Ziqi Ke
Real-Time Radio Technology and Modulation Classification via an LSTM Auto-Encoder
Nov 16, 2020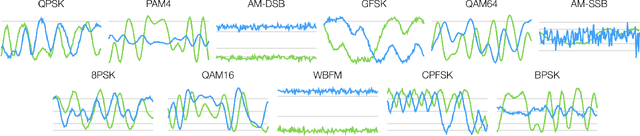
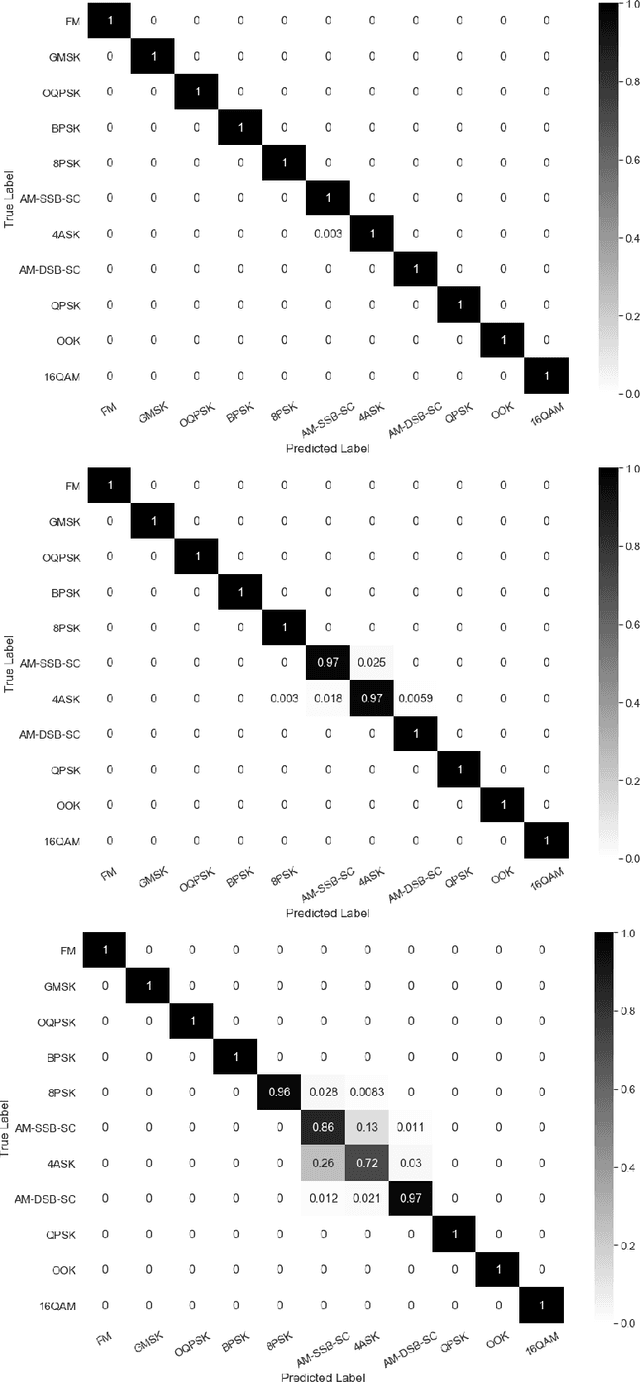
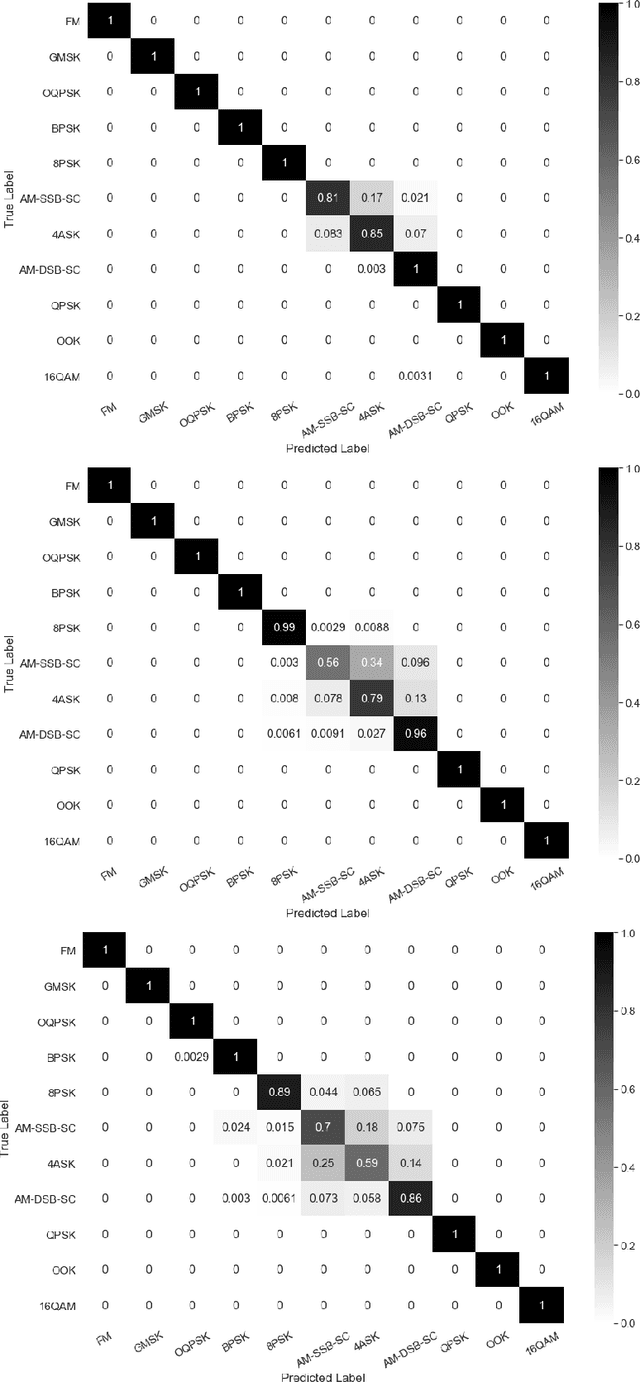
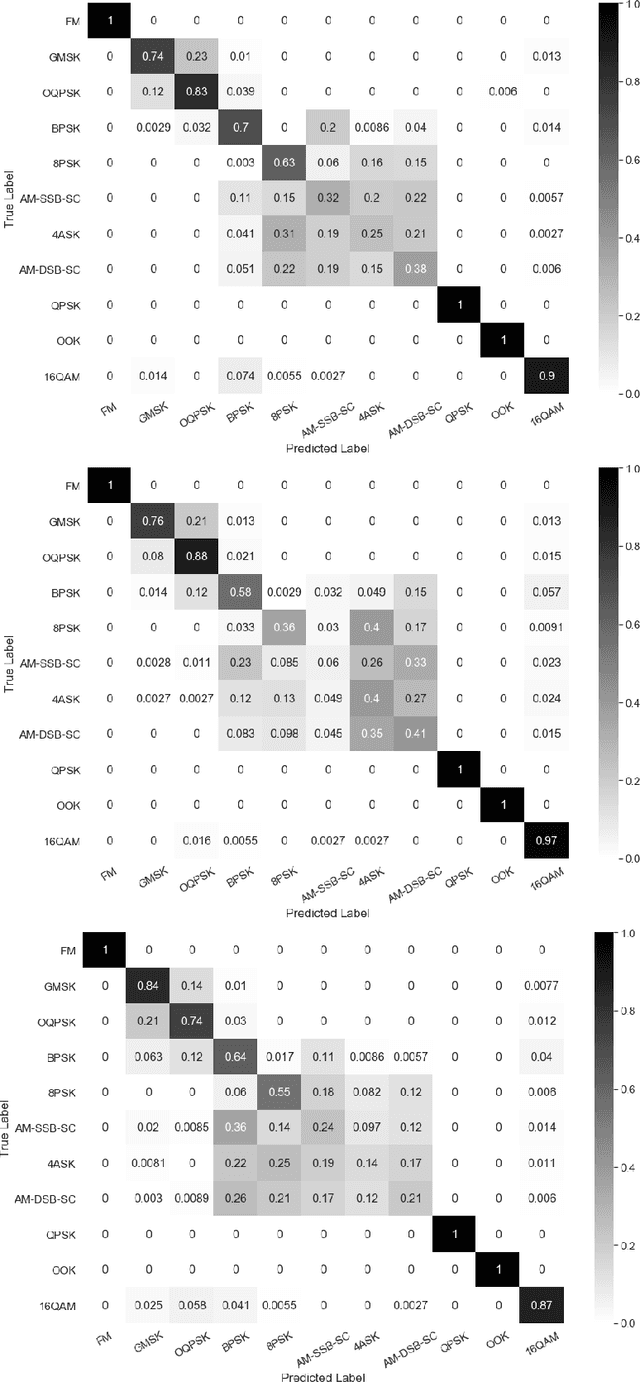
Abstract:Identification of the type of communication technology and/or modulation scheme based on detected radio signal are challenging problems encountered in a variety of applications including spectrum allocation and radio interference mitigation. They are rendered difficult due to a growing number of emitter types and varied effects of real-world channels upon the radio signal. Existing spectrum monitoring techniques are capable of acquiring massive amounts of radio and real-time spectrum data using compact sensors deployed in a variety of settings. However, state-of-the-art methods that use such data to classify emitter types and detect communication schemes struggle to achieve required levels of accuracy at a computational efficiency that would allow their implementation on low-cost computational platforms. In this paper, we present a learning framework based on an LSTM denoising auto-encoder designed to automatically extract stable and robust features from noisy radio signals, and infer modulation or technology type using the learned features. The algorithm utilizes a compact neural network architecture readily implemented on a low-cost computational platform while exceeding state-of-the-art accuracy. Results on realistic synthetic as well as over-the-air radio data demonstrate that the proposed framework reliably and efficiently classifies received radio signals, often demonstrating superior performance compared to state-of-the-art methods.
A Graph Auto-Encoder for Haplotype Assembly and Viral Quasispecies Reconstruction
Nov 13, 2019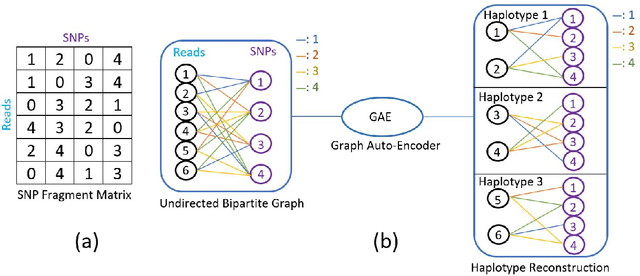
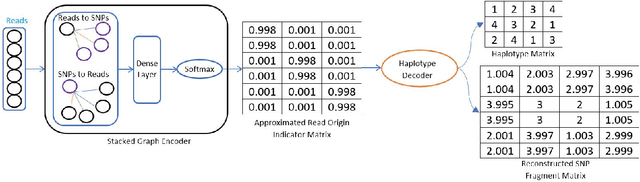
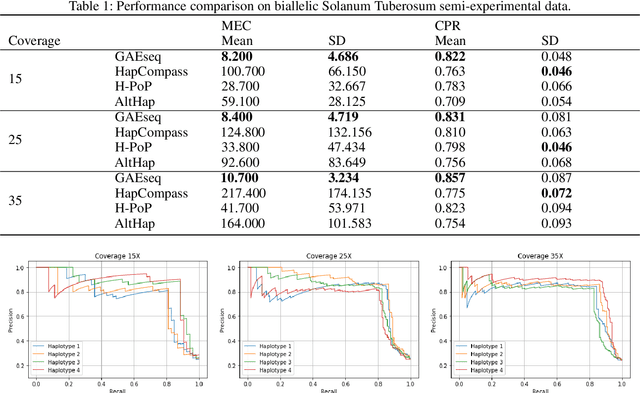

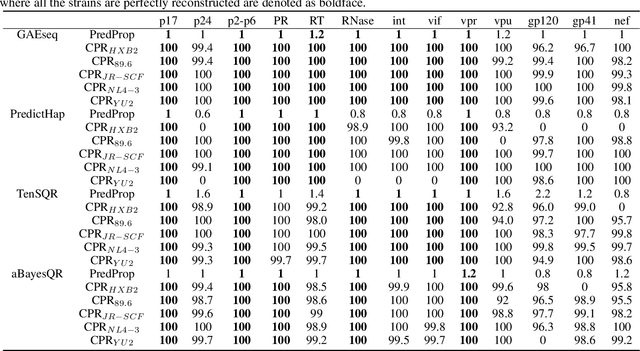
Abstract:Reconstructing components of a genomic mixture from data obtained by means of DNA sequencing is a challenging problem encountered in a variety of applications including single individual haplotyping and studies of viral communities. High-throughput DNA sequencing platforms oversample mixture components to provide massive amounts of reads whose relative positions can be determined by mapping the reads to a known reference genome; assembly of the components, however, requires discovery of the reads' origin -- an NP-hard problem that the existing methods struggle to solve with the required level of accuracy. In this paper, we present a learning framework based on a graph auto-encoder designed to exploit structural properties of sequencing data. The algorithm is a neural network which essentially trains to ignore sequencing errors and infers the posteriori probabilities of the origin of sequencing reads. Mixture components are then reconstructed by finding consensus of the reads determined to originate from the same genomic component. Results on realistic synthetic as well as experimental data demonstrate that the proposed framework reliably assembles haplotypes and reconstructs viral communities, often significantly outperforming state-of-the-art techniques.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge