Ashwin Srinivasan
One-way Explainability Isn't The Message
May 05, 2022



Abstract:Recent engineering developments in specialised computational hardware, data-acquisition and storage technology have seen the emergence of Machine Learning (ML) as a powerful form of data analysis with widespread applicability beyond its historical roots in the design of autonomous agents. However -- possibly because of its origins in the development of agents capable of self-discovery -- relatively little attention has been paid to the interaction between people and ML. In this paper we are concerned with the use of ML in automated or semi-automated tools that assist one or more human decision makers. We argue that requirements on both human and machine in this context are significantly different to the use of ML either as part of autonomous agents for self-discovery or as part statistical data analysis. Our principal position is that the design of such human-machine systems should be driven by repeated, two-way intelligibility of information rather than one-way explainability of the ML-system's recommendations. Iterated rounds of intelligible information exchange, we think, will characterise the kinds of collaboration that will be needed to understand complex phenomena for which neither man or machine have complete answers. We propose operational principles -- we call them Intelligibility Axioms -- to guide the design of a collaborative decision-support system. The principles are concerned with: (a) what it means for information provided by the human to be intelligible to the ML system; and (b) what it means for an explanation provided by an ML system to be intelligible to a human. Using examples from the literature on the use of ML for drug-design and in medicine, we demonstrate cases where the conditions of the axioms are met. We describe some additional requirements needed for the design of a truly collaborative decision-support system.
Solving Visual Analogies Using Neural Algorithmic Reasoning
Nov 19, 2021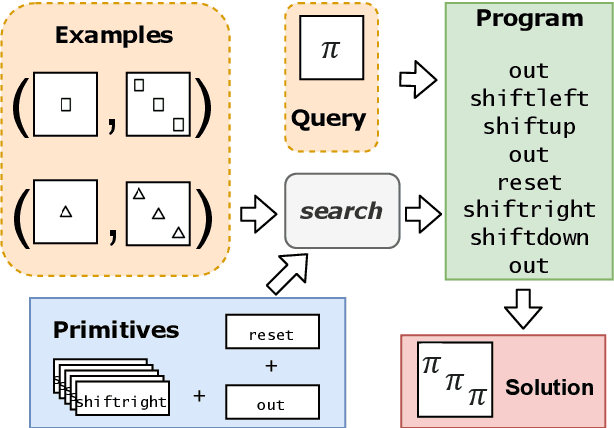

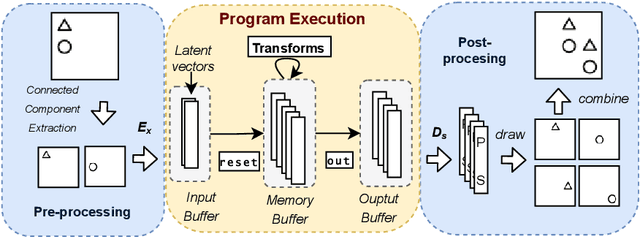
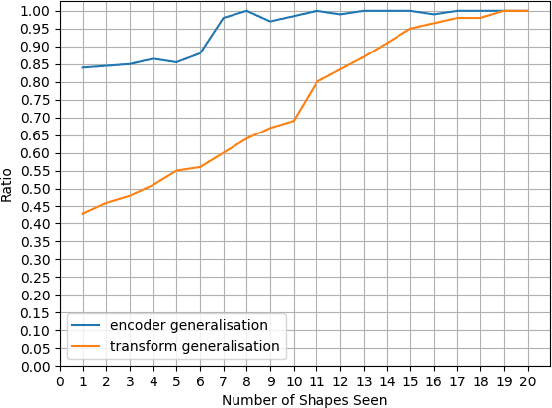
Abstract:We consider a class of visual analogical reasoning problems that involve discovering the sequence of transformations by which pairs of input/output images are related, so as to analogously transform future inputs. This program synthesis task can be easily solved via symbolic search. Using a variation of the `neural analogical reasoning' approach of (Velickovic and Blundell 2021), we instead search for a sequence of elementary neural network transformations that manipulate distributed representations derived from a symbolic space, to which input images are directly encoded. We evaluate the extent to which our `neural reasoning' approach generalizes for images with unseen shapes and positions.
Using Program Synthesis and Inductive Logic Programming to solve Bongard Problems
Oct 19, 2021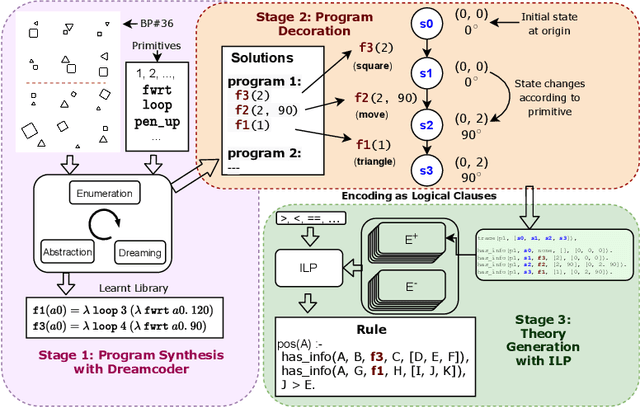
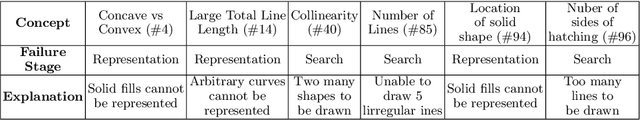
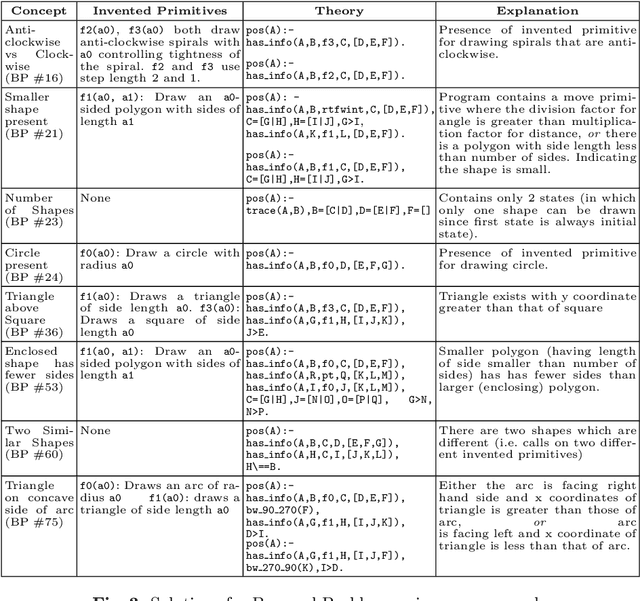
Abstract:The ability to recognise and make analogies is often used as a measure or test of human intelligence. The ability to solve Bongard problems is an example of such a test. It has also been postulated that the ability to rapidly construct novel abstractions is critical to being able to solve analogical problems. Given an image, the ability to construct a program that would generate that image is one form of abstraction, as exemplified in the Dreamcoder project. In this paper, we present a preliminary examination of whether programs constructed by Dreamcoder can be used for analogical reasoning to solve certain Bongard problems. We use Dreamcoder to discover programs that generate the images in a Bongard problem and represent each of these as a sequence of state transitions. We decorate the states using positional information in an automated manner and then encode the resulting sequence into logical facts in Prolog. We use inductive logic programming (ILP), to learn an (interpretable) theory for the abstract concept involved in an instance of a Bongard problem. Experiments on synthetically created Bongard problems for concepts such as 'above/below' and 'clockwise/counterclockwise' demonstrate that our end-to-end system can solve such problems. We study the importance and completeness of each component of our approach, highlighting its current limitations and pointing to directions for improvement in our formulation as well as in elements of any Dreamcoder-like program synthesis system used for such an approach.
Zero-Shot Dense Retrieval with Momentum Adversarial Domain Invariant Representations
Oct 14, 2021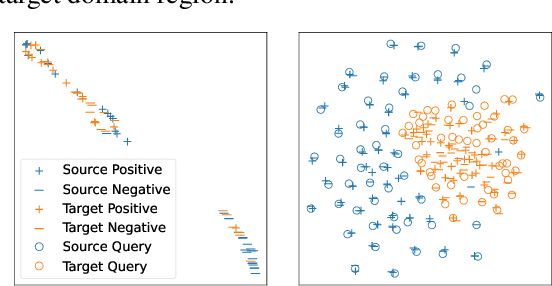
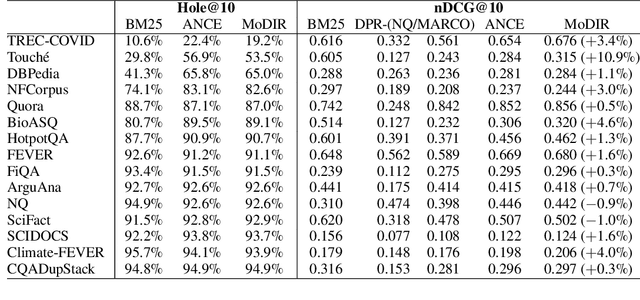
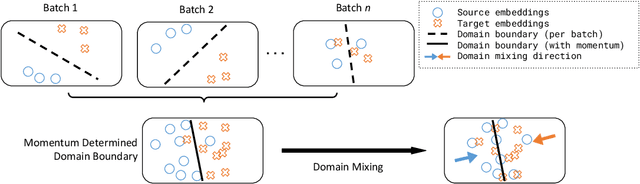
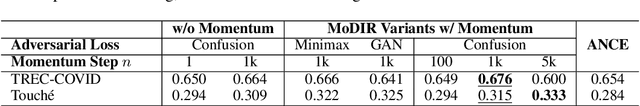
Abstract:Dense retrieval (DR) methods conduct text retrieval by first encoding texts in the embedding space and then matching them by nearest neighbor search. This requires strong locality properties from the representation space, i.e, the close allocations of each small group of relevant texts, which are hard to generalize to domains without sufficient training data. In this paper, we aim to improve the generalization ability of DR models from source training domains with rich supervision signals to target domains without any relevant labels, in the zero-shot setting. To achieve that, we propose Momentum adversarial Domain Invariant Representation learning (MoDIR), which introduces a momentum method in the DR training process to train a domain classifier distinguishing source versus target, and then adversarially updates the DR encoder to learn domain invariant representations. Our experiments show that MoDIR robustly outperforms its baselines on 10+ ranking datasets from the BEIR benchmark in the zero-shot setup, with more than 10% relative gains on datasets with enough sensitivity for DR models' evaluation. Source code of this paper will be released.
How to Tell Deep Neural Networks What We Know
Jul 21, 2021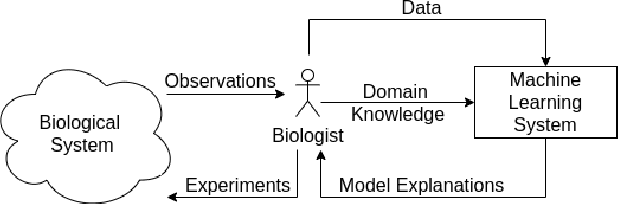
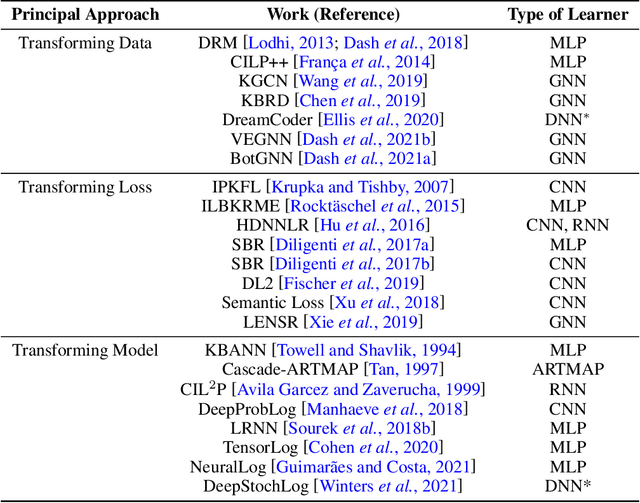
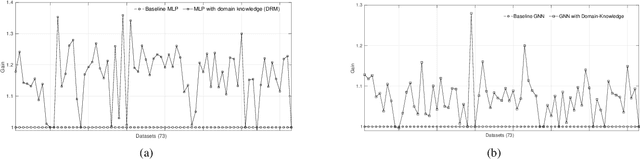
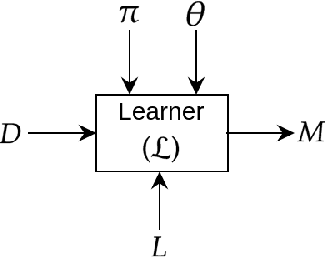
Abstract:We present a short survey of ways in which existing scientific knowledge are included when constructing models with neural networks. The inclusion of domain-knowledge is of special interest not just to constructing scientific assistants, but also, many other areas that involve understanding data using human-machine collaboration. In many such instances, machine-based model construction may benefit significantly from being provided with human-knowledge of the domain encoded in a sufficiently precise form. This paper examines the inclusion of domain-knowledge by means of changes to: the input, the loss-function, and the architecture of deep networks. The categorisation is for ease of exposition: in practice we expect a combination of such changes will be employed. In each category, we describe techniques that have been shown to yield significant changes in network performance.
Inclusion of Domain-Knowledge into GNNs using Mode-Directed Inverse Entailment
May 22, 2021
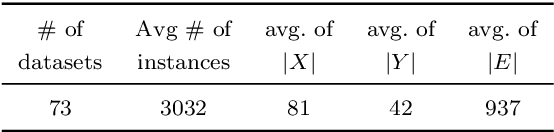
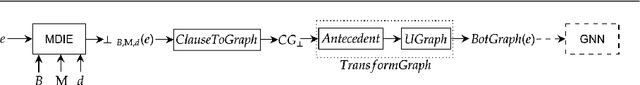
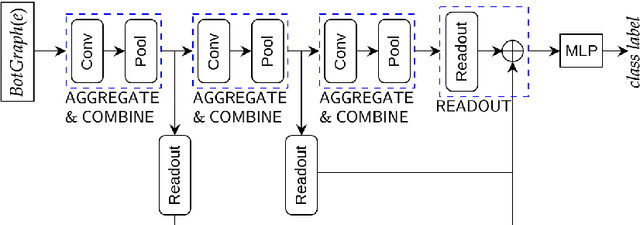
Abstract:We present a general technique for constructing Graph Neural Networks (GNNs) capable of using multi-relational domain knowledge. The technique is based on mode-directed inverse entailment (MDIE) developed in Inductive Logic Programming (ILP). Given a data instance $e$ and background knowledge $B$, MDIE identifies a most-specific logical formula $\bot_B(e)$ that contains all the relational information in $B$ that is related to $e$. We transform $\bot_B(e)$ into a corresponding "bottom-graph" that can be processed for use by standard GNN implementations. This transformation allows a principled way of incorporating generic background knowledge into GNNs: we use the term `BotGNN' for this form of graph neural networks. For several GNN variants, using real-world datasets with substantial background knowledge, we show that BotGNNs perform significantly better than both GNNs without background knowledge and a recently proposed simplified technique for including domain knowledge into GNNs. We also provide experimental evidence comparing BotGNNs favourably to multi-layer perceptrons (MLPs) that use features representing a "propositionalised" form of the background knowledge; and BotGNNs to a standard ILP based on the use of most-specific clauses. Taken together, these results point to BotGNNs as capable of combining the computational efficacy of GNNs with the representational versatility of ILP.
Incorporating Domain Knowledge into Deep Neural Networks
Mar 15, 2021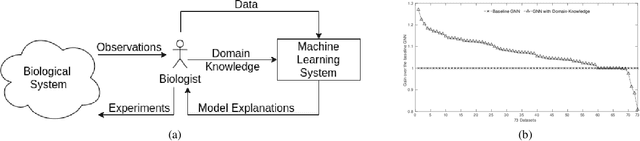
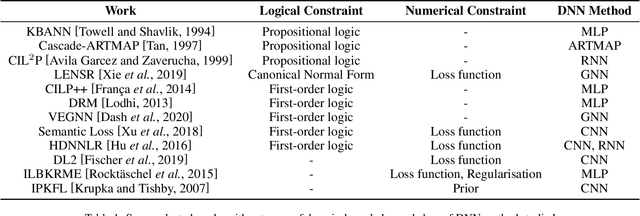
Abstract:We present a survey of ways in which domain-knowledge has been included when constructing models with neural networks. The inclusion of domain-knowledge is of special interest not just to constructing scientific assistants, but also, many other areas that involve understanding data using human-machine collaboration. In many such instances, machine-based model construction may benefit significantly from being provided with human-knowledge of the domain encoded in a sufficiently precise form. This paper examines two broad approaches to encode such knowledge--as logical and numerical constraints--and describes techniques and results obtained in several sub-categories under each of these approaches.
Constructing and Evaluating an Explainable Model for COVID-19 Diagnosis from Chest X-rays
Dec 19, 2020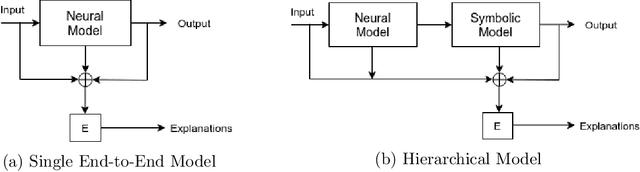
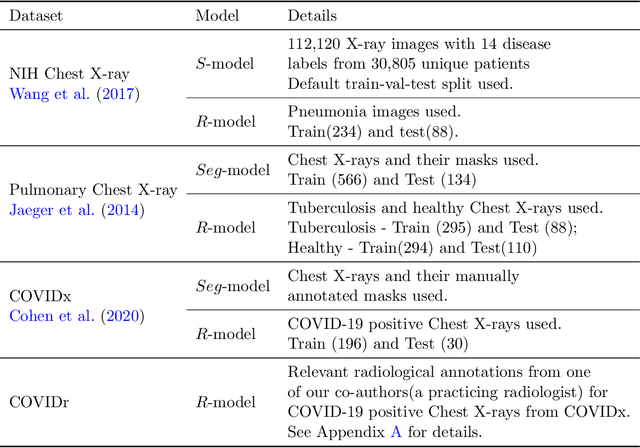
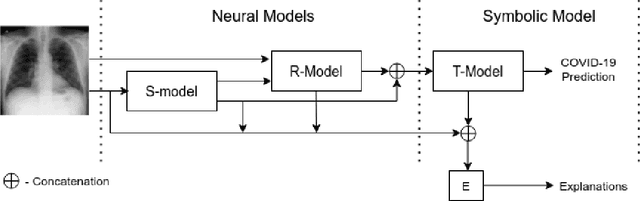
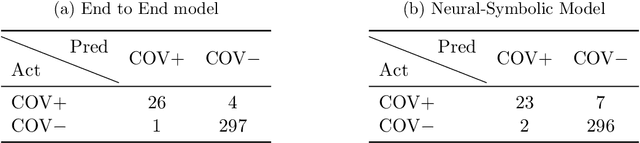
Abstract:In this paper, our focus is on constructing models to assist a clinician in the diagnosis of COVID-19 patients in situations where it is easier and cheaper to obtain X-ray data than to obtain high-quality images like those from CT scans. Deep neural networks have repeatedly been shown to be capable of constructing highly predictive models for disease detection directly from image data. However, their use in assisting clinicians has repeatedly hit a stumbling block due to their black-box nature. Some of this difficulty can be alleviated if predictions were accompanied by explanations expressed in clinically relevant terms. In this paper, deep neural networks are used to extract domain-specific features(morphological features like ground-glass opacity and disease indications like pneumonia) directly from the image data. Predictions about these features are then used to construct a symbolic model (a decision tree) for the diagnosis of COVID-19 from chest X-rays, accompanied with two kinds of explanations: visual (saliency maps, derived from the neural stage), and textual (logical descriptions, derived from the symbolic stage). A radiologist rates the usefulness of the visual and textual explanations. Our results demonstrate that neural models can be employed usefully in identifying domain-specific features from low-level image data; that textual explanations in terms of clinically relevant features may be useful; and that visual explanations will need to be clinically meaningful to be useful.
Incorporating Symbolic Domain Knowledge into Graph Neural Networks
Oct 23, 2020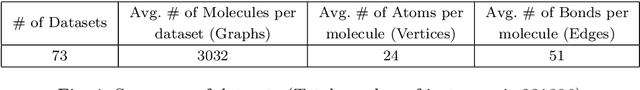
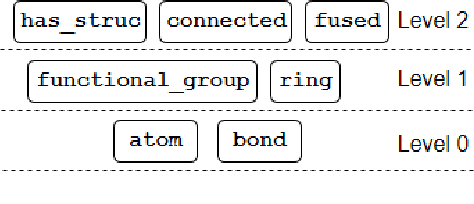
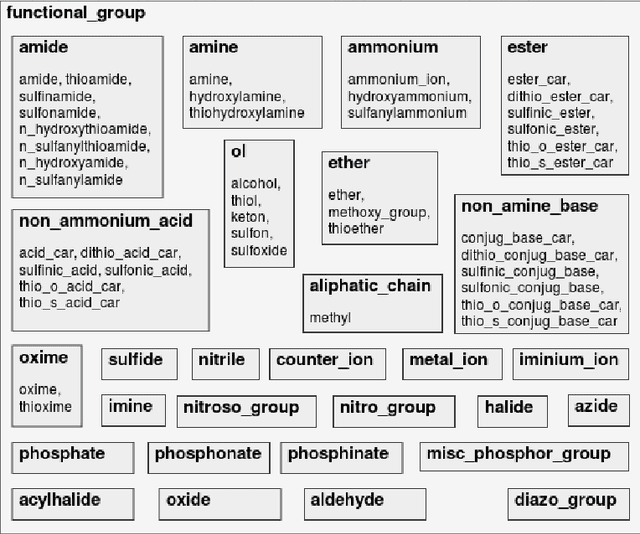
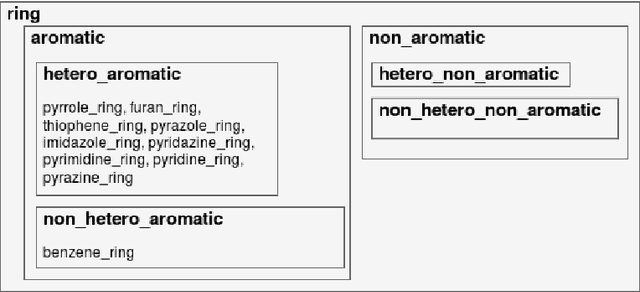
Abstract:Our interest is in scientific problems with the following characteristics: (1) Data are naturally represented as graphs; (2) The amount of data available is typically small; and (3) There is significant domain-knowledge, usually expressed in some symbolic form. These kinds of problems have been addressed effectively in the past by Inductive Logic Programming (ILP), by virtue of 2 important characteristics: (a) The use of a representation language that easily captures the relation encoded in graph-structured data, and (b) The inclusion of prior information encoded as domain-specific relations, that can alleviate problems of data scarcity, and construct new relations. Recent advances have seen the emergence of deep neural networks specifically developed for graph-structured data (Graph-based Neural Networks, or GNNs). While GNNs have been shown to be able to handle graph-structured data, less has been done to investigate the inclusion of domain-knowledge. Here we investigate this aspect of GNNs empirically by employing an operation we term "vertex-enrichment" and denote the corresponding GNNs as "VEGNNs". Using over 70 real-world datasets and substantial amounts of symbolic domain-knowledge, we examine the result of vertex-enrichment across 5 different variants of GNNs. Our results provide support for the following: (a) Inclusion of domain-knowledge by vertex-enrichment can significantly improve the performance of a GNN. That is, the performance VEGNNs is significantly better than GNNs across all GNN variants; (b) The inclusion of domain-specific relations constructed using ILP improves the performance of VEGNNs, across all GNN variants. Taken together, the results provide evidence that it is possible to incorporate symbolic domain knowledge into a GNN, and that ILP can play an important role in providing high-level relationships that are not easily discovered by a GNN.
Dr.Quad at MEDIQA 2019: Towards Textual Inference and Question Entailment using contextualized representations
Jul 23, 2019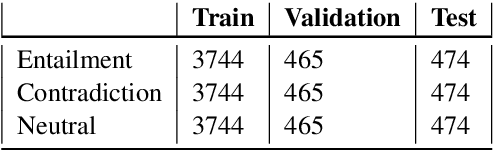
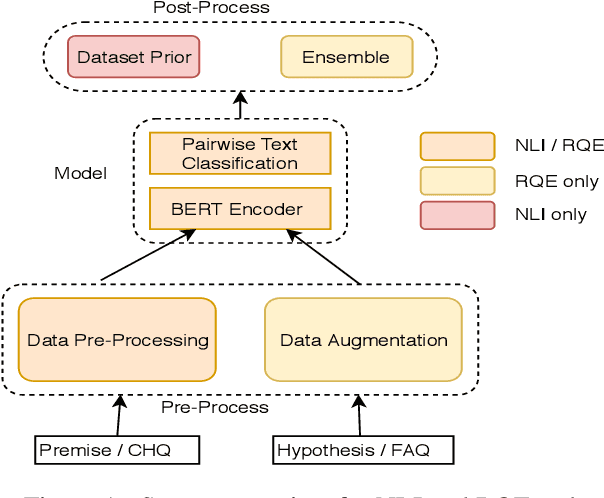

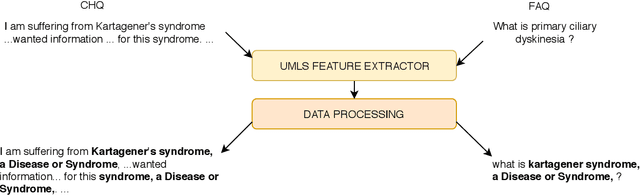
Abstract:This paper presents the submissions by Team Dr.Quad to the ACL-BioNLP 2019 shared task on Textual Inference and Question Entailment in the Medical Domain. Our system is based on the prior work Liu et al. (2019) which uses a multi-task objective function for textual entailment. In this work, we explore different strategies for generalizing state-of-the-art language understanding models to the specialized medical domain. Our results on the shared task demonstrate that incorporating domain knowledge through data augmentation is a powerful strategy for addressing challenges posed by specialized domains such as medicine.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge