Alexis Joly
ZENITH, LIRMM, UM
Revisiting Human-in-the-Loop Object Retrieval with Pre-Trained Vision Transformers
Apr 01, 2026Abstract:Building on existing approaches, we revisit Human-in-the-Loop Object Retrieval, a task that consists of iteratively retrieving images containing objects of a class-of-interest, specified by a user-provided query. Starting from a large unlabeled image collection, the aim is to rapidly identify diverse instances of an object category relying solely on the initial query and the user's Relevance Feedback, with no prior labels. The retrieval process is formulated as a binary classification task, where the system continuously learns to distinguish between relevant and non-relevant images to the query, through iterative user interaction. This interaction is guided by an Active Learning loop: at each iteration, the system selects informative samples for user annotation, thereby refining the retrieval performance. This task is particularly challenging in multi-object datasets, where the object of interest may occupy only a small region of the image within a complex, cluttered scene. Unlike object-centered settings where global descriptors often suffice, multi-object images require more adapted, localized descriptors. In this work, we formulate and revisit the Human-in-the-Loop Object Retrieval task by leveraging pre-trained ViT representations, and addressing key design questions, including which object instances to consider in an image, what form the annotations should take, how Active Selection should be applied, and which representation strategies best capture the object's features. We compare several representation strategies across multi-object datasets highlighting trade-offs between capturing the global context and focusing on fine-grained local object details. Our results offer practical insights for the design of effective interactive retrieval pipelines based on Active Learning for object class retrieval.
Positive-First Most Ambiguous: A Simple Active Learning Criterion for Interactive Retrieval of Rare Categories
Mar 25, 2026Abstract:Real-world fine-grained visual retrieval often requires discovering a rare concept from large unlabeled collections with minimal supervision. This is especially critical in biodiversity monitoring, ecological studies, and long-tailed visual domains, where the target may represent only a tiny fraction of the data, creating highly imbalanced binary problems. Interactive retrieval with relevance feedback offers a practical solution: starting from a small query, the system selects candidates for binary user annotation and iteratively refines a lightweight classifier. While Active Learning (AL) is commonly used to guide selection, conventional AL assumes symmetric class priors and large annotation budgets, limiting effectiveness in imbalanced, low-budget, low-latency settings. We introduce Positive-First Most Ambiguous (PF-MA), a simple yet effective AL criterion that explicitly addresses the class imbalance asymmetry: it prioritizes near-boundary samples while favoring likely positives, enabling rapid discovery of subtle visual categories while maintaining informativeness. Unlike standard methods that oversample negatives, PF-MA consistently returns small batches with a high proportion of relevant samples, improving early retrieval and user satisfaction. To capture retrieval diversity, we also propose a class coverage metric that measures how well selected positives span the visual variability of the target class. Experiments on long-tailed datasets, including fine-grained botanical data, demonstrate that PF-MA consistently outperforms strong baselines in both coverage and classifier performance, across varying class sizes and descriptors. Our results highlight that aligning AL with the asymmetric and user-centric objectives of interactive fine-grained retrieval enables simple yet powerful solutions for retrieving rare and visually subtle categories in realistic human-in-the-loop settings.
How to Optimize Multispecies Set Predictions in Presence-Absence Modeling ?
Feb 12, 2026Abstract:Species distribution models (SDMs) commonly produce probabilistic occurrence predictions that must be converted into binary presence-absence maps for ecological inference and conservation planning. However, this binarization step is typically heuristic and can substantially distort estimates of species prevalence and community composition. We present MaxExp, a decision-driven binarization framework that selects the most probable species assemblage by directly maximizing a chosen evaluation metric. MaxExp requires no calibration data and is flexible across several scores. We also introduce the Set Size Expectation (SSE) method, a computationally efficient alternative that predicts assemblages based on expected species richness. Using three case studies spanning diverse taxa, species counts, and performance metrics, we show that MaxExp consistently matches or surpasses widely used thresholding and calibration methods, especially under strong class imbalance and high rarity. SSE offers a simpler yet competitive option. Together, these methods provide robust, reproducible tools for multispecies SDM binarization.
Self-Supervised Learning as Discrete Communication
Feb 10, 2026Abstract:Most self-supervised learning (SSL) methods learn continuous visual representations by aligning different views of the same input, offering limited control over how information is structured across representation dimensions. In this work, we frame visual self-supervised learning as a discrete communication process between a teacher and a student network, where semantic information is transmitted through a fixed-capacity binary channel. Rather than aligning continuous features, the student predicts multi-label binary messages produced by the teacher. Discrete agreement is enforced through an element-wise binary cross-entropy objective, while a coding-rate regularization term encourages effective utilization of the constrained channel, promoting structured representations. We further show that periodically reinitializing the projection head strengthens this effect by encouraging embeddings that remain predictive across multiple discrete encodings. Extensive experiments demonstrate consistent improvements over continuous agreement baselines on image classification, retrieval, and dense visual prediction tasks, as well as under domain shift through self-supervised adaptation. Beyond backbone representations, we analyze the learned binary codes and show that they form a compact and informative discrete language, capturing semantic factors reusable across classes.
Audio-to-Image Bird Species Retrieval without Audio-Image Pairs via Text Distillation
Jan 31, 2026Abstract:Audio-to-image retrieval offers an interpretable alternative to audio-only classification for bioacoustic species recognition, but learning aligned audio-image representations is challenging due to the scarcity of paired audio-image data. We propose a simple and data-efficient approach that enables audio-to-image retrieval without any audio-image supervision. Our proposed method uses text as a semantic intermediary: we distill the text embedding space of a pretrained image-text model (BioCLIP-2), which encodes rich visual and taxonomic structure, into a pretrained audio-text model (BioLingual) by fine-tuning its audio encoder with a contrastive objective. This distillation transfers visually grounded semantics into the audio representation, inducing emergent alignment between audio and image embeddings without using images during training. We evaluate the resulting model on multiple bioacoustic benchmarks. The distilled audio encoder preserves audio discriminative power while substantially improving audio-text alignment on focal recordings and soundscape datasets. Most importantly, on the SSW60 benchmark, the proposed approach achieves strong audio-to-image retrieval performance exceeding baselines based on zero-shot model combinations or learned mappings between text embeddings, despite not training on paired audio-image data. These results demonstrate that indirect semantic transfer through text is sufficient to induce meaningful audio-image alignment, providing a practical solution for visually grounded species recognition in data-scarce bioacoustic settings.
Compact Hypercube Embeddings for Fast Text-based Wildlife Observation Retrieval
Jan 30, 2026Abstract:Large-scale biodiversity monitoring platforms increasingly rely on multimodal wildlife observations. While recent foundation models enable rich semantic representations across vision, audio, and language, retrieving relevant observations from massive archives remains challenging due to the computational cost of high-dimensional similarity search. In this work, we introduce compact hypercube embeddings for fast text-based wildlife observation retrieval, a framework that enables efficient text-based search over large-scale wildlife image and audio databases using compact binary representations. Building on the cross-view code alignment hashing framework, we extend lightweight hashing beyond a single-modality setup to align natural language descriptions with visual or acoustic observations in a shared Hamming space. Our approach leverages pretrained wildlife foundation models, including BioCLIP and BioLingual, and adapts them efficiently for hashing using parameter-efficient fine-tuning. We evaluate our method on large-scale benchmarks, including iNaturalist2024 for text-to-image retrieval and iNatSounds2024 for text-to-audio retrieval, as well as multiple soundscape datasets to assess robustness under domain shift. Results show that retrieval using discrete hypercube embeddings achieves competitive, and in several cases superior, performance compared to continuous embeddings, while drastically reducing memory and search cost. Moreover, we observe that the hashing objective consistently improves the underlying encoder representations, leading to stronger retrieval and zero-shot generalization. These results demonstrate that binary, language-based retrieval enables scalable and efficient search over large wildlife archives for biodiversity monitoring systems.
GeoPl@ntNet: A Platform for Exploring Essential Biodiversity Variables
Nov 16, 2025Abstract:This paper describes GeoPl@ntNet, an interactive web application designed to make Essential Biodiversity Variables accessible and understandable to everyone through dynamic maps and fact sheets. Its core purpose is to allow users to explore high-resolution AI-generated maps of species distributions, habitat types, and biodiversity indicators across Europe. These maps, developed through a cascading pipeline involving convolutional neural networks and large language models, provide an intuitive yet information-rich interface to better understand biodiversity, with resolutions as precise as 50x50 meters. The website also enables exploration of specific regions, allowing users to select areas of interest on the map (e.g., urban green spaces, protected areas, or riverbanks) to view local species and their coverage. Additionally, GeoPl@ntNet generates comprehensive reports for selected regions, including insights into the number of protected species, invasive species, and endemic species.
Image Hashing via Cross-View Code Alignment in the Age of Foundation Models
Nov 03, 2025Abstract:Efficient large-scale retrieval requires representations that are both compact and discriminative. Foundation models provide powerful visual and multimodal embeddings, but nearest neighbor search in these high-dimensional spaces is computationally expensive. Hashing offers an efficient alternative by enabling fast Hamming distance search with binary codes, yet existing approaches often rely on complex pipelines, multi-term objectives, designs specialized for a single learning paradigm, and long training times. We introduce CroVCA (Cross-View Code Alignment), a simple and unified principle for learning binary codes that remain consistent across semantically aligned views. A single binary cross-entropy loss enforces alignment, while coding-rate maximization serves as an anti-collapse regularizer to promote balanced and diverse codes. To implement this, we design HashCoder, a lightweight MLP hashing network with a final batch normalization layer to enforce balanced codes. HashCoder can be used as a probing head on frozen embeddings or to adapt encoders efficiently via LoRA fine-tuning. Across benchmarks, CroVCA achieves state-of-the-art results in just 5 training epochs. At 16 bits, it particularly well-for instance, unsupervised hashing on COCO completes in under 2 minutes and supervised hashing on ImageNet100 in about 3 minutes on a single GPU. These results highlight CroVCA's efficiency, adaptability, and broad applicability.
Unlocking Zero-Shot Plant Segmentation with Pl@ntNet Intelligence
Oct 14, 2025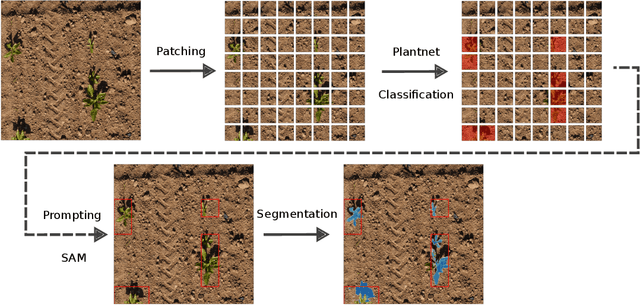
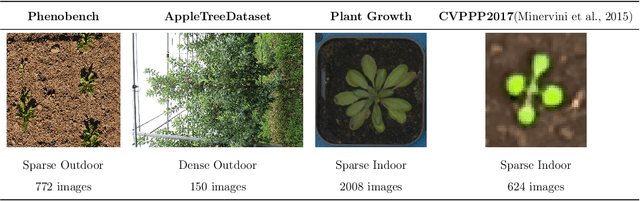

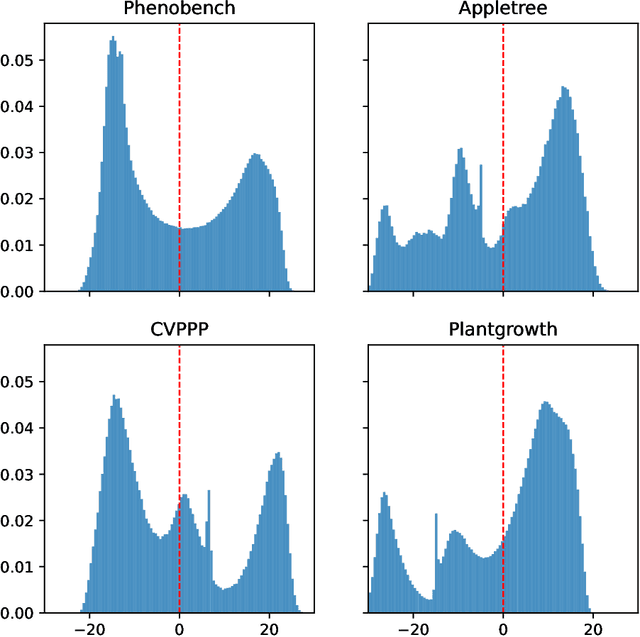
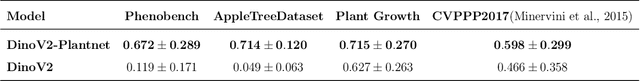
Abstract:We present a zero-shot segmentation approach for agricultural imagery that leverages Plantnet, a large-scale plant classification model, in conjunction with its DinoV2 backbone and the Segment Anything Model (SAM). Rather than collecting and annotating new datasets, our method exploits Plantnet's specialized plant representations to identify plant regions and produce coarse segmentation masks. These masks are then refined by SAM to yield detailed segmentations. We evaluate on four publicly available datasets of various complexity in terms of contrast including some where the limited size of the training data and complex field conditions often hinder purely supervised methods. Our results show consistent performance gains when using Plantnet-fine-tuned DinoV2 over the base DinoV2 model, as measured by the Jaccard Index (IoU). These findings highlight the potential of combining foundation models with specialized plant-centric models to alleviate the annotation bottleneck and enable effective segmentation in diverse agricultural scenarios.
Overview of PlantCLEF 2024: multi-species plant identification in vegetation plot images
Sep 19, 2025



Abstract:Plot images are essential for ecological studies, enabling standardized sampling, biodiversity assessment, long-term monitoring and remote, large-scale surveys. Plot images are typically fifty centimetres or one square meter in size, and botanists meticulously identify all the species found there. The integration of AI could significantly improve the efficiency of specialists, helping them to extend the scope and coverage of ecological studies. To evaluate advances in this regard, the PlantCLEF 2024 challenge leverages a new test set of thousands of multi-label images annotated by experts and covering over 800 species. In addition, it provides a large training set of 1.7 million individual plant images as well as state-of-the-art vision transformer models pre-trained on this data. The task is evaluated as a (weakly-labeled) multi-label classification task where the aim is to predict all the plant species present on a high-resolution plot image (using the single-label training data). In this paper, we provide an detailed description of the data, the evaluation methodology, the methods and models employed by the participants and the results achieved.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge