Xingjian Yan
Ensemble-Conditional Gaussian Processes (Ens-CGP): Representation, Geometry, and Inference
Feb 14, 2026Abstract:We formulate Ensemble-Conditional Gaussian Processes (Ens-CGP), a finite-dimensional synthesis that centers ensemble-based inference on the conditional Gaussian law. Conditional Gaussian processes (CGP) arise directly from Gaussian processes under conditioning and, in linear-Gaussian settings, define the full posterior distribution for a Gaussian prior and linear observations. Classical Kalman filtering is a recursive algorithm that computes this same conditional law under dynamical assumptions; the conditional Gaussian law itself is therefore the underlying representational object, while the filter is one computational realization. In this sense, CGP provides the probabilistic foundation for Kalman-type methods as well as equivalent formulations as a strictly convex quadratic program (MAP estimation), RKHS-regularized regression, and classical regularization. Ens-CGP is the ensemble instantiation of this object, obtained by treating empirical ensemble moments as a (possibly low-rank) Gaussian prior and performing exact conditioning. By separating representation (GP -> CGP -> Ens-CGP) from computation (Kalman filters, EnKF variants, and iterative ensemble schemes), the framework links an earlier-established representational foundation for inference to ensemble-derived priors and clarifies the relationships among probabilistic, variational, and ensemble perspectives.
Surrogate Supervision for Medical Image Analysis: Effective Deep Learning From Limited Quantities of Labeled Data
Jan 25, 2019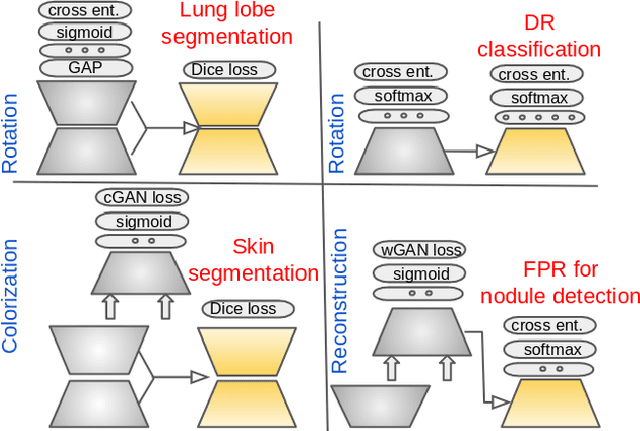

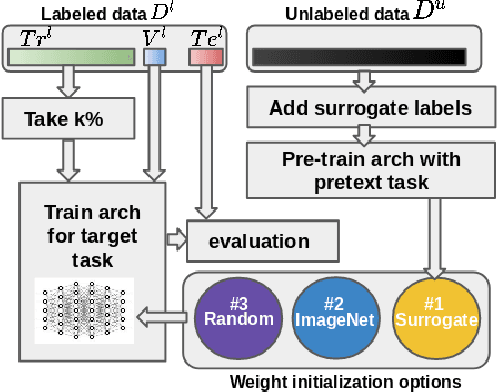
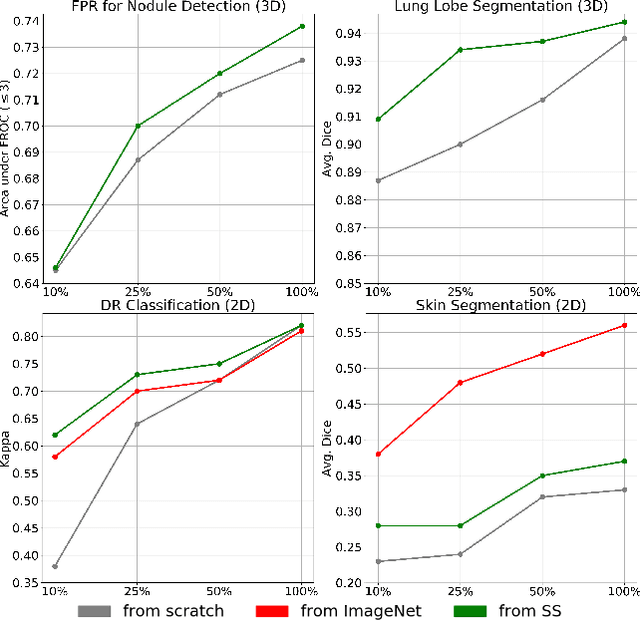
Abstract:We investigate the effectiveness of a simple solution to the common problem of deep learning in medical image analysis with limited quantities of labeled training data. The underlying idea is to assign artificial labels to abundantly available unlabeled medical images and, through a process known as surrogate supervision, pre-train a deep neural network model for the target medical image analysis task lacking sufficient labeled training data. In particular, we employ 3 surrogate supervision schemes, namely rotation, reconstruction, and colorization, in 4 different medical imaging applications representing classification and segmentation for both 2D and 3D medical images. 3 key findings emerge from our research: 1) pre-training with surrogate supervision is effective for small training sets; 2) deep models trained from initial weights pre-trained through surrogate supervision outperform the same models when trained from scratch, suggesting that pre-training with surrogate supervision should be considered prior to training any deep 3D models; 3) pre-training models in the medical domain with surrogate supervision is more effective than transfer learning from an unrelated domain (e.g., natural images), indicating the practical value of abundant unlabeled medical image data.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge