Zhen Yuan
M100: An Orchestrated Dataflow Architecture Powering General AI Computing
Apr 20, 2026Abstract:As deep learning-based AI technologies gain momentum, the demand for general-purpose AI computing architectures continues to grow. While GPGPU-based architectures offer versatility for diverse AI workloads, they often fall short in efficiency and cost-effectiveness. Various Domain-Specific Architectures (DSAs) excel at particular AI tasks but struggle to extend across broader applications or adapt to the rapidly evolving AI landscape. M100 is Li Auto's response: a performant, cost-effective architecture for AI inference in Autonomous Driving (AD), Large Language Models (LLMs), and intelligent human interactions, domains crucial to today's most competitive automobile platforms. M100 employs a dataflow parallel architecture, where compiler-architecture co-design orchestrates not only computation but, more critically, data movement across time and space. Leveraging dataflow computing efficiency, our hardware-software co-design improves system performance while reducing hardware complexity and cost. M100 largely eliminates caching: tensor computations are driven by compiler- and runtime-managed data streams flowing between computing elements and on/off-chip memories, yielding greater efficiency and scalability than cache-based systems. Another key principle was selecting the right operational granularity for scheduling, issuing, and execution across compiler, firmware, and hardware. Recognizing commonalities in AI workloads, we chose the tensor as the fundamental data element. M100 demonstrates general AI computing capability across diverse inference applications, including UniAD (for AD) and LLaMA (for LLMs). Benchmarks show M100 outperforms GPGPU architectures in AD applications with higher utilization, representing a promising direction for future general AI computing.
MODIX: A Training-Free Multimodal Information-Driven Positional Index Scaling for Vision-Language Models
Apr 14, 2026Abstract:Vision-Language Models (VLMs) have achieved remarkable progress in multimodal understanding, yet their positional encoding mechanisms remain suboptimal. Existing approaches uniformly assign positional indices to all tokens, overlooking variations in information density within and across modalities, which leads to inefficient attention allocation where redundant visual regions dominate while informative content is underrepresented. We identify positional granularity as an implicit resource and propose MODIX (Multimodal Information-Driven Positional IndeX Scaling), a training-free framework that dynamically adapts positional strides based on modality-specific contributions. MODIX jointly models intra-modal density via covariance-based entropy and inter-modal interaction via cross-modal alignment to derive unified scores, which rescale positional indices to allocate finer granularity to informative modalities while compressing redundant ones, without requiring any modification to model parameters or architecture. Experiments across diverse architectures and benchmarks demonstrate that MODIX consistently improves multimodal reasoning and adaptively reallocates attention according to task-dependent information distributions, suggesting that positional encoding should be treated as an adaptive resource in Transformers for multimodal sequence modeling.
Deep Learning-Based Fetal Lung Segmentation from Diffusion-weighted MRI Images and Lung Maturity Evaluation for Fetal Growth Restriction
Jul 17, 2025Abstract:Fetal lung maturity is a critical indicator for predicting neonatal outcomes and the need for post-natal intervention, especially for pregnancies affected by fetal growth restriction. Intra-voxel incoherent motion analysis has shown promising results for non-invasive assessment of fetal lung development, but its reliance on manual segmentation is time-consuming, thus limiting its clinical applicability. In this work, we present an automated lung maturity evaluation pipeline for diffusion-weighted magnetic resonance images that consists of a deep learning-based fetal lung segmentation model and a model-fitting lung maturity assessment. A 3D nnU-Net model was trained on manually segmented images selected from the baseline frames of 4D diffusion-weighted MRI scans. The segmentation model demonstrated robust performance, yielding a mean Dice coefficient of 82.14%. Next, voxel-wise model fitting was performed based on both the nnU-Net-predicted and manual lung segmentations to quantify IVIM parameters reflecting tissue microstructure and perfusion. The results suggested no differences between the two. Our work shows that a fully automated pipeline is possible for supporting fetal lung maturity assessment and clinical decision-making.
DeepSPV: An Interpretable Deep Learning Pipeline for 3D Spleen Volume Estimation from 2D Ultrasound Images
Nov 17, 2024



Abstract:Splenomegaly, the enlargement of the spleen, is an important clinical indicator for various associated medical conditions, such as sickle cell disease (SCD). Spleen length measured from 2D ultrasound is the most widely used metric for characterising spleen size. However, it is still considered a surrogate measure, and spleen volume remains the gold standard for assessing spleen size. Accurate spleen volume measurement typically requires 3D imaging modalities, such as computed tomography or magnetic resonance imaging, but these are not widely available, especially in the Global South which has a high prevalence of SCD. In this work, we introduce a deep learning pipeline, DeepSPV, for precise spleen volume estimation from single or dual 2D ultrasound images. The pipeline involves a segmentation network and a variational autoencoder for learning low-dimensional representations from the estimated segmentations. We investigate three approaches for spleen volume estimation and our best model achieves 86.62%/92.5% mean relative volume accuracy (MRVA) under single-view/dual-view settings, surpassing the performance of human experts. In addition, the pipeline can provide confidence intervals for the volume estimates as well as offering benefits in terms of interpretability, which further support clinicians in decision-making when identifying splenomegaly. We evaluate the full pipeline using a highly realistic synthetic dataset generated by a diffusion model, achieving an overall MRVA of 83.0% from a single 2D ultrasound image. Our proposed DeepSPV is the first work to use deep learning to estimate 3D spleen volume from 2D ultrasound images and can be seamlessly integrated into the current clinical workflow for spleen assessment.
Deep Learning Framework for Spleen Volume Estimation from 2D Cross-sectional Views
Aug 17, 2023



Abstract:Abnormal spleen enlargement (splenomegaly) is regarded as a clinical indicator for a range of conditions, including liver disease, cancer and blood diseases. While spleen length measured from ultrasound images is a commonly used surrogate for spleen size, spleen volume remains the gold standard metric for assessing splenomegaly and the severity of related clinical conditions. Computed tomography is the main imaging modality for measuring spleen volume, but it is less accessible in areas where there is a high prevalence of splenomegaly (e.g., the Global South). Our objective was to enable automated spleen volume measurement from 2D cross-sectional segmentations, which can be obtained from ultrasound imaging. In this study, we describe a variational autoencoder-based framework to measure spleen volume from single- or dual-view 2D spleen segmentations. We propose and evaluate three volume estimation methods within this framework. We also demonstrate how 95% confidence intervals of volume estimates can be produced to make our method more clinically useful. Our best model achieved mean relative volume accuracies of 86.62% and 92.58% for single- and dual-view segmentations, respectively, surpassing the performance of the clinical standard approach of linear regression using manual measurements and a comparative deep learning-based 2D-3D reconstruction-based approach. The proposed spleen volume estimation framework can be integrated into standard clinical workflows which currently use 2D ultrasound images to measure spleen length. To the best of our knowledge, this is the first work to achieve direct 3D spleen volume estimation from 2D spleen segmentations.
Deep Learning for Automatic Spleen Length Measurement in Sickle Cell Disease Patients
Sep 06, 2020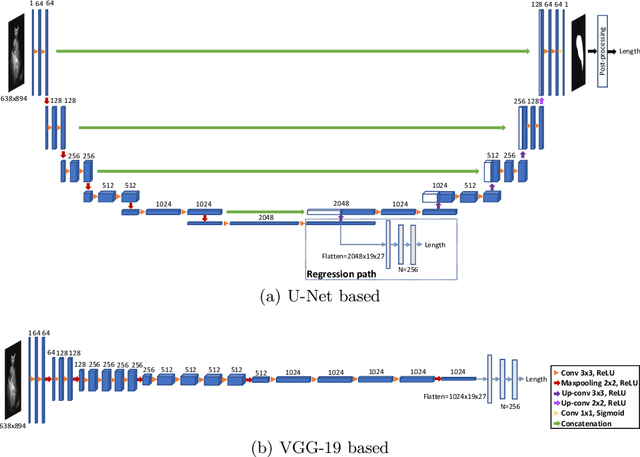
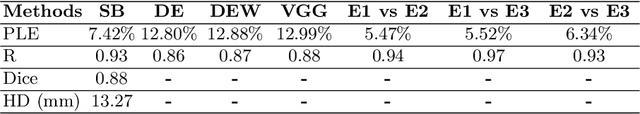
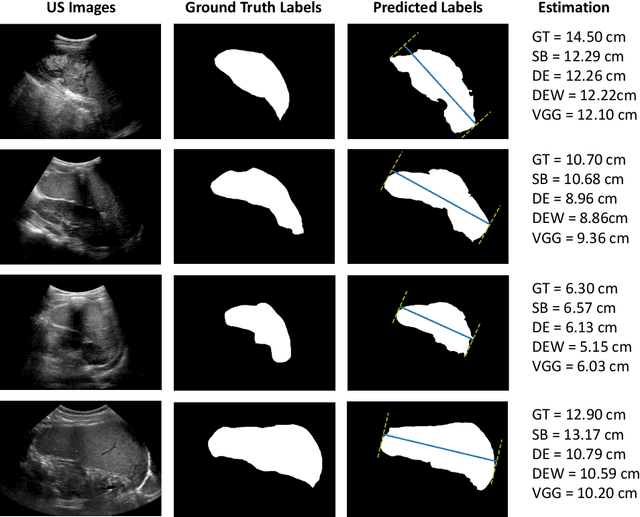
Abstract:Sickle Cell Disease (SCD) is one of the most common genetic diseases in the world. Splenomegaly (abnormal enlargement of the spleen) is frequent among children with SCD. If left untreated, splenomegaly can be life-threatening. The current workflow to measure spleen size includes palpation, possibly followed by manual length measurement in 2D ultrasound imaging. However, this manual measurement is dependent on operator expertise and is subject to intra- and inter-observer variability. We investigate the use of deep learning to perform automatic estimation of spleen length from ultrasound images. We investigate two types of approach, one segmentation-based and one based on direct length estimation, and compare the results against measurements made by human experts. Our best model (segmentation-based) achieved a percentage length error of 7.42%, which is approaching the level of inter-observer variability (5.47%-6.34%). To the best of our knowledge, this is the first attempt to measure spleen size in a fully automated way from ultrasound images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge