Andrea Bozoki
Topology-Aware Graph Augmentation for Predicting Clinical Trajectories in Neurocognitive Disorders
Oct 31, 2024



Abstract:Brain networks/graphs derived from resting-state functional MRI (fMRI) help study underlying pathophysiology of neurocognitive disorders by measuring neuronal activities in the brain. Some studies utilize learning-based methods for brain network analysis, but typically suffer from low model generalizability caused by scarce labeled fMRI data. As a notable self-supervised strategy, graph contrastive learning helps leverage auxiliary unlabeled data. But existing methods generally arbitrarily perturb graph nodes/edges to generate augmented graphs, without considering essential topology information of brain networks. To this end, we propose a topology-aware graph augmentation (TGA) framework, comprising a pretext model to train a generalizable encoder on large-scale unlabeled fMRI cohorts and a task-specific model to perform downstream tasks on a small target dataset. In the pretext model, we design two novel topology-aware graph augmentation strategies: (1) hub-preserving node dropping that prioritizes preserving brain hub regions according to node importance, and (2) weight-dependent edge removing that focuses on keeping important functional connectivities based on edge weights. Experiments on 1, 688 fMRI scans suggest that TGA outperforms several state-of-the-art methods.
Functional Imaging Constrained Diffusion for Brain PET Synthesis from Structural MRI
May 03, 2024



Abstract:Magnetic resonance imaging (MRI) and positron emission tomography (PET) are increasingly used in multimodal analysis of neurodegenerative disorders. While MRI is broadly utilized in clinical settings, PET is less accessible. Many studies have attempted to use deep generative models to synthesize PET from MRI scans. However, they often suffer from unstable training and inadequately preserve brain functional information conveyed by PET. To this end, we propose a functional imaging constrained diffusion (FICD) framework for 3D brain PET image synthesis with paired structural MRI as input condition, through a new constrained diffusion model (CDM). The FICD introduces noise to PET and then progressively removes it with CDM, ensuring high output fidelity throughout a stable training phase. The CDM learns to predict denoised PET with a functional imaging constraint introduced to ensure voxel-wise alignment between each denoised PET and its ground truth. Quantitative and qualitative analyses conducted on 293 subjects with paired T1-weighted MRI and 18F-fluorodeoxyglucose (FDG)-PET scans suggest that FICD achieves superior performance in generating FDG-PET data compared to state-of-the-art methods. We further validate the effectiveness of the proposed FICD on data from a total of 1,262 subjects through three downstream tasks, with experimental results suggesting its utility and generalizability.
Source-Free Collaborative Domain Adaptation via Multi-Perspective Feature Enrichment for Functional MRI Analysis
Aug 24, 2023



Abstract:Resting-state functional MRI (rs-fMRI) is increasingly employed in multi-site research to aid neurological disorder analysis. Existing studies usually suffer from significant cross-site/domain data heterogeneity caused by site effects such as differences in scanners/protocols. Many methods have been proposed to reduce fMRI heterogeneity between source and target domains, heavily relying on the availability of source data. But acquiring source data is challenging due to privacy concerns and/or data storage burdens in multi-site studies. To this end, we design a source-free collaborative domain adaptation (SCDA) framework for fMRI analysis, where only a pretrained source model and unlabeled target data are accessible. Specifically, a multi-perspective feature enrichment method (MFE) is developed for target fMRI analysis, consisting of multiple collaborative branches to dynamically capture fMRI features of unlabeled target data from multiple views. Each branch has a data-feeding module, a spatiotemporal feature encoder, and a class predictor. A mutual-consistency constraint is designed to encourage pair-wise consistency of latent features of the same input generated from these branches for robust representation learning. To facilitate efficient cross-domain knowledge transfer without source data, we initialize MFE using parameters of a pretrained source model. We also introduce an unsupervised pretraining strategy using 3,806 unlabeled fMRIs from three large-scale auxiliary databases, aiming to obtain a general feature encoder. Experimental results on three public datasets and one private dataset demonstrate the efficacy of our method in cross-scanner and cross-study prediction tasks. The model pretrained on large-scale rs-fMRI data has been released to the public.
Attention-Guided Autoencoder for Automated Progression Prediction of Subjective Cognitive Decline with Structural MRI
Jun 24, 2022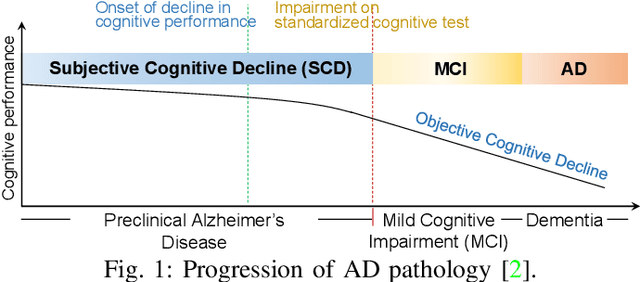
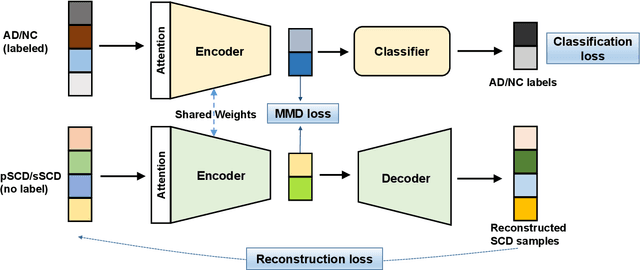
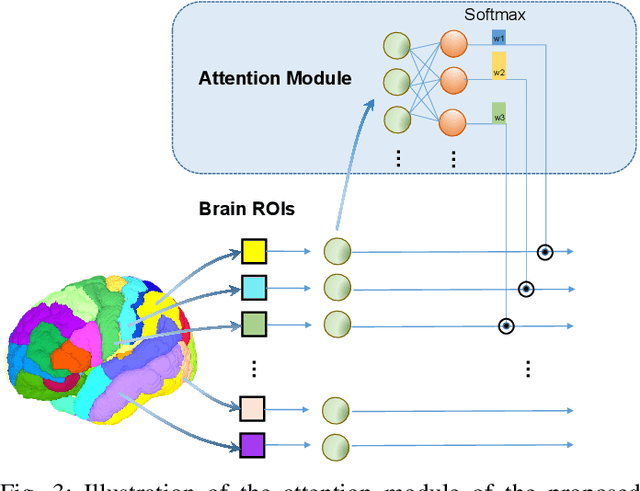
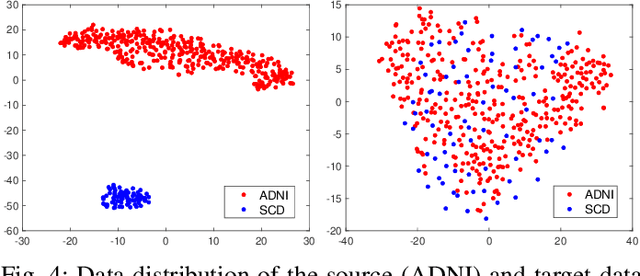
Abstract:Subjective cognitive decline (SCD) is a preclinical stage of Alzheimer's disease (AD) which occurs even before mild cognitive impairment (MCI). Progressive SCD will convert to MCI with the potential of further evolving to AD. Therefore, early identification of progressive SCD with neuroimaging techniques (e.g., structural MRI) is of great clinical value for early intervention of AD. However, existing MRI-based machine/deep learning methods usually suffer the small-sample-size problem which poses a great challenge to related neuroimaging analysis. The central question we aim to tackle in this paper is how to leverage related domains (e.g., AD/NC) to assist the progression prediction of SCD. Meanwhile, we are concerned about which brain areas are more closely linked to the identification of progressive SCD. To this end, we propose an attention-guided autoencoder model for efficient cross-domain adaptation which facilitates the knowledge transfer from AD to SCD. The proposed model is composed of four key components: 1) a feature encoding module for learning shared subspace representations of different domains, 2) an attention module for automatically locating discriminative brain regions of interest defined in brain atlases, 3) a decoding module for reconstructing the original input, 4) a classification module for identification of brain diseases. Through joint training of these four modules, domain invariant features can be learned. Meanwhile, the brain disease related regions can be highlighted by the attention mechanism. Extensive experiments on the publicly available ADNI dataset and a private CLAS dataset have demonstrated the effectiveness of the proposed method. The proposed model is straightforward to train and test with only 5-10 seconds on CPUs and is suitable for medical tasks with small datasets.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge