Linked Component Analysis from Matrices to High Order Tensors: Applications to Biomedical Data
Aug 29, 2015Guoxu Zhou, Qibin Zhao, Yu Zhang, Tülay Adalı, Shengli Xie, Andrzej Cichocki
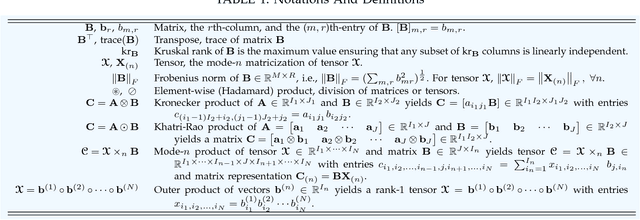
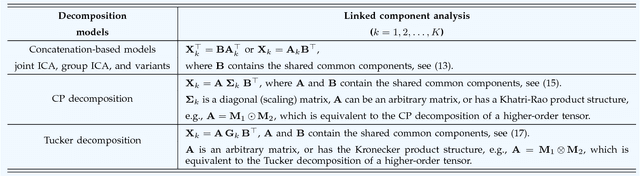
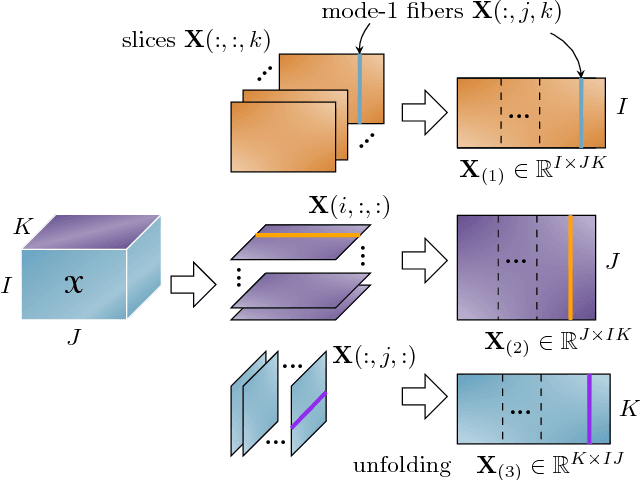

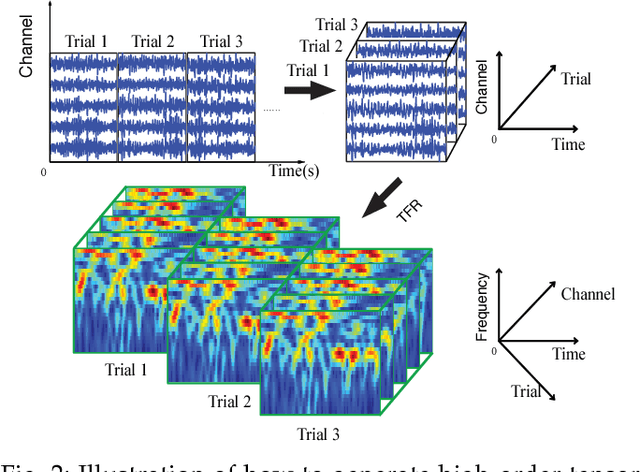

With the increasing availability of various sensor technologies, we now have access to large amounts of multi-block (also called multi-set, multi-relational, or multi-view) data that need to be jointly analyzed to explore their latent connections. Various component analysis methods have played an increasingly important role for the analysis of such coupled data. In this paper, we first provide a brief review of existing matrix-based (two-way) component analysis methods for the joint analysis of such data with a focus on biomedical applications. Then, we discuss their important extensions and generalization to multi-block multiway (tensor) data. We show how constrained multi-block tensor decomposition methods are able to extract similar or statistically dependent common features that are shared by all blocks, by incorporating the multiway nature of data. Special emphasis is given to the flexible common and individual feature analysis of multi-block data with the aim to simultaneously extract common and individual latent components with desired properties and types of diversity. Illustrative examples are given to demonstrate their effectiveness for biomedical data analysis.
Bayesian Sparse Tucker Models for Dimension Reduction and Tensor Completion
May 10, 2015Qibin Zhao, Liqing Zhang, Andrzej Cichocki
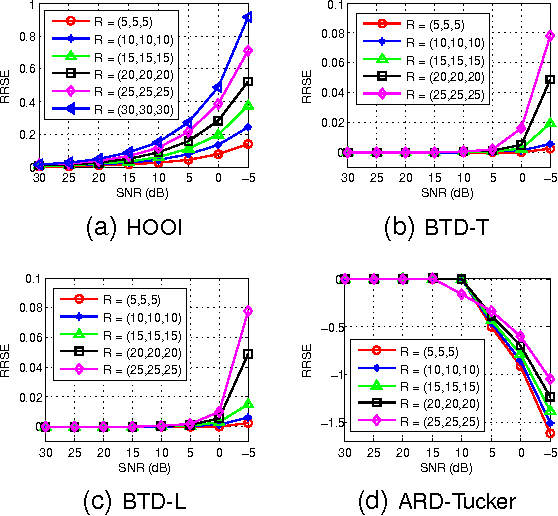
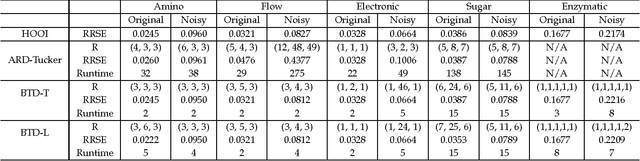
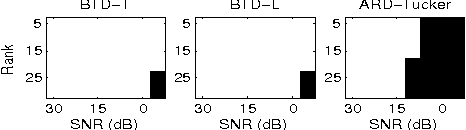
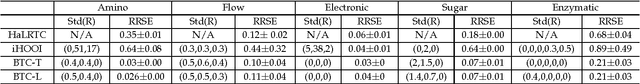
Tucker decomposition is the cornerstone of modern machine learning on tensorial data analysis, which have attracted considerable attention for multiway feature extraction, compressive sensing, and tensor completion. The most challenging problem is related to determination of model complexity (i.e., multilinear rank), especially when noise and missing data are present. In addition, existing methods cannot take into account uncertainty information of latent factors, resulting in low generalization performance. To address these issues, we present a class of probabilistic generative Tucker models for tensor decomposition and completion with structural sparsity over multilinear latent space. To exploit structural sparse modeling, we introduce two group sparsity inducing priors by hierarchial representation of Laplace and Student-t distributions, which facilitates fully posterior inference. For model learning, we derived variational Bayesian inferences over all model (hyper)parameters, and developed efficient and scalable algorithms based on multilinear operations. Our methods can automatically adapt model complexity and infer an optimal multilinear rank by the principle of maximum lower bound of model evidence. Experimental results and comparisons on synthetic, chemometrics and neuroimaging data demonstrate remarkable performance of our models for recovering ground-truth of multilinear rank and missing entries.
Bayesian Robust Tensor Factorization for Incomplete Multiway Data
Apr 16, 2015Qibin Zhao, Guoxu Zhou, Liqing Zhang, Andrzej Cichocki, Shun-ichi Amari
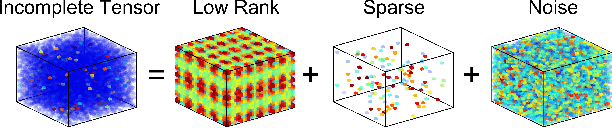
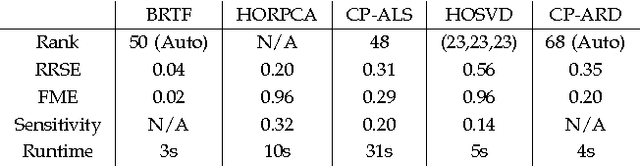
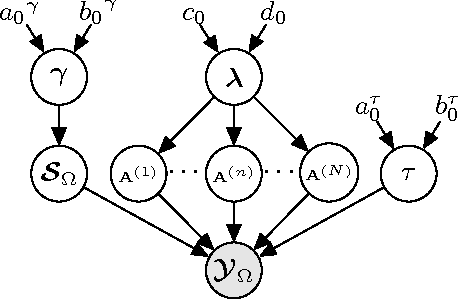
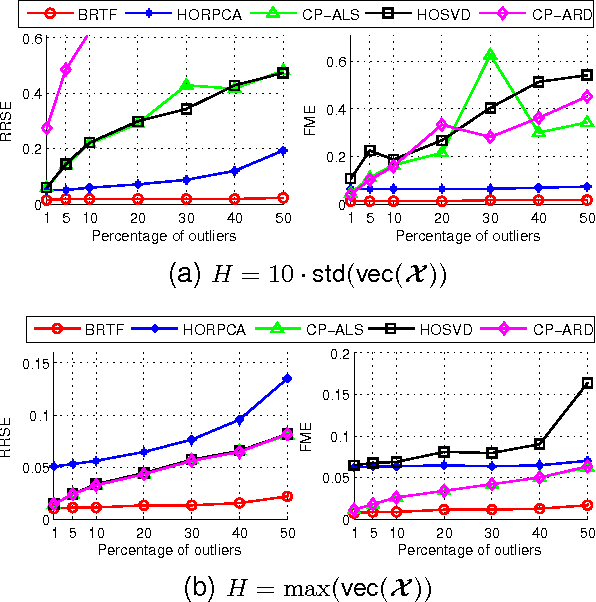
We propose a generative model for robust tensor factorization in the presence of both missing data and outliers. The objective is to explicitly infer the underlying low-CP-rank tensor capturing the global information and a sparse tensor capturing the local information (also considered as outliers), thus providing the robust predictive distribution over missing entries. The low-CP-rank tensor is modeled by multilinear interactions between multiple latent factors on which the column sparsity is enforced by a hierarchical prior, while the sparse tensor is modeled by a hierarchical view of Student-$t$ distribution that associates an individual hyperparameter with each element independently. For model learning, we develop an efficient closed-form variational inference under a fully Bayesian treatment, which can effectively prevent the overfitting problem and scales linearly with data size. In contrast to existing related works, our method can perform model selection automatically and implicitly without need of tuning parameters. More specifically, it can discover the groundtruth of CP rank and automatically adapt the sparsity inducing priors to various types of outliers. In addition, the tradeoff between the low-rank approximation and the sparse representation can be optimized in the sense of maximum model evidence. The extensive experiments and comparisons with many state-of-the-art algorithms on both synthetic and real-world datasets demonstrate the superiorities of our method from several perspectives.
Bayesian CP Factorization of Incomplete Tensors with Automatic Rank Determination
Oct 09, 2014Qibin Zhao, Liqing Zhang, Andrzej Cichocki
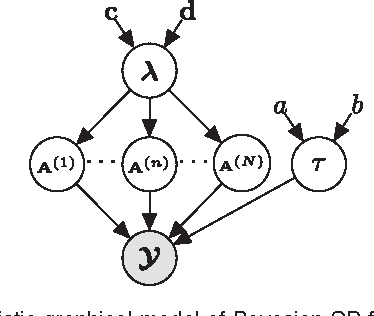
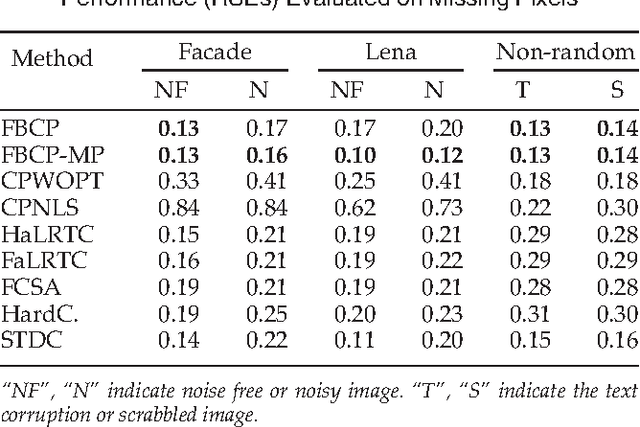
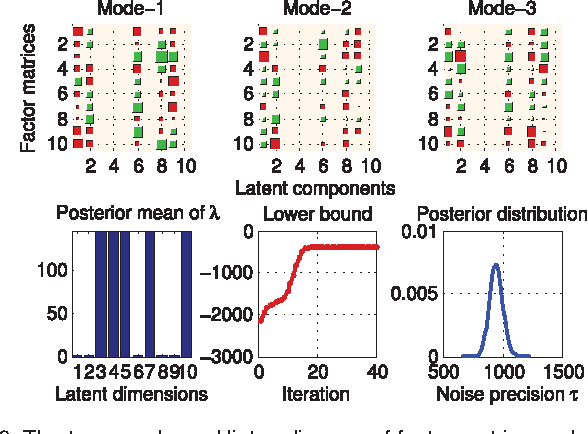
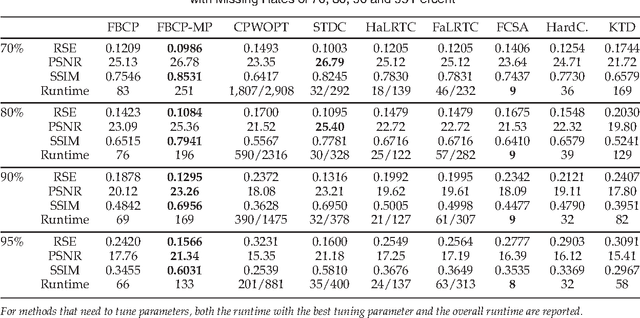
CANDECOMP/PARAFAC (CP) tensor factorization of incomplete data is a powerful technique for tensor completion through explicitly capturing the multilinear latent factors. The existing CP algorithms require the tensor rank to be manually specified, however, the determination of tensor rank remains a challenging problem especially for CP rank. In addition, existing approaches do not take into account uncertainty information of latent factors, as well as missing entries. To address these issues, we formulate CP factorization using a hierarchical probabilistic model and employ a fully Bayesian treatment by incorporating a sparsity-inducing prior over multiple latent factors and the appropriate hyperpriors over all hyperparameters, resulting in automatic rank determination. To learn the model, we develop an efficient deterministic Bayesian inference algorithm, which scales linearly with data size. Our method is characterized as a tuning parameter-free approach, which can effectively infer underlying multilinear factors with a low-rank constraint, while also providing predictive distributions over missing entries. Extensive simulations on synthetic data illustrate the intrinsic capability of our method to recover the ground-truth of CP rank and prevent the overfitting problem, even when a large amount of entries are missing. Moreover, the results from real-world applications, including image inpainting and facial image synthesis, demonstrate that our method outperforms state-of-the-art approaches for both tensor factorization and tensor completion in terms of predictive performance.
Feature Learning from Incomplete EEG with Denoising Autoencoder
Oct 03, 2014Junhua Li, Zbigniew Struzik, Liqing Zhang, Andrzej Cichocki
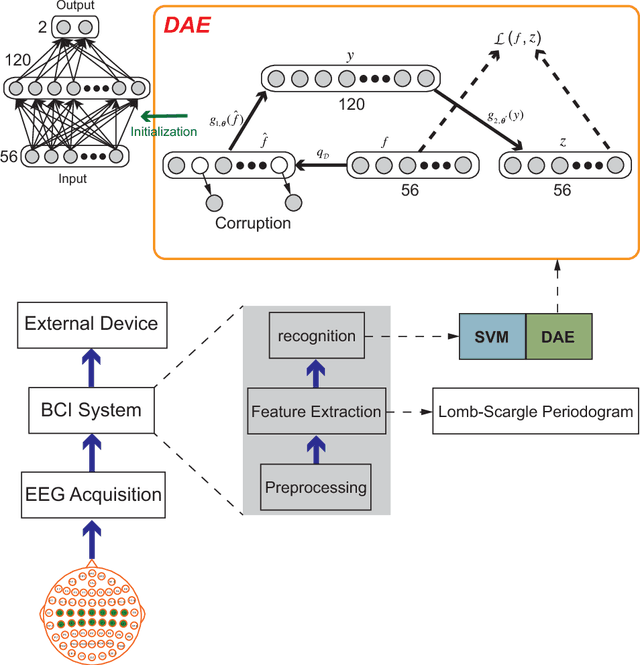
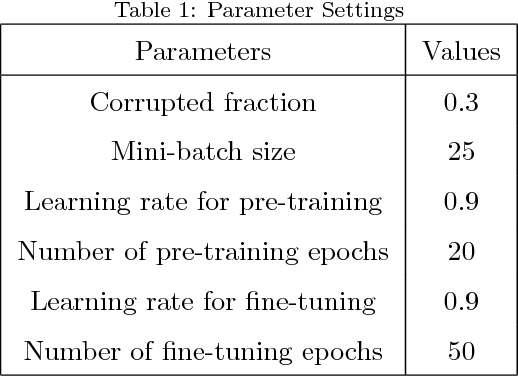
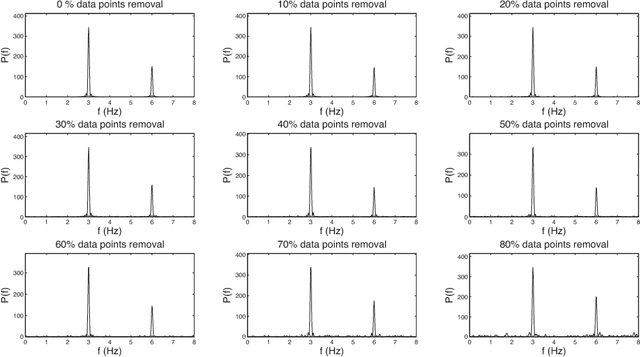
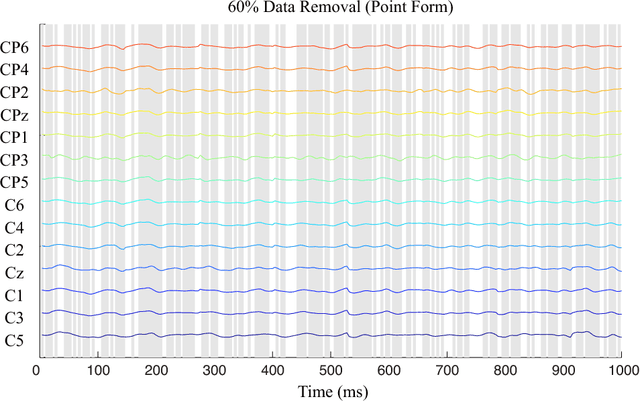
An alternative pathway for the human brain to communicate with the outside world is by means of a brain computer interface (BCI). A BCI can decode electroencephalogram (EEG) signals of brain activities, and then send a command or an intent to an external interactive device, such as a wheelchair. The effectiveness of the BCI depends on the performance in decoding the EEG. Usually, the EEG is contaminated by different kinds of artefacts (e.g., electromyogram (EMG), background activity), which leads to a low decoding performance. A number of filtering methods can be utilized to remove or weaken the effects of artefacts, but they generally fail when the EEG contains extreme artefacts. In such cases, the most common approach is to discard the whole data segment containing extreme artefacts. This causes the fatal drawback that the BCI cannot output decoding results during that time. In order to solve this problem, we employ the Lomb-Scargle periodogram to estimate the spectral power from incomplete EEG (after removing only parts contaminated by artefacts), and Denoising Autoencoder (DAE) for learning. The proposed method is evaluated with motor imagery EEG data. The results show that our method can successfully decode incomplete EEG to good effect.
Multi-tensor Completion for Estimating Missing Values in Video Data
Sep 01, 2014Chao Li, Lili Guo, Andrzej Cichocki
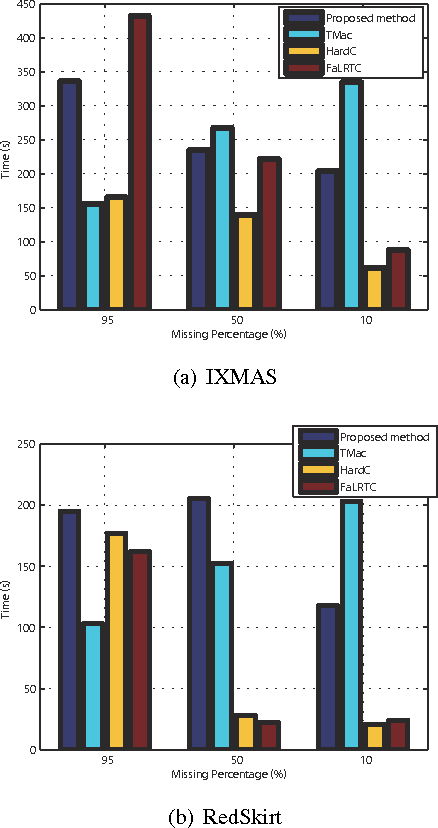
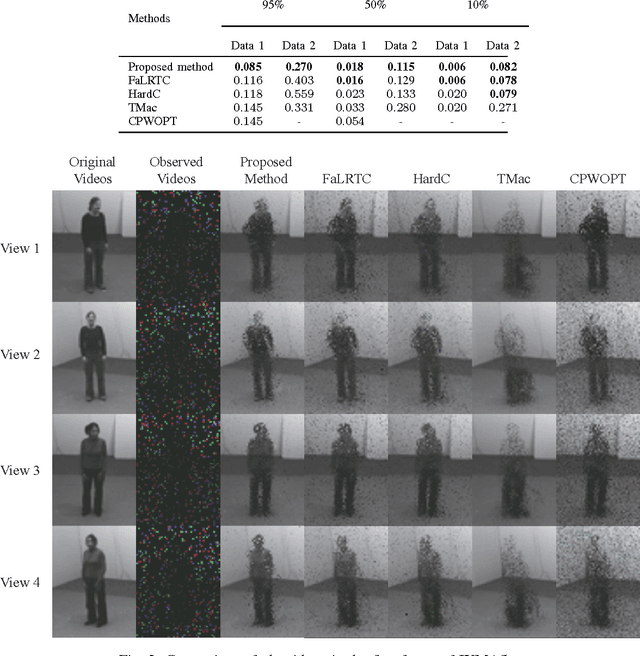
Many tensor-based data completion methods aim to solve image and video in-painting problems. But, all methods were only developed for a single dataset. In most of real applications, we can usually obtain more than one dataset to reflect one phenomenon, and all the datasets are mutually related in some sense. Thus one question raised whether such the relationship can improve the performance of data completion or not? In the paper, we proposed a novel and efficient method by exploiting the relationship among datasets for multi-video data completion. Numerical results show that the proposed method significantly improve the performance of video in-painting, particularly in the case of very high missing percentage.
Frequency Recognition in SSVEP-based BCI using Multiset Canonical Correlation Analysis
Jan 16, 2014Yu Zhang, Guoxu Zhou, Jing Jin, Xingyu Wang, Andrzej Cichocki
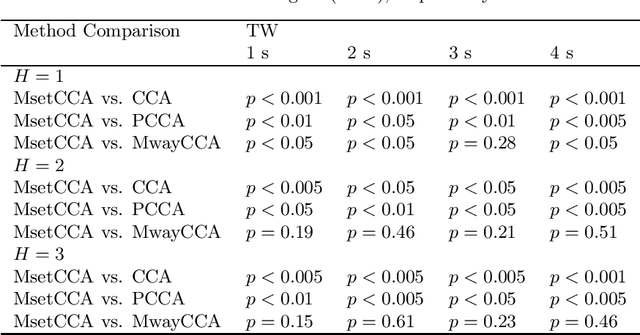
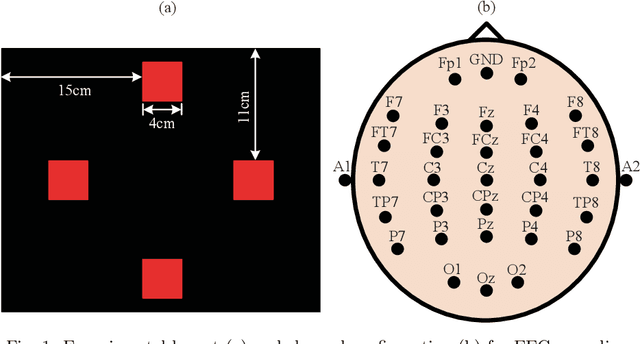

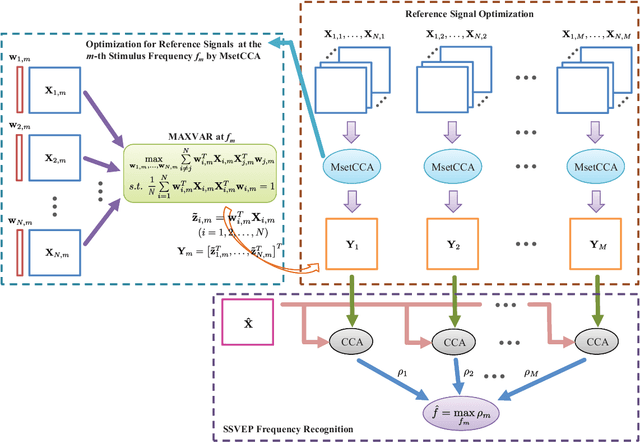
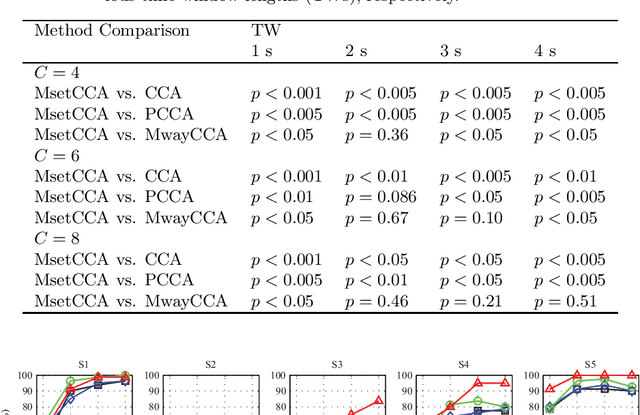
Canonical correlation analysis (CCA) has been one of the most popular methods for frequency recognition in steady-state visual evoked potential (SSVEP)-based brain-computer interfaces (BCIs). Despite its efficiency, a potential problem is that using pre-constructed sine-cosine waves as the required reference signals in the CCA method often does not result in the optimal recognition accuracy due to their lack of features from the real EEG data. To address this problem, this study proposes a novel method based on multiset canonical correlation analysis (MsetCCA) to optimize the reference signals used in the CCA method for SSVEP frequency recognition. The MsetCCA method learns multiple linear transforms that implement joint spatial filtering to maximize the overall correlation among canonical variates, and hence extracts SSVEP common features from multiple sets of EEG data recorded at the same stimulus frequency. The optimized reference signals are formed by combination of the common features and completely based on training data. Experimental study with EEG data from ten healthy subjects demonstrates that the MsetCCA method improves the recognition accuracy of SSVEP frequency in comparison with the CCA method and other two competing methods (multiway CCA (MwayCCA) and phase constrained CCA (PCCA)), especially for a small number of channels and a short time window length. The superiority indicates that the proposed MsetCCA method is a new promising candidate for frequency recognition in SSVEP-based BCIs.
Accelerated Canonical Polyadic Decomposition by Using Mode Reduction
Jun 25, 2013Guoxu Zhou, Andrzej Cichocki, Shengli Xie
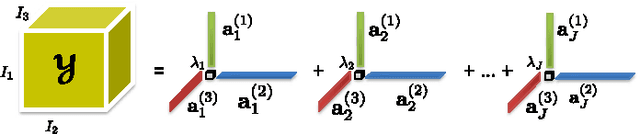
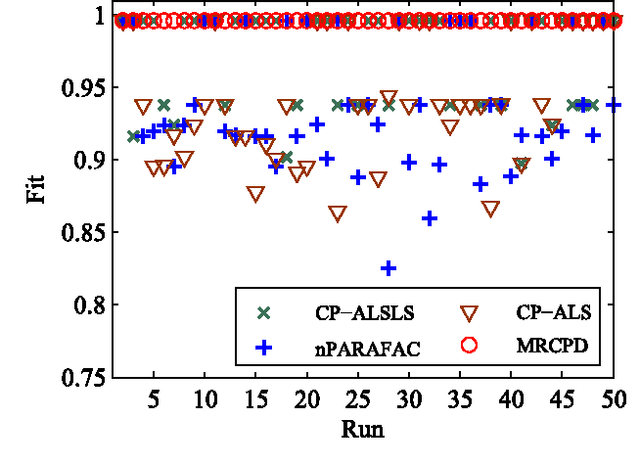
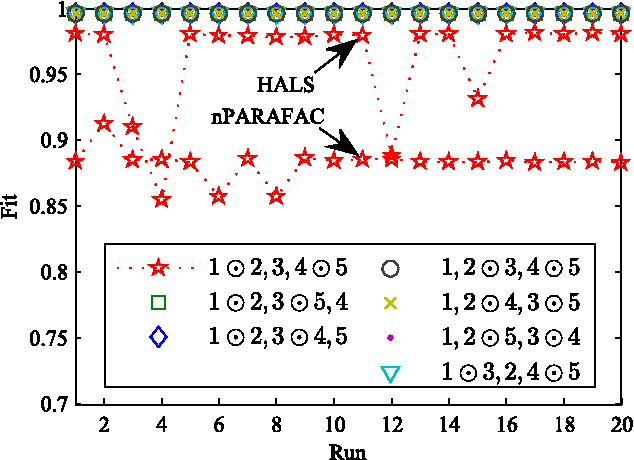
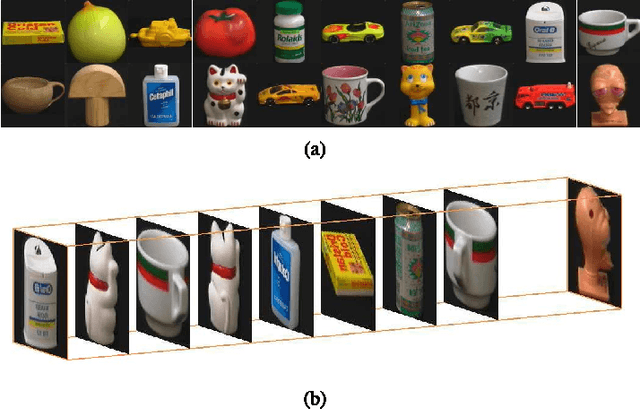
Canonical Polyadic (or CANDECOMP/PARAFAC, CP) decompositions (CPD) are widely applied to analyze high order tensors. Existing CPD methods use alternating least square (ALS) iterations and hence need to unfold tensors to each of the $N$ modes frequently, which is one major bottleneck of efficiency for large-scale data and especially when $N$ is large. To overcome this problem, in this paper we proposed a new CPD method which converts the original $N$th ($N>3$) order tensor to a 3rd-order tensor first. Then the full CPD is realized by decomposing this mode reduced tensor followed by a Khatri-Rao product projection procedure. This way is quite efficient as unfolding to each of the $N$ modes are avoided, and dimensionality reduction can also be easily incorporated to further improve the efficiency. We show that, under mild conditions, any $N$th-order CPD can be converted into a 3rd-order case but without destroying the essential uniqueness, and theoretically gives the same results as direct $N$-way CPD methods. Simulations show that, compared with state-of-the-art CPD methods, the proposed method is more efficient and escape from local solutions more easily.
Tensor Decompositions: A New Concept in Brain Data Analysis?
May 02, 2013Andrzej Cichocki
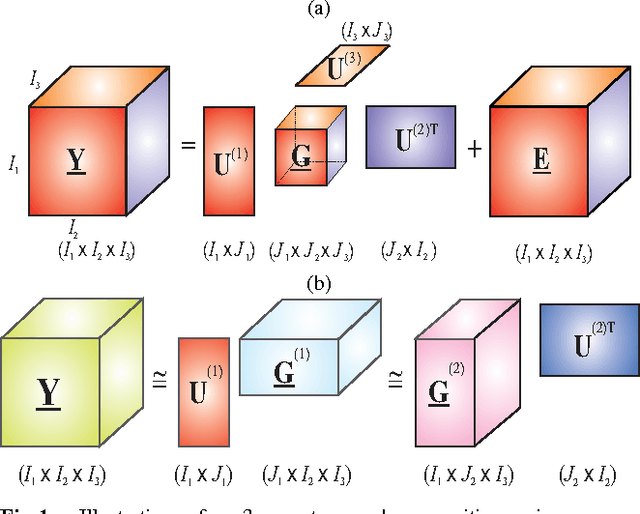
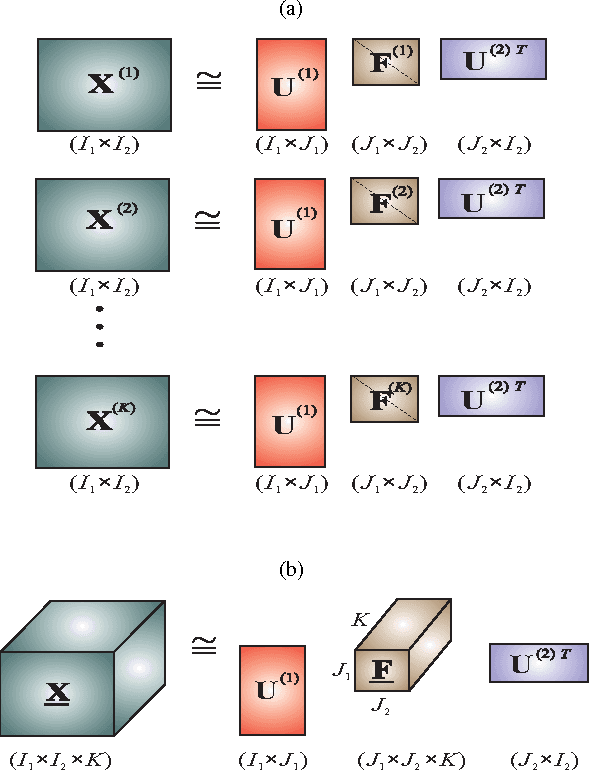
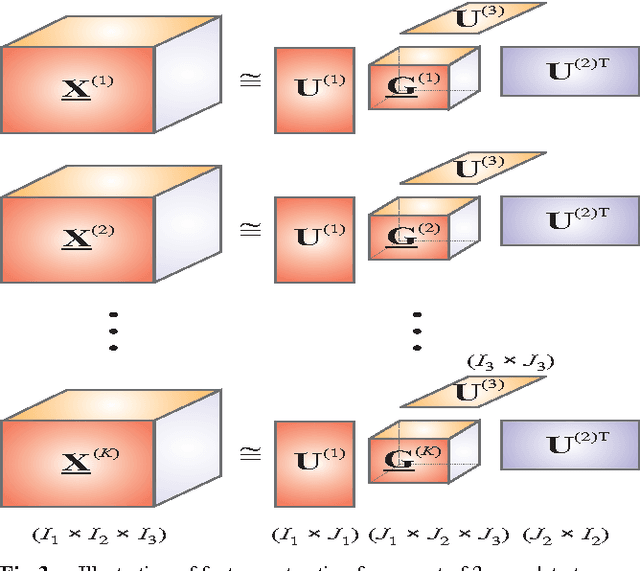
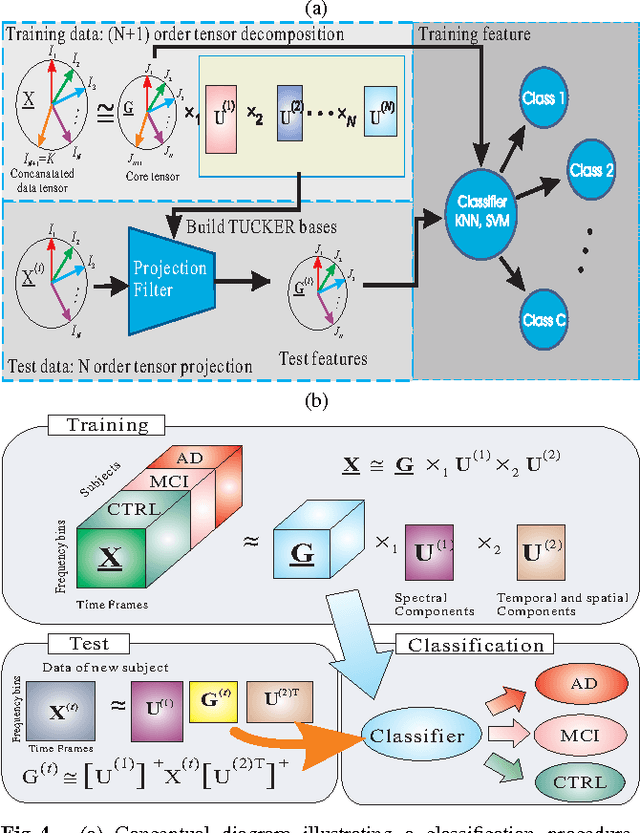
Matrix factorizations and their extensions to tensor factorizations and decompositions have become prominent techniques for linear and multilinear blind source separation (BSS), especially multiway Independent Component Analysis (ICA), NonnegativeMatrix and Tensor Factorization (NMF/NTF), Smooth Component Analysis (SmoCA) and Sparse Component Analysis (SCA). Moreover, tensor decompositions have many other potential applications beyond multilinear BSS, especially feature extraction, classification, dimensionality reduction and multiway clustering. In this paper, we briefly overview new and emerging models and approaches for tensor decompositions in applications to group and linked multiway BSS/ICA, feature extraction, classification andMultiway Partial Least Squares (MPLS) regression problems. Keywords: Multilinear BSS, linked multiway BSS/ICA, tensor factorizations and decompositions, constrained Tucker and CP models, Penalized Tensor Decompositions (PTD), feature extraction, classification, multiway PLS and CCA.
Low Complexity Damped Gauss-Newton Algorithms for CANDECOMP/PARAFAC
Sep 13, 2012Anh Huy Phan, Petr Tichavský, Andrzej Cichocki
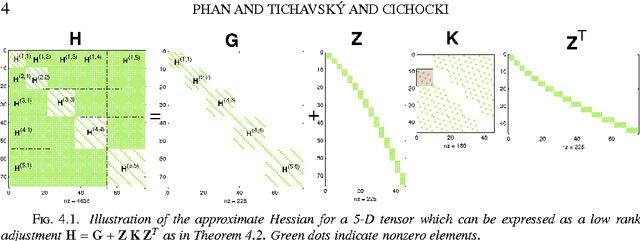
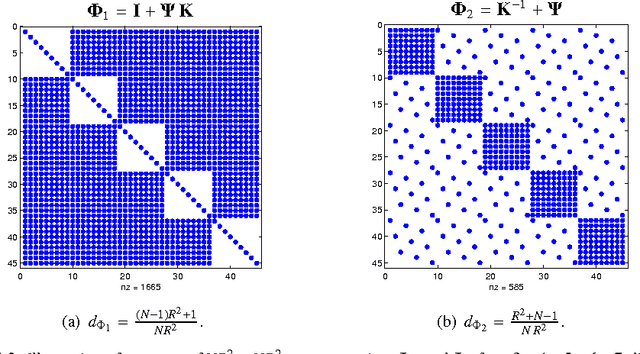
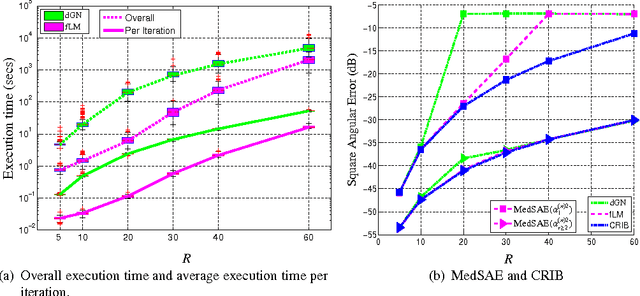
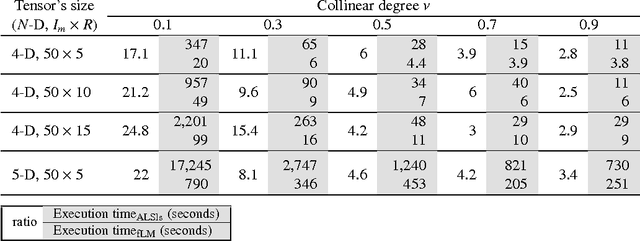
The damped Gauss-Newton (dGN) algorithm for CANDECOMP/PARAFAC (CP) decomposition can handle the challenges of collinearity of factors and different magnitudes of factors; nevertheless, for factorization of an $N$-D tensor of size $I_1\times I_N$ with rank $R$, the algorithm is computationally demanding due to construction of large approximate Hessian of size $(RT \times RT)$ and its inversion where $T = \sum_n I_n$. In this paper, we propose a fast implementation of the dGN algorithm which is based on novel expressions of the inverse approximate Hessian in block form. The new implementation has lower computational complexity, besides computation of the gradient (this part is common to both methods), requiring the inversion of a matrix of size $NR^2\times NR^2$, which is much smaller than the whole approximate Hessian, if $T \gg NR$. In addition, the implementation has lower memory requirements, because neither the Hessian nor its inverse never need to be stored in their entirety. A variant of the algorithm working with complex valued data is proposed as well. Complexity and performance of the proposed algorithm is compared with those of dGN and ALS with line search on examples of difficult benchmark tensors.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge