Ziaurrehman Tanoli
A Survey of the Self Supervised Learning Mechanisms for Vision Transformers
Aug 30, 2024



Abstract:Deep supervised learning models require high volume of labeled data to attain sufficiently good results. Although, the practice of gathering and annotating such big data is costly and laborious. Recently, the application of self supervised learning (SSL) in vision tasks has gained significant attention. The intuition behind SSL is to exploit the synchronous relationships within the data as a form of self-supervision, which can be versatile. In the current big data era, most of the data is unlabeled, and the success of SSL thus relies in finding ways to improve this vast amount of unlabeled data available. Thus its better for deep learning algorithms to reduce reliance on human supervision and instead focus on self-supervision based on the inherent relationships within the data. With the advent of ViTs, which have achieved remarkable results in computer vision, it is crucial to explore and understand the various SSL mechanisms employed for training these models specifically in scenarios where there is less label data available. In this survey we thus develop a comprehensive taxonomy of systematically classifying the SSL techniques based upon their representations and pre-training tasks being applied. Additionally, we discuss the motivations behind SSL, review popular pre-training tasks, and highlight the challenges and advancements in this field. Furthermore, we present a comparative analysis of different SSL methods, evaluate their strengths and limitations, and identify potential avenues for future research.
R-BERT-CNN: Drug-target interactions extraction from biomedical literature
Oct 31, 2021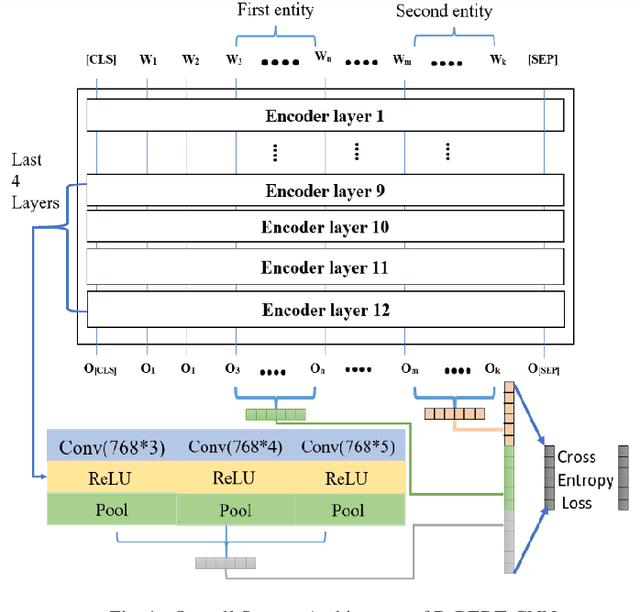
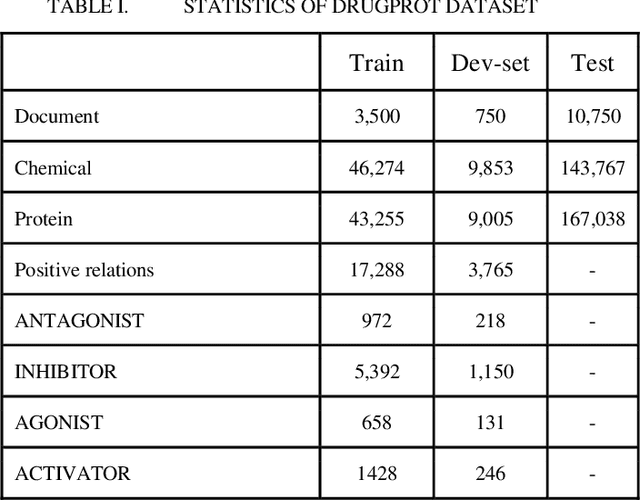
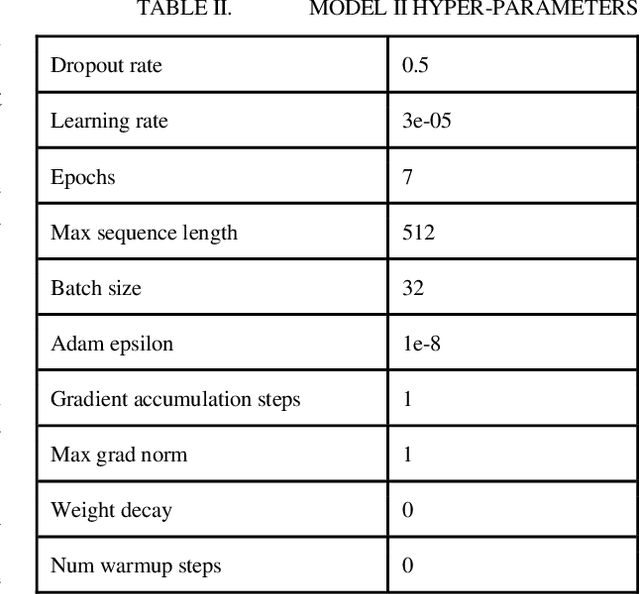
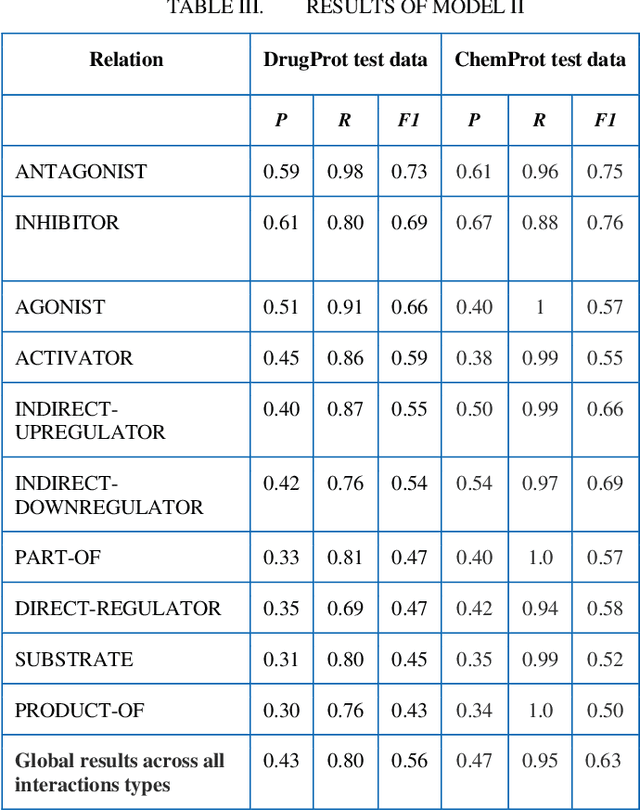
Abstract:In this research, we present our work participation for the DrugProt task of BioCreative VII challenge. Drug-target interactions (DTIs) are critical for drug discovery and repurposing, which are often manually extracted from the experimental articles. There are >32M biomedical articles on PubMed and manually extracting DTIs from such a huge knowledge base is challenging. To solve this issue, we provide a solution for Track 1, which aims to extract 10 types of interactions between drug and protein entities. We applied an Ensemble Classifier model that combines BioMed-RoBERTa, a state of art language model, with Convolutional Neural Networks (CNN) to extract these relations. Despite the class imbalances in the BioCreative VII DrugProt test corpus, our model achieves a good performance compared to the average of other submissions in the challenge, with the micro F1 score of 55.67% (and 63% on BioCreative VI ChemProt test corpus). The results show the potential of deep learning in extracting various types of DTIs.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge