Arash A. Amini
Laplacian Kernelized Bandit
Jan 01, 2026Abstract:We study multi-user contextual bandits where users are related by a graph and their reward functions exhibit both non-linear behavior and graph homophily. We introduce a principled joint penalty for the collection of user reward functions $\{f_u\}$, combining a graph smoothness term based on RKHS distances with an individual roughness penalty. Our central contribution is proving that this penalty is equivalent to the squared norm within a single, unified \emph{multi-user RKHS}. We explicitly derive its reproducing kernel, which elegantly fuses the graph Laplacian with the base arm kernel. This unification allows us to reframe the problem as learning a single ''lifted'' function, enabling the design of principled algorithms, \texttt{LK-GP-UCB} and \texttt{LK-GP-TS}, that leverage Gaussian Process posteriors over this new kernel for exploration. We provide high-probability regret bounds that scale with an \emph{effective dimension} of the multi-user kernel, replacing dependencies on user count or ambient dimension. Empirically, our methods outperform strong linear and non-graph-aware baselines in non-linear settings and remain competitive even when the true rewards are linear. Our work delivers a unified, theoretically grounded, and practical framework that bridges Laplacian regularization with kernelized bandits for structured exploration.
Sharp Bounds for Poly-GNNs and the Effect of Graph Noise
Jul 28, 2024Abstract:We investigate the classification performance of graph neural networks with graph-polynomial features, poly-GNNs, on the problem of semi-supervised node classification. We analyze poly-GNNs under a general contextual stochastic block model (CSBM) by providing a sharp characterization of the rate of separation between classes in their output node representations. A question of interest is whether this rate depends on the depth of the network $k$, i.e., whether deeper networks can achieve a faster separation? We provide a negative answer to this question: for a sufficiently large graph, a depth $k > 1$ poly-GNN exhibits the same rate of separation as a depth $k=1$ counterpart. Our analysis highlights and quantifies the impact of ``graph noise'' in deep GNNs and shows how noise in the graph structure can dominate other sources of signal in the graph, negating any benefit further aggregation provides. Our analysis also reveals subtle differences between even and odd-layered GNNs in how the feature noise propagates.
Graph Neural Thompson Sampling
Jun 15, 2024



Abstract:We consider an online decision-making problem with a reward function defined over graph-structured data. We formally formulate the problem as an instance of graph action bandit. We then propose \texttt{GNN-TS}, a Graph Neural Network (GNN) powered Thompson Sampling (TS) algorithm which employs a GNN approximator for estimating the mean reward function and the graph neural tangent features for uncertainty estimation. We prove that, under certain boundness assumptions on the reward function, GNN-TS achieves a state-of-the-art regret bound which is (1) sub-linear of order $\tilde{\mathcal{O}}((\tilde{d} T)^{1/2})$ in the number of interaction rounds, $T$, and a notion of effective dimension $\tilde{d}$, and (2) independent of the number of graph nodes. Empirical results validate that our proposed \texttt{GNN-TS} exhibits competitive performance and scales well on graph action bandit problems.
Network two-sample test for block models
Jun 10, 2024



Abstract:We consider the two-sample testing problem for networks, where the goal is to determine whether two sets of networks originated from the same stochastic model. Assuming no vertex correspondence and allowing for different numbers of nodes, we address a fundamental network testing problem that goes beyond simple adjacency matrix comparisons. We adopt the stochastic block model (SBM) for network distributions, due to their interpretability and the potential to approximate more general models. The lack of meaningful node labels and vertex correspondence translate to a graph matching challenge when developing a test for SBMs. We introduce an efficient algorithm to match estimated network parameters, allowing us to properly combine and contrast information within and across samples, leading to a powerful test. We show that the matching algorithm, and the overall test are consistent, under mild conditions on the sparsity of the networks and the sample sizes, and derive a chi-squared asymptotic null distribution for the test. Through a mixture of theoretical insights and empirical validations, including experiments with both synthetic and real-world data, this study advances robust statistical inference for complex network data.
Step and Smooth Decompositions as Topological Clustering
Nov 09, 2023



Abstract:We investigate a class of recovery problems for which observations are a noisy combination of continuous and step functions. These problems can be seen as non-injective instances of non-linear ICA with direct applications to image decontamination for magnetic resonance imaging. Alternately, the problem can be viewed as clustering in the presence of structured (smooth) contaminant. We show that a global topological property (graph connectivity) interacts with a local property (the degree of smoothness of the continuous component) to determine conditions under which the components are identifiable. Additionally, a practical estimation algorithm is provided for the case when the contaminant lies in a reproducing kernel Hilbert space of continuous functions. Algorithm effectiveness is demonstrated through a series of simulations and real-world studies.
Simplifying GNN Performance with Low Rank Kernel Models
Oct 08, 2023Abstract:We revisit recent spectral GNN approaches to semi-supervised node classification (SSNC). We posit that many of the current GNN architectures may be over-engineered. Instead, simpler, traditional methods from nonparametric estimation, applied in the spectral domain, could replace many deep-learning inspired GNN designs. These conventional techniques appear to be well suited for a variety of graph types reaching state-of-the-art performance on many of the common SSNC benchmarks. Additionally, we show that recent performance improvements in GNN approaches may be partially attributed to shifts in evaluation conventions. Lastly, an ablative study is conducted on the various hyperparameters associated with GNN spectral filtering techniques. Code available at: https://github.com/lucianoAvinas/lowrank-gnn-kernels
Nested stochastic block model for simultaneously clustering networks and nodes
Jul 18, 2023



Abstract:We introduce the nested stochastic block model (NSBM) to cluster a collection of networks while simultaneously detecting communities within each network. NSBM has several appealing features including the ability to work on unlabeled networks with potentially different node sets, the flexibility to model heterogeneous communities, and the means to automatically select the number of classes for the networks and the number of communities within each network. This is accomplished via a Bayesian model, with a novel application of the nested Dirichlet process (NDP) as a prior to jointly model the between-network and within-network clusters. The dependency introduced by the network data creates nontrivial challenges for the NDP, especially in the development of efficient samplers. For posterior inference, we propose several Markov chain Monte Carlo algorithms including a standard Gibbs sampler, a collapsed Gibbs sampler, and two blocked Gibbs samplers that ultimately return two levels of clustering labels from both within and across the networks. Extensive simulation studies are carried out which demonstrate that the model provides very accurate estimates of both levels of the clustering structure. We also apply our model to two social network datasets that cannot be analyzed using any previous method in the literature due to the anonymity of the nodes and the varying number of nodes in each network.
LapGM: A Multisequence MR Bias Correction and Normalization Model
Sep 27, 2022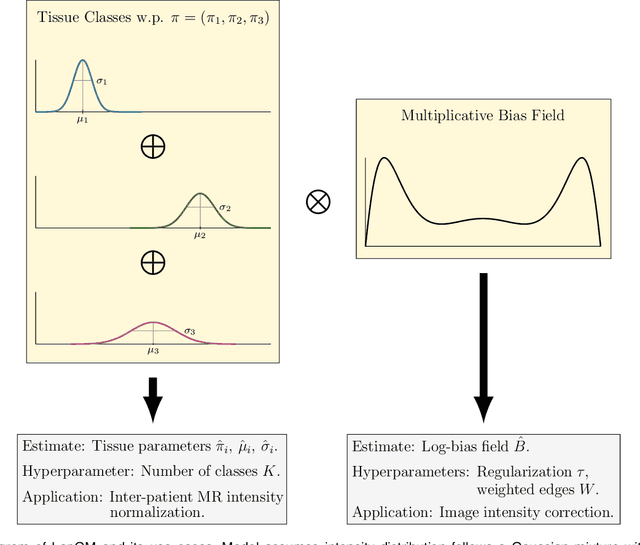
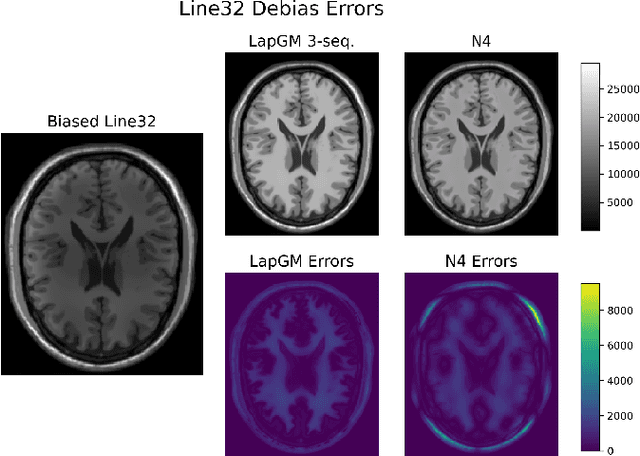
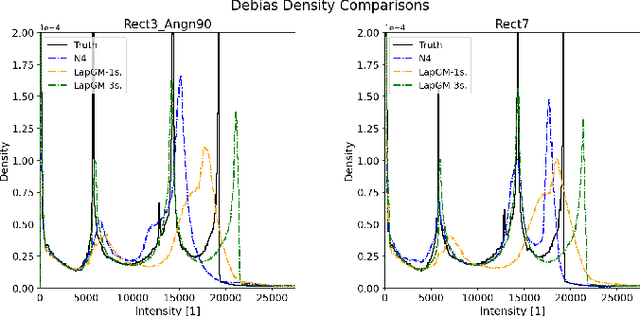
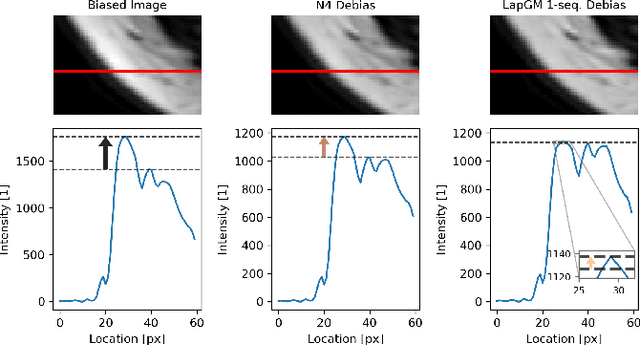
Abstract:A spatially regularized Gaussian mixture model, LapGM, is proposed for the bias field correction and magnetic resonance normalization problem. The proposed spatial regularizer gives practitioners fine-tuned control between balancing bias field removal and preserving image contrast preservation for multi-sequence, magnetic resonance images. The fitted Gaussian parameters of LapGM serve as control values which can be used to normalize image intensities across different patient scans. LapGM is compared to well-known debiasing algorithm N4ITK in both the single and multi-sequence setting. As a normalization procedure, LapGM is compared to known techniques such as: max normalization, Z-score normalization, and a water-masked region-of-interest normalization. Lastly a CUDA-accelerated Python package $\texttt{lapgm}$ is provided from the authors for use.
Target alignment in truncated kernel ridge regression
Jun 28, 2022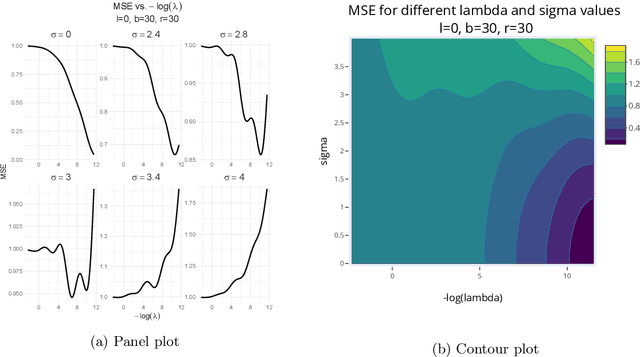
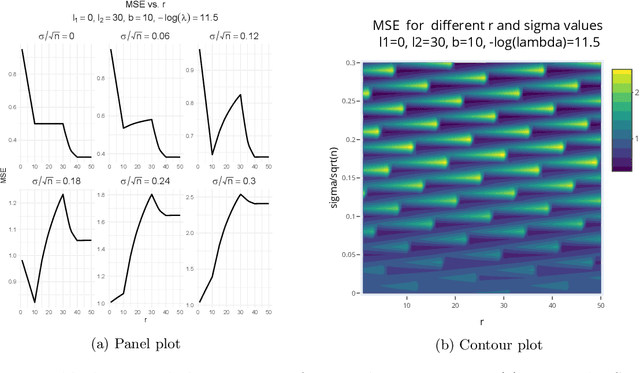
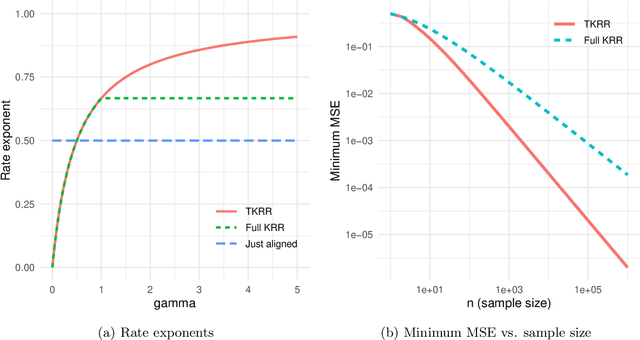
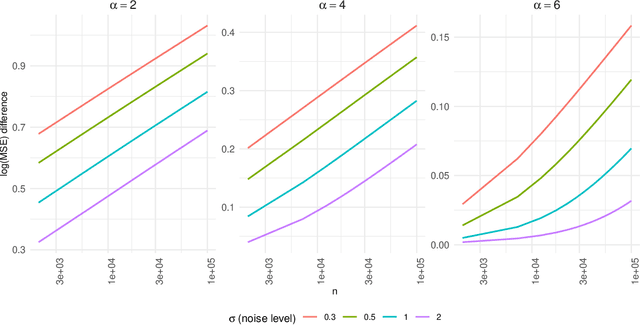
Abstract:Kernel ridge regression (KRR) has recently attracted renewed interest due to its potential for explaining the transient effects, such as double descent, that emerge during neural network training. In this work, we study how the alignment between the target function and the kernel affects the performance of the KRR. We focus on the truncated KRR (TKRR) which utilizes an additional parameter that controls the spectral truncation of the kernel matrix. We show that for polynomial alignment, there is an \emph{over-aligned} regime, in which TKRR can achieve a faster rate than what is achievable by full KRR. The rate of TKRR can improve all the way to the parametric rate, while that of full KRR is capped at a sub-optimal value. This shows that target alignemnt can be better leveraged by utilizing spectral truncation in kernel methods. We also consider the bandlimited alignment setting and show that the regularization surface of TKRR can exhibit transient effects including multiple descent and non-monotonic behavior. Our results show that there is a strong and quantifable relation between the shape of the \emph{alignment spectrum} and the generalization performance of kernel methods, both in terms of rates and in finite samples.
A non-graphical representation of conditional independence via the neighbourhood lattice
Jun 12, 2022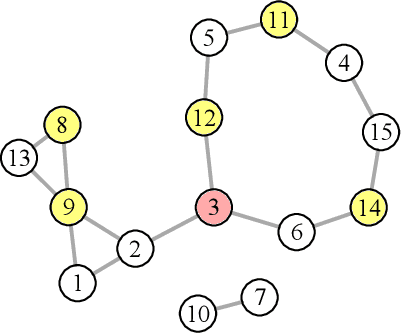
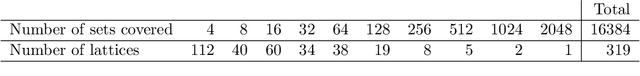
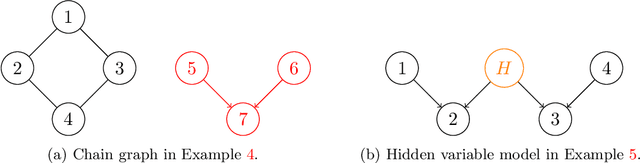
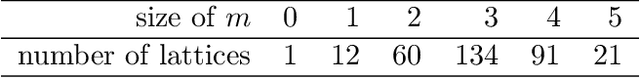
Abstract:We introduce and study the neighbourhood lattice decomposition of a distribution, which is a compact, non-graphical representation of conditional independence that is valid in the absence of a faithful graphical representation. The idea is to view the set of neighbourhoods of a variable as a subset lattice, and partition this lattice into convex sublattices, each of which directly encodes a collection of conditional independence relations. We show that this decomposition exists in any compositional graphoid and can be computed efficiently and consistently in high-dimensions. {In particular, this gives a way to encode all of independence relations implied by a distribution that satisfies the composition axiom, which is strictly weaker than the faithfulness assumption that is typically assumed by graphical approaches.} We also discuss various special cases such as graphical models and projection lattices, each of which has intuitive interpretations. Along the way, we see how this problem is closely related to neighbourhood regression, which has been extensively studied in the context of graphical models and structural equations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge