Ana G. Maguitman
Métodos para la Selección y el Ajuste de Características en el Problema de la Detección de Spam
Oct 14, 2010Abstract:The email is used daily by millions of people to communicate around the globe and it is a mission-critical application for many businesses. Over the last decade, unsolicited bulk email has become a major problem for email users. An overwhelming amount of spam is flowing into users' mailboxes daily. In 2004, an estimated 62% of all email was attributed to spam. Spam is not only frustrating for most email users, it strains the IT infrastructure of organizations and costs businesses billions of dollars in lost productivity. In recent years, spam has evolved from an annoyance into a serious security threat, and is now a prime medium for phishing of sensitive information, as well the spread of malicious software. This work presents a first approach to attack the spam problem. We propose an algorithm that will improve a classifier's results by adjusting its training set data. It improves the document's vocabulary representation by detecting good topic descriptors and discriminators.
* 5 pages, 1 figure, Workshop de Investigadores en Ciencias de la Computaci\'{o}n, WICC 2010, pp 48-52
Learning Better Context Characterizations: An Intelligent Information Retrieval Approach
Apr 27, 2010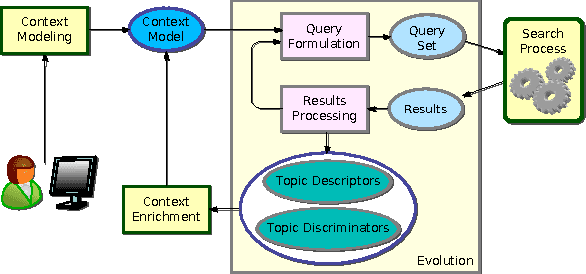

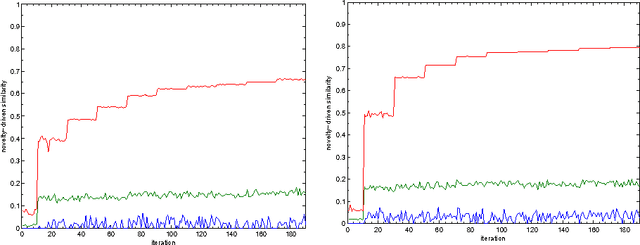
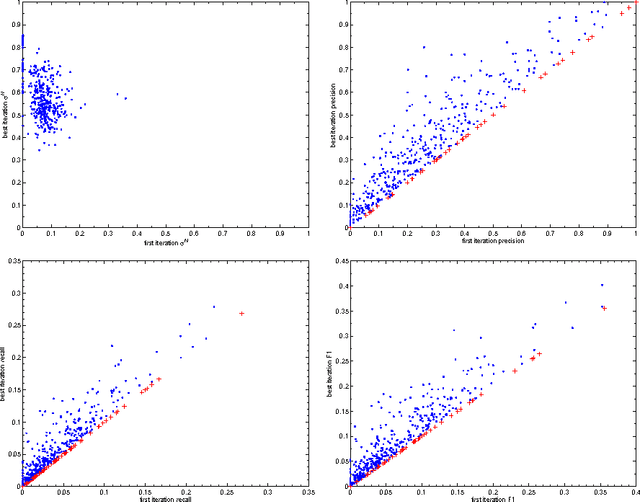
Abstract:This paper proposes an incremental method that can be used by an intelligent system to learn better descriptions of a thematic context. The method starts with a small number of terms selected from a simple description of the topic under analysis and uses this description as the initial search context. Using these terms, a set of queries are built and submitted to a search engine. New documents and terms are used to refine the learned vocabulary. Evaluations performed on a large number of topics indicate that the learned vocabulary is much more effective than the original one at the time of constructing queries to retrieve relevant material.
* 10 pages, 3 figures, CLEI 2008
Uncovering protein interaction in abstracts and text using a novel linear model and word proximity networks
Dec 04, 2008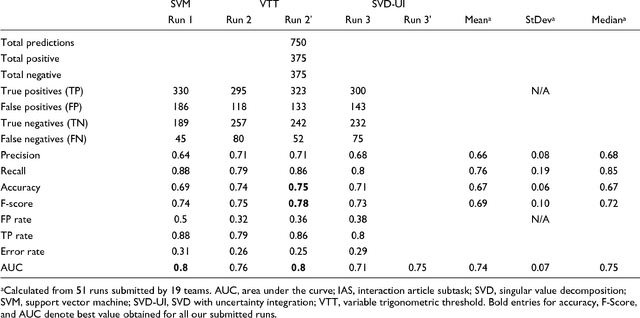
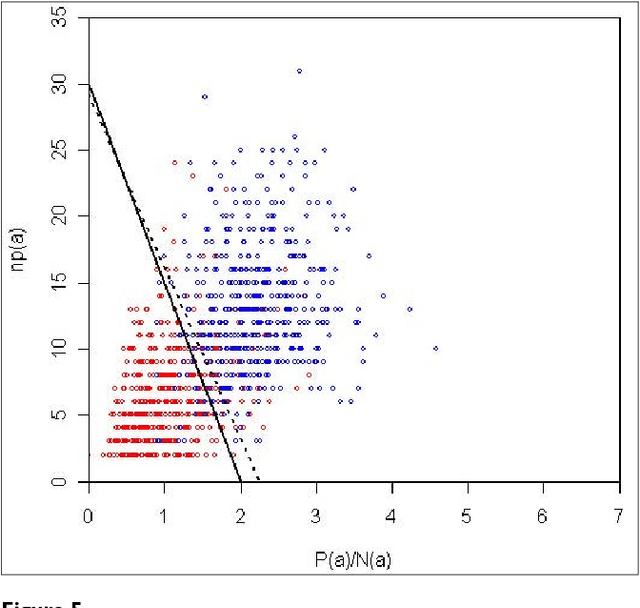
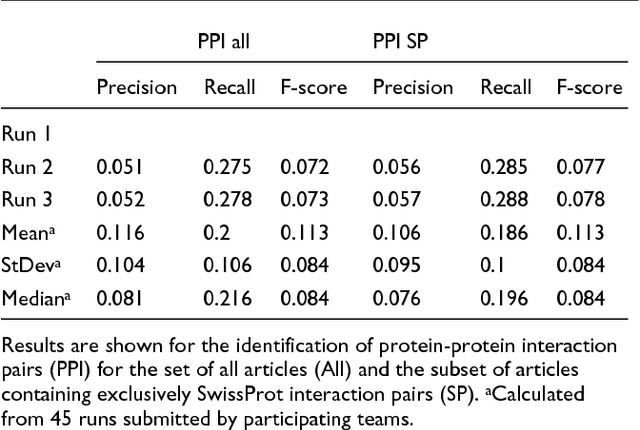
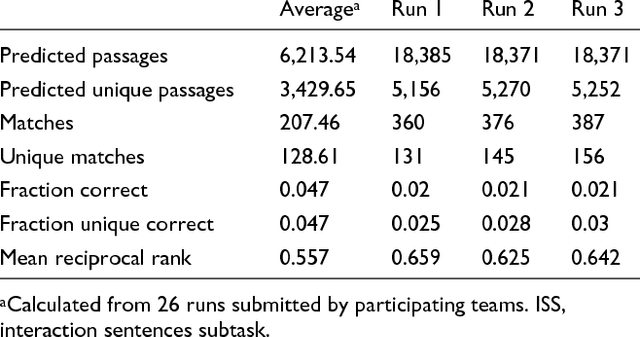
Abstract:We participated in three of the protein-protein interaction subtasks of the Second BioCreative Challenge: classification of abstracts relevant for protein-protein interaction (IAS), discovery of protein pairs (IPS) and text passages characterizing protein interaction (ISS) in full text documents. We approached the abstract classification task with a novel, lightweight linear model inspired by spam-detection techniques, as well as an uncertainty-based integration scheme. We also used a Support Vector Machine and the Singular Value Decomposition on the same features for comparison purposes. Our approach to the full text subtasks (protein pair and passage identification) includes a feature expansion method based on word-proximity networks. Our approach to the abstract classification task (IAS) was among the top submissions for this task in terms of the measures of performance used in the challenge evaluation (accuracy, F-score and AUC). We also report on a web-tool we produced using our approach: the Protein Interaction Abstract Relevance Evaluator (PIARE). Our approach to the full text tasks resulted in one of the highest recall rates as well as mean reciprocal rank of correct passages. Our approach to abstract classification shows that a simple linear model, using relatively few features, is capable of generalizing and uncovering the conceptual nature of protein-protein interaction from the bibliome. Since the novel approach is based on a very lightweight linear model, it can be easily ported and applied to similar problems. In full text problems, the expansion of word features with word-proximity networks is shown to be useful, though the need for some improvements is discussed.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge