. The result is a model that is a fraction of the size of the original with comparable predictive performance (test accuracy). Here, we reassess and evaluate whether the use of test accuracy alone in the terminating condition is sufficient to ensure that the resulting model performs well across a wide spectrum of "harder" metrics such as generalization to out-of-distribution data and resilience to noise. Across evaluations on varying architectures and data sets, we find that pruned networks effectively approximate the unpruned model, however, the prune ratio at which pruned networks achieve commensurate performance varies significantly across tasks. These results call into question the extent of \emph{genuine} overparameterization in deep learning and raise concerns about the practicability of deploying pruned networks, specifically in the context of safety-critical systems, unless they are widely evaluated beyond test accuracy to reliably predict their performance. Our code is available at https://github.com/lucaslie/torchprune.
Maximum n-times Coverage for COVID-19 Vaccine Design
Jan 24, 2021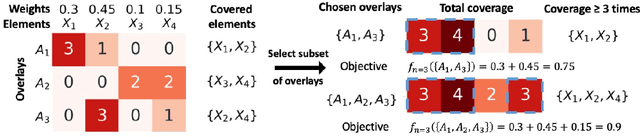
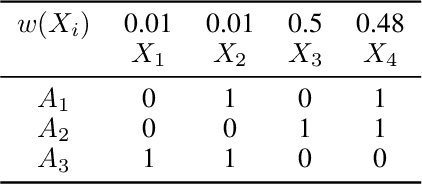

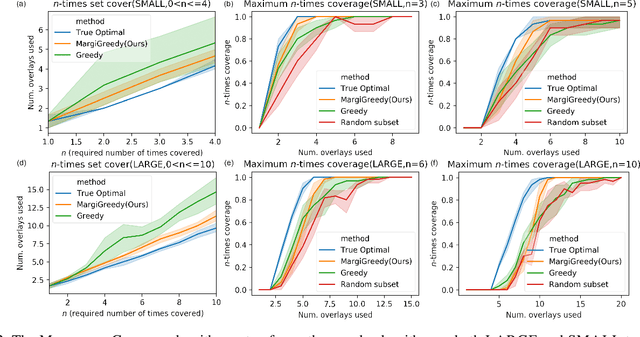
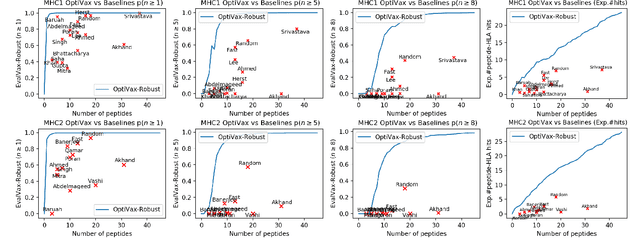
Abstract:In the maximum $n$-times coverage problem, we are provided a set of elements, a weight for each element, and a set of overlays where each overlay specifies an element specific coverage of zero or more times. The goal is to select up to $k$ overlays such that the sum of the weights of elements that are covered at least $n$ times is maximized. We also define the min-cost $n$-times coverage problem where the objective is to select the minimum set of overlays such that the sum of the weights of elements that are covered at least $n$ times is at least $\tau$. We show that the $n$-times coverage objective is not submodular, and we present an efficient solution by sequential greedy optimization. We frame the design of a peptide vaccine for COVID-19 as maximum $n$-times coverage using machine learning defined candidate peptide sets, and show that our solution is superior to 29 other published COVID-19 peptide vaccine designs in predicted population coverage and the expected number of peptides displayed by each individual's HLA molecules.
Overinterpretation reveals image classification model pathologies
Mar 19, 2020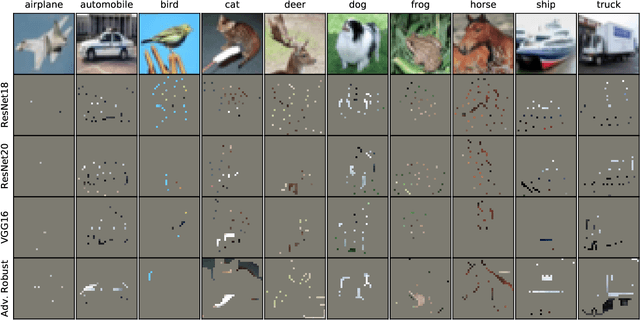
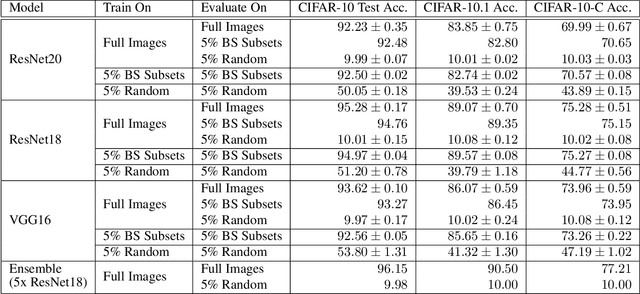
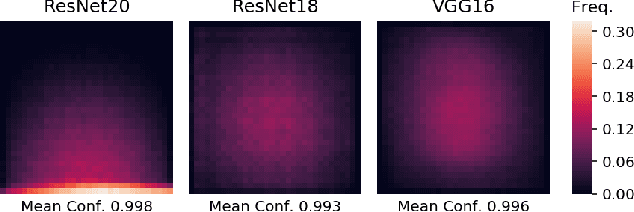
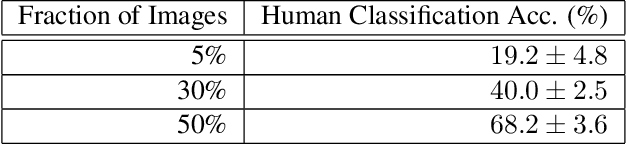
Abstract:Image classifiers are typically scored on their test set accuracy, but high accuracy can mask a subtle type of model failure. We find that high scoring convolutional neural networks (CNN) exhibit troubling pathologies that allow them to display high accuracy even in the absence of semantically salient features. When a model provides a high-confidence decision without salient supporting input features we say that the classifier has overinterpreted its input, finding too much class-evidence in patterns that appear nonsensical to humans. Here, we demonstrate that state of the art neural networks for CIFAR-10 and ImageNet suffer from overinterpretation, and find CIFAR-10 trained models make confident predictions even when 95% of an input image has been masked and humans are unable to discern salient features in the remaining pixel subset. Although these patterns portend potential model fragility in real-world deployment, they are in fact valid statistical patterns of the image classification benchmark that alone suffice to attain high test accuracy. We find that ensembling strategies can help mitigate model overinterpretation, and classifiers which rely on more semantically meaningful features can improve accuracy over both the test set and out-of-distribution images from a different source than the training data.
Embedding Comparator: Visualizing Differences in Global Structure and Local Neighborhoods via Small Multiples
Dec 10, 2019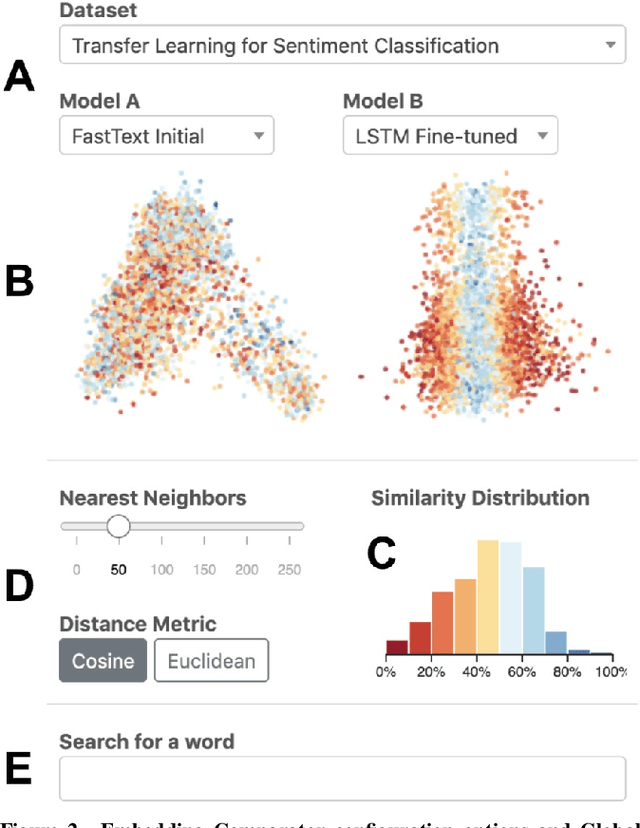
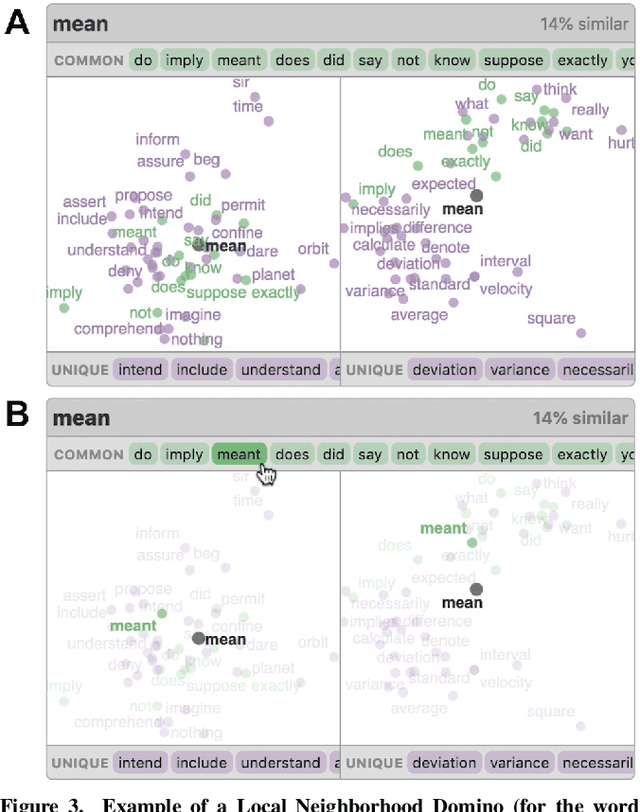
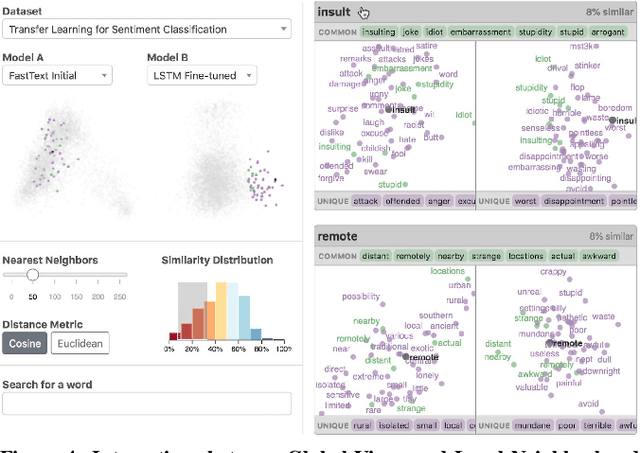
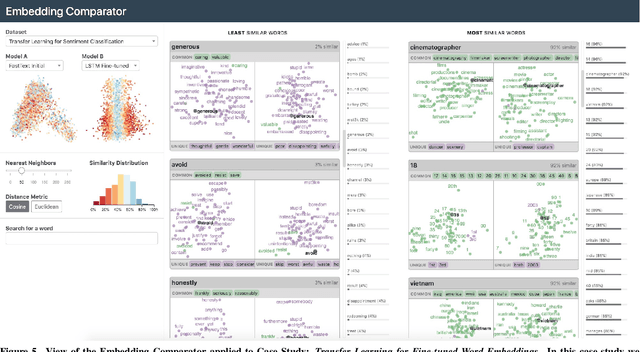
Abstract:Embeddings -- mappings from high-dimensional discrete input to lower-dimensional continuous vector spaces -- have been widely adopted in machine learning, linguistics, and computational biology as they often surface interesting and unexpected domain semantics. Through semi-structured interviews with embedding model researchers and practitioners, we find that current tools poorly support a central concern: comparing different embeddings when developing fairer, more robust models. In response, we present the Embedding Comparator, an interactive system that balances gaining an overview of the embedding spaces with making fine-grained comparisons of local neighborhoods. For a pair of models, we compute the similarity of the k-nearest neighbors of every embedded object, and visualize the results as Local Neighborhood Dominoes: small multiples that facilitate rapid comparisons. Using case studies, we illustrate the types of insights the Embedding Comparator reveals including how fine-tuning embeddings changes semantics, how language changes over time, and how training data differences affect two seemingly similar models.
What made you do this? Understanding black-box decisions with sufficient input subsets
Oct 09, 2018
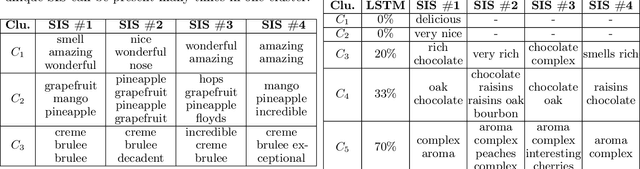

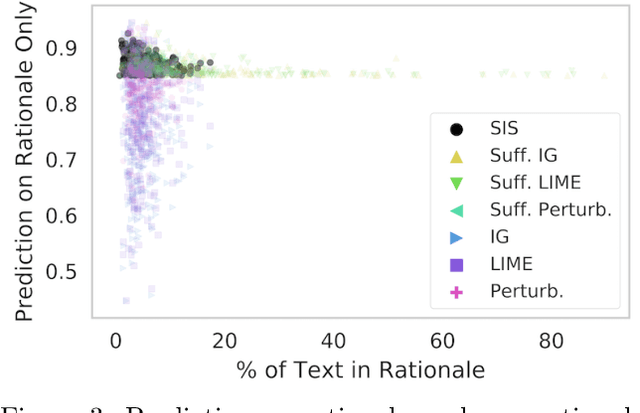
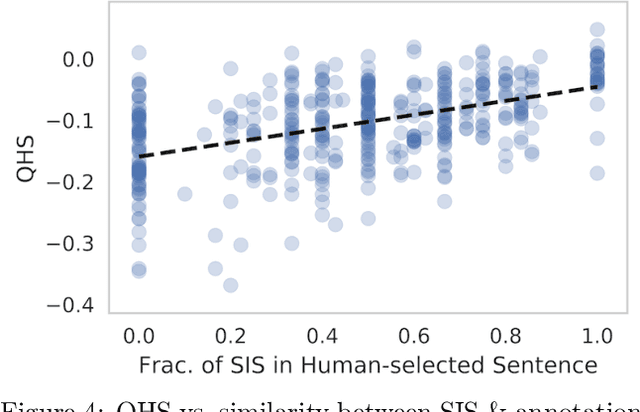
Abstract:Local explanation frameworks aim to rationalize particular decisions made by a black-box prediction model. Existing techniques are often restricted to a specific type of predictor or based on input saliency, which may be undesirably sensitive to factors unrelated to the model's decision making process. We instead propose sufficient input subsets that identify minimal subsets of features whose observed values alone suffice for the same decision to be reached, even if all other input feature values are missing. General principles that globally govern a model's decision-making can also be revealed by searching for clusters of such input patterns across many data points. Our approach is conceptually straightforward, entirely model-agnostic, simply implemented using instance-wise backward selection, and able to produce more concise rationales than existing techniques. We demonstrate the utility of our interpretation method on various neural network models trained on text, image, and genomic data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge