Alexander Kenneth Clarke
Unlocking the Full Potential of High-Density Surface EMG: Novel Non-Invasive High-Yield Motor Unit Decomposition
Oct 18, 2024Abstract:The decomposition of high-density surface electromyography (HD-sEMG) signals into motor unit discharge patterns has become a powerful tool for investigating the neural control of movement, providing insights into motor neuron recruitment and discharge behavior. However, current algorithms, while very effective under certain conditions, face significant challenges in complex scenarios, as their accuracy and motor unit yield are highly dependent on anatomical differences among individuals. This can limit the number of decomposed motor units, particularly in challenging conditions. To address this issue, we recently introduced Swarm-Contrastive Decomposition (SCD), which dynamically adjusts the separation function based on the distribution of the data and prevents convergence to the same source. Initially applied to intramuscular EMG signals, SCD is here adapted for HD-sEMG signals. We demonstrated its ability to address key challenges faced by existing methods, particularly in identifying low-amplitude motor unit action potentials and effectively handling complex decomposition scenarios, like high-interference signals. We extensively validated SCD using simulated and experimental HD-sEMG recordings and compared it with current state-of-the-art decomposition methods under varying conditions, including different excitation levels, noise intensities, force profiles, sexes, and muscle groups. The proposed method consistently outperformed existing techniques in both the quantity of decoded motor units and the precision of their firing time identification. For instance, under certain experimental conditions, SCD detected more than three times as many motor units compared to previous methods, while also significantly improving accuracy. These advancements represent a major step forward in non-invasive EMG technology for studying motor unit activity in complex scenarios.
HarmonICA: Neural non-stationarity correction and source separation for motor neuron interfaces
Jun 28, 2024Abstract:A major outstanding problem when interfacing with spinal motor neurons is how to accurately compensate for non-stationary effects in the signal during source separation routines, particularly when they cannot be estimated in advance. This forces current systems to instead use undifferentiated bulk signal, which limits the potential degrees of freedom for control. In this study we propose a potential solution, using an unsupervised learning algorithm to blindly correct for the effects of latent processes which drive the signal non-stationarities. We implement this methodology within the theoretical framework of a quasilinear version of independent component analysis (ICA). The proposed design, HarmonICA, sidesteps the identifiability problems of nonlinear ICA, allowing for equivalent predictability to linear ICA whilst retaining the ability to learn complex nonlinear relationships between non-stationary latents and their effects on the signal. We test HarmonICA on both invasive and non-invasive recordings both simulated and real, demonstrating an ability to blindly compensate for the non-stationary effects specific to each, and thus to significantly enhance the quality of a source separation routine.
Human Biophysics as Network Weights: Conditional Generative Models for Ultra-fast Simulation
Nov 03, 2022



Abstract:Simulations of biophysical systems have provided a huge contribution to our fundamental understanding of human physiology and remain a central pillar for developments in medical devices and human machine interfaces. However, despite their successes, such simulations usually rely on highly computationally expensive numerical modelling, which is often inefficient to adapt to new simulation parameters. This limits their use in dynamic models of human behavior, for example in modelling the electric fields generated by muscles in a moving arm. We propose the alternative approach to use conditional generative models, which can learn complex relationships between the underlying generative conditions whilst remaining inexpensive to sample from. As a demonstration of this concept, we present BioMime, a hybrid architecture that combines elements of deep latent variable models and conditional adversarial training to construct a generative model that can both transform existing data samples to reflect new modelling assumptions and sample new data from a conditioned distribution. We demonstrate that BioMime can learn to accurately mimic a complex numerical model of human muscle biophysics and then use this knowledge to continuously sample from a dynamically changing system in real-time. We argue that transfer learning approaches with conditional generative models are a viable solution for dynamic simulation with any numerical model.
Deep Metric Learning with Locality Sensitive Angular Loss for Self-Correcting Source Separation of Neural Spiking Signals
Oct 13, 2021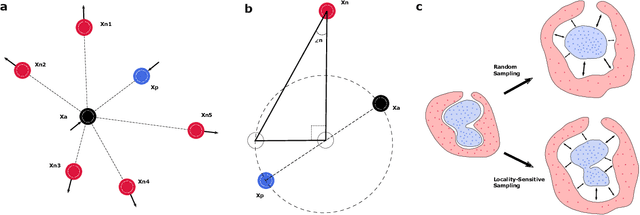
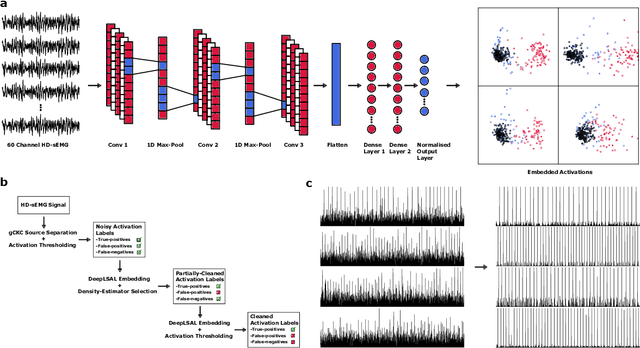
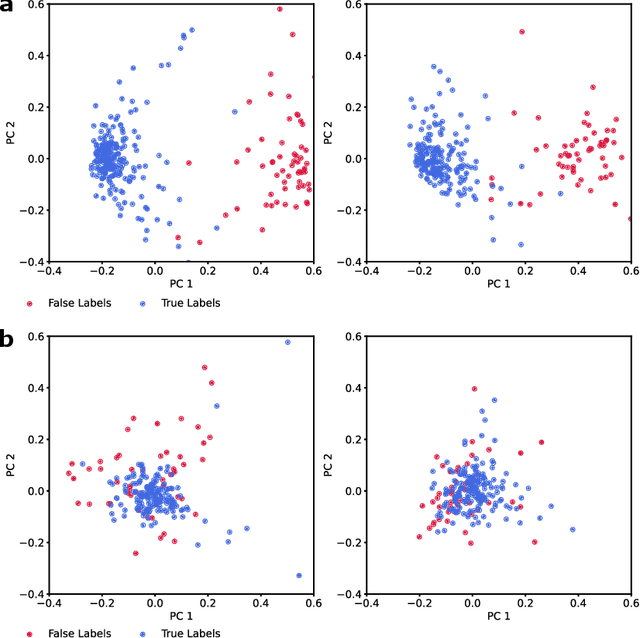
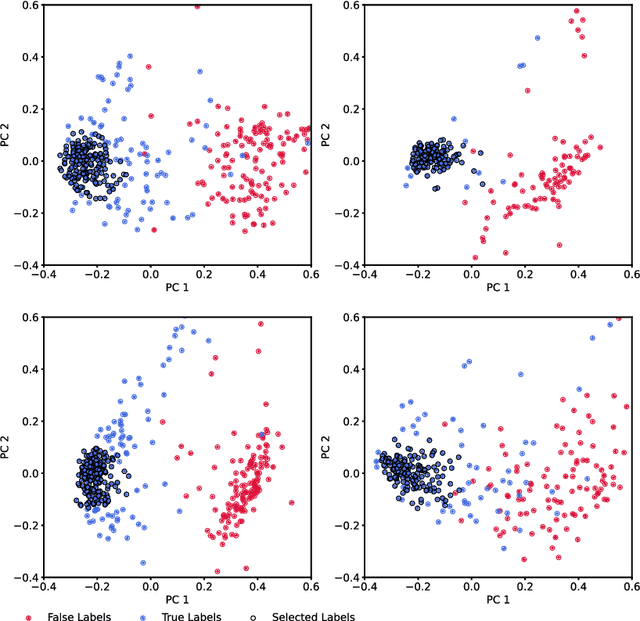
Abstract:Neurophysiological time series, such as electromyographic signal and intracortical recordings, are typically composed of many individual spiking sources, the recovery of which can give fundamental insights into the biological system of interest or provide neural information for man-machine interfaces. For this reason, source separation algorithms have become an increasingly important tool in neuroscience and neuroengineering. However, in noisy or highly multivariate recordings these decomposition techniques often make a large number of errors, which degrades human-machine interfacing applications and often requires costly post-hoc manual cleaning of the output label set of spike timestamps. To address both the need for automated post-hoc cleaning and robust separation filters we propose a methodology based on deep metric learning, using a novel loss function which maintains intra-class variance, creating a rich embedding space suitable for both label cleaning and the discovery of new activations. We then validate this method with an artificially corrupted label set based on source-separated high-density surface electromyography recordings, recovering the original timestamps even in extreme degrees of feature and class-dependent label noise. This approach enables a neural network to learn to accurately decode neurophysiological time series using any imperfect method of labelling the signal.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge