Alan R. Moody
Domain Adaptation using Silver Standard Masks for Lateral Ventricle Segmentation in FLAIR MRI
Jul 17, 2023



Abstract:Lateral ventricular volume (LVV) is an important biomarker for clinical investigation. We present the first transfer learning-based LVV segmentation method for fluid-attenuated inversion recovery (FLAIR) MRI. To mitigate covariate shifts between source and target domains, this work proposes an domain adaptation method that optimizes performance on three target datasets. Silver standard (SS) masks were generated from the target domain using a novel conventional image processing ventricular segmentation algorithm and used to supplement the gold standard (GS) data from the source domain, Canadian Atherosclerosis Imaging Network (CAIN). Four models were tested on held-out test sets from four datasets: 1) SS+GS: trained on target SS masks and fine-tuned on source GS masks, 2) GS+SS: trained on source GS masks and fine-tuned on target SS masks, 3) trained on source GS (GS CAIN Only) and 4) trained on target SS masks (SS Only). The SS+GS model had the best and most consistent performance (mean DSC = 0.89, CoV = 0.05) and showed significantly (p < 0.05) higher DSC compared to the GS-only model on three target domains. Results suggest pre-training with noisy labels from the target domain allows the model to adapt to the dataset-specific characteristics and provides robust parameter initialization while fine-tuning with GS masks allows the model to learn detailed features. This method has wide application to other medical imaging problems where labeled data is scarce, and can be used as a per-dataset calibration method to accelerate wide-scale adoption.
Effect of Intensity Standardization on Deep Learning for WML Segmentation in Multi-Centre FLAIR MRI
Jul 07, 2023Abstract:Deep learning (DL) methods for white matter lesion (WML) segmentation in MRI suffer a reduction in performance when applied on data from a scanner or centre that is out-of-distribution (OOD) from the training data. This is critical for translation and widescale adoption, since current models cannot be readily applied to data from new institutions. In this work, we evaluate several intensity standardization methods for MRI as a preprocessing step for WML segmentation in multi-centre Fluid-Attenuated Inversion Recovery (FLAIR) MRI. We evaluate a method specifically developed for FLAIR MRI called IAMLAB along with other popular normalization techniques such as White-strip, Nyul and Z-score. We proposed an Ensemble model that combines predictions from each of these models. A skip-connection UNet (SC UNet) was trained on the standardized images, as well as the original data and segmentation performance was evaluated over several dimensions. The training (in-distribution) data consists of a single study, of 60 volumes, and the test (OOD) data is 128 unseen volumes from three clinical cohorts. Results show IAMLAB and Ensemble provide higher WML segmentation performance compared to models from original data or other normalization methods. IAMLAB & Ensemble have the highest dice similarity coefficient (DSC) on the in-distribution data (0.78 & 0.80) and on clinical OOD data. DSC was significantly higher for IAMLAB compared to the original data (p<0.05) for all lesion categories (LL>25mL: 0.77 vs. 0.71; 10mL<= LL<25mL: 0.66 vs. 0.61; LL<10mL: 0.53 vs. 0.52). The IAMLAB and Ensemble normalization methods are mitigating MRI domain shift and are optimal for DL-based WML segmentation in unseen FLAIR data.
MLP-SRGAN: A Single-Dimension Super Resolution GAN using MLP-Mixer
Mar 11, 2023



Abstract:We propose a novel architecture called MLP-SRGAN, which is a single-dimension Super Resolution Generative Adversarial Network (SRGAN) that utilizes Multi-Layer Perceptron Mixers (MLP-Mixers) along with convolutional layers to upsample in the slice direction. MLP-SRGAN is trained and validated using high resolution (HR) FLAIR MRI from the MSSEG2 challenge dataset. The method was applied to three multicentre FLAIR datasets (CAIN, ADNI, CCNA) of images with low spatial resolution in the slice dimension to examine performance on held-out (unseen) clinical data. Upsampled results are compared to several state-of-the-art SR networks. For images with high resolution (HR) ground truths, peak-signal-to-noise-ratio (PSNR) and structural similarity index (SSIM) are used to measure upsampling performance. Several new structural, no-reference image quality metrics were proposed to quantify sharpness (edge strength), noise (entropy), and blurriness (low frequency information) in the absence of ground truths. Results show MLP-SRGAN results in sharper edges, less blurring, preserves more texture and fine-anatomical detail, with fewer parameters, faster training/evaluation time, and smaller model size than existing methods. Code for MLP-SRGAN training and inference, data generators, models and no-reference image quality metrics will be available at https://github.com/IAMLAB-Ryerson/MLP-SRGAN.
Adapting to Unseen Vendor Domains for MRI Lesion Segmentation
Aug 14, 2021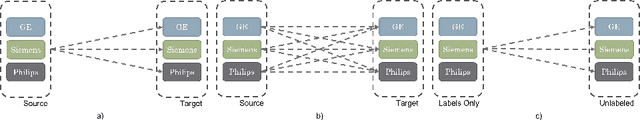
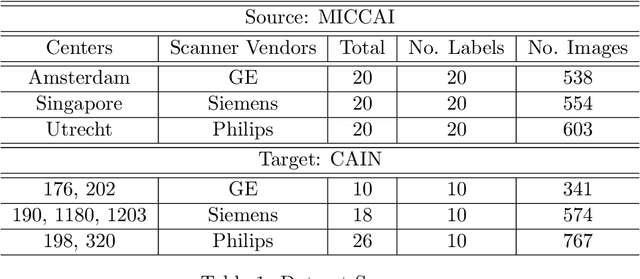
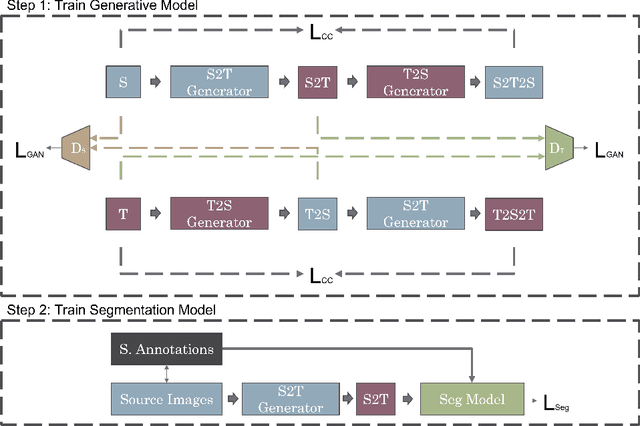
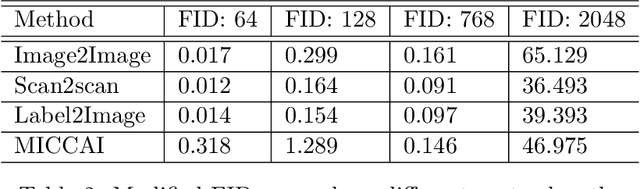
Abstract:One of the key limitations in machine learning models is poor performance on data that is out of the domain of the training distribution. This is especially true for image analysis in magnetic resonance (MR) imaging, as variations in hardware and software create non-standard intensities, contrasts, and noise distributions across scanners. Recently, image translation models have been proposed to augment data across domains to create synthetic data points. In this paper, we investigate the application an unsupervised image translation model to augment MR images from a source dataset to a target dataset. Specifically, we want to evaluate how well these models can create synthetic data points representative of the target dataset through image translation, and to see if a segmentation model trained these synthetic data points would approach the performance of a model trained directly on the target dataset. We consider three configurations of augmentation between datasets consisting of translation between images, between scanner vendors, and from labels to images. It was found that the segmentation models trained on synthetic data from labels to images configuration yielded the closest performance to the segmentation model trained directly on the target dataset. The Dice coeffcient score per each target vendor (GE, Siemens, Philips) for training on synthetic data was 0.63, 0.64, and 0.58, compared to training directly on target dataset was 0.65, 0.72, and 0.61.
FlowReg: Fast Deformable Unsupervised Medical Image Registration using Optical Flow
Jan 24, 2021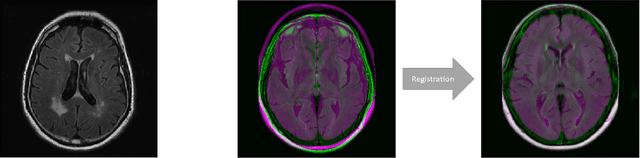
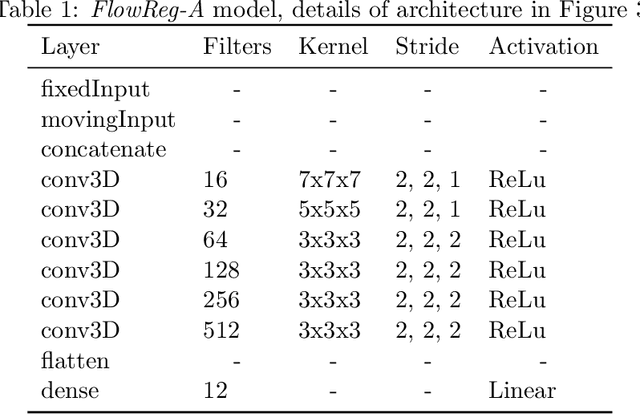
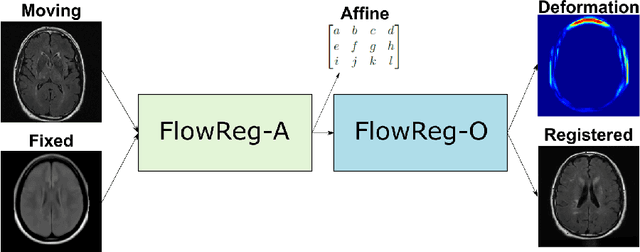
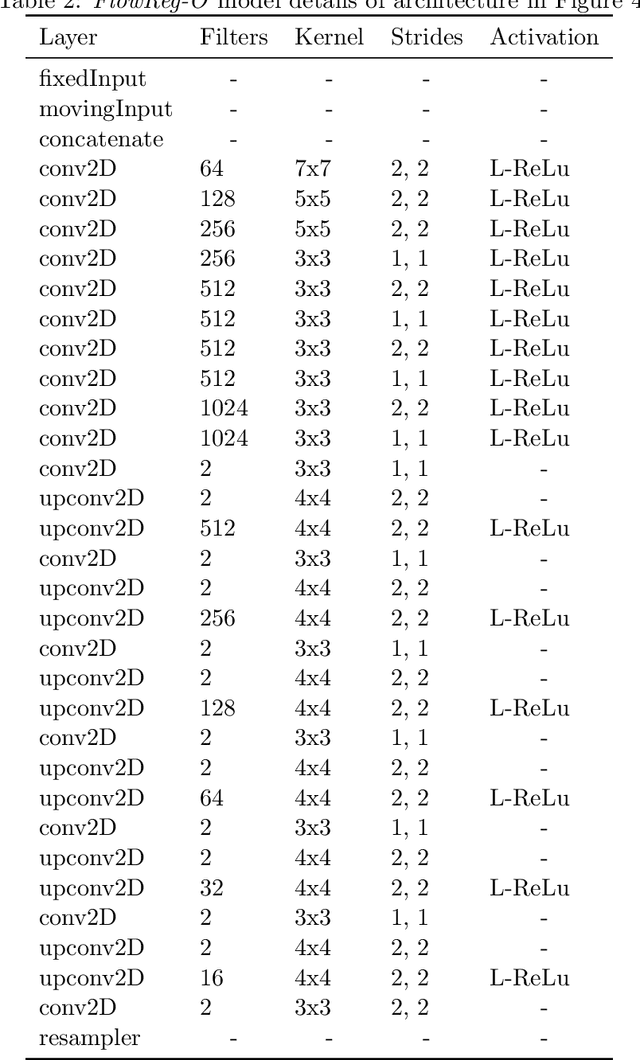
Abstract:We propose FlowReg, a deep learning-based framework for unsupervised image registration for neuroimaging applications. The system is composed of two architectures that are trained sequentially: FlowReg-A which affinely corrects for gross differences between moving and fixed volumes in 3D followed by FlowReg-O which performs pixel-wise deformations on a slice-by-slice basis for fine tuning in 2D. The affine network regresses the 3D affine matrix based on a correlation loss function that enforces global similarity. The deformable network operates on 2D image slices based on the optical flow network FlowNet-Simple but with three loss components. The photometric loss minimizes pixel intensity differences differences, the smoothness loss encourages similar magnitudes between neighbouring vectors, and a correlation loss that is used to maintain the intensity similarity between fixed and moving image slices. The proposed method is compared to four open source registration techniques ANTs, Demons, SE, and Voxelmorph. In total, 4643 FLAIR MR imaging volumes are used from dementia and vascular disease cohorts, acquired from over 60 international centres with varying acquisition parameters. A battery of quantitative novel registration validation metrics are proposed that focus on the structural integrity of tissues, spatial alignment, and intensity similarity. Experimental results show FlowReg (FlowReg-A+O) performs better than iterative-based registration algorithms for intensity and spatial alignment metrics with a Pixelwise Agreement of 0.65, correlation coefficient of 0.80, and Mutual Information of 0.29. Among the deep learning frameworks, FlowReg-A or FlowReg-A+O provided the highest performance over all but one of the metrics. Results show that FlowReg is able to obtain high intensity and spatial similarity while maintaining the shape and structure of anatomy and pathology.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge