Won-Jin Moon
Unpaired Deep Learning for Pharmacokinetic Parameter Estimation from Dynamic Contrast-Enhanced MRI
Jun 07, 2023



Abstract:DCE-MRI provides information about vascular permeability and tissue perfusion through the acquisition of pharmacokinetic parameters. However, traditional methods for estimating these pharmacokinetic parameters involve fitting tracer kinetic models, which often suffer from computational complexity and low accuracy due to noisy arterial input function (AIF) measurements. Although some deep learning approaches have been proposed to tackle these challenges, most existing methods rely on supervised learning that requires paired input DCE-MRI and labeled pharmacokinetic parameter maps. This dependency on labeled data introduces significant time and resource constraints, as well as potential noise in the labels, making supervised learning methods often impractical. To address these limitations, here we present a novel unpaired deep learning method for estimating both pharmacokinetic parameters and the AIF using a physics-driven CycleGAN approach. Our proposed CycleGAN framework is designed based on the underlying physics model, resulting in a simpler architecture with a single generator and discriminator pair. Crucially, our experimental results indicate that our method, which does not necessitate separate AIF measurements, produces more reliable pharmacokinetic parameters than other techniques.
Which Contrast Does Matter? Towards a Deep Understanding of MR Contrast using Collaborative GAN
May 10, 2019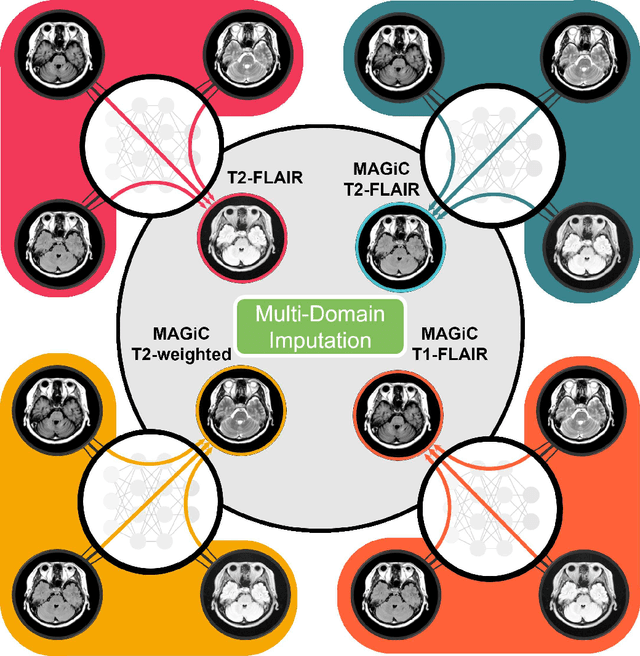
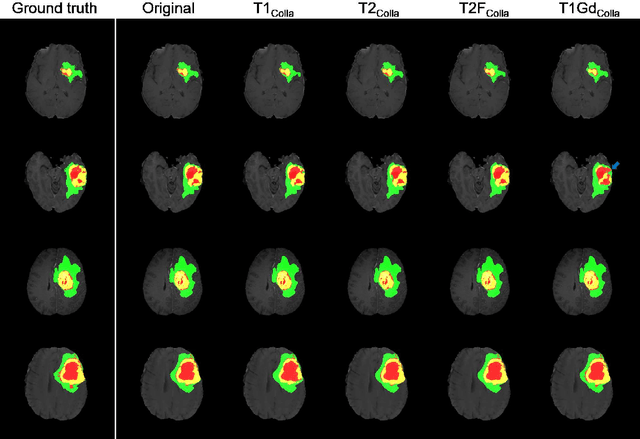
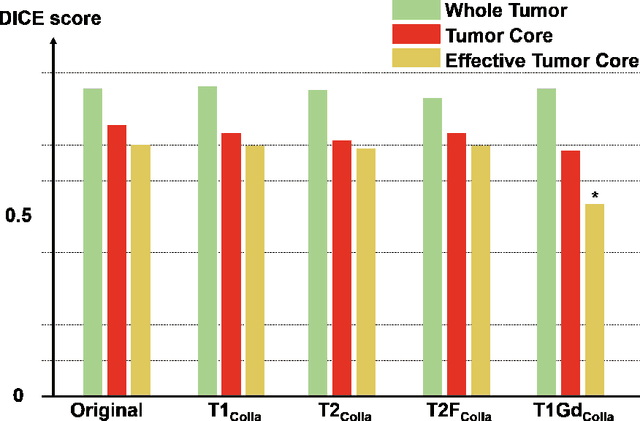
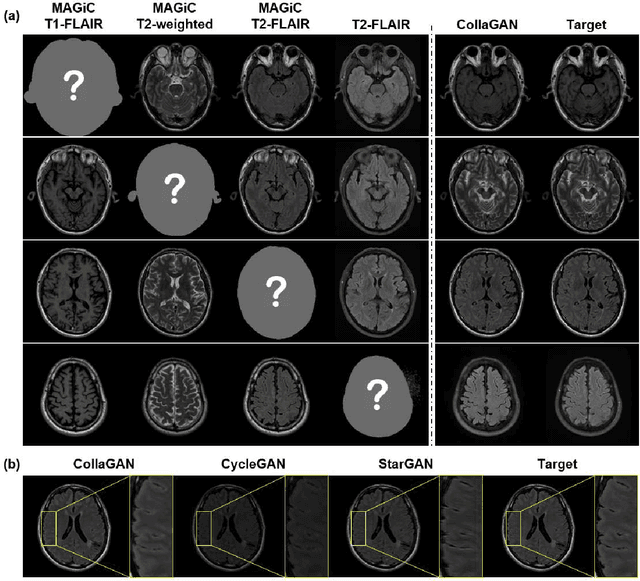
Abstract:Thanks to the recent success of generative adversarial network (GAN) for image synthesis, there are many exciting GAN approaches that successfully synthesize MR image contrast from other images with different contrasts. These approaches are potentially important for image imputation problems, where complete set of data is often difficult to obtain and image synthesis is one of the key solutions for handling the missing data problem. Unfortunately, the lack of the scalability of the existing GAN-based image translation approaches poses a fundamental challenge to understand the nature of the MR contrast imputation problem: which contrast does matter? Here, we present a systematic approach using Collaborative Generative Adversarial Networks (CollaGAN), which enable the learning of the joint image manifold of multiple MR contrasts to investigate which contrasts are essential. Our experimental results showed that the exogenous contrast from contrast agents is not replaceable, but other endogenous contrast such as T1, T2, etc can be synthesized from other contrast. These findings may give important guidance to the acquisition protocol design for MR in real clinical environment.
CollaGAN : Collaborative GAN for Missing Image Data Imputation
Jan 28, 2019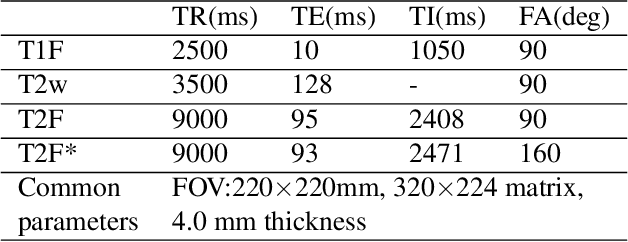
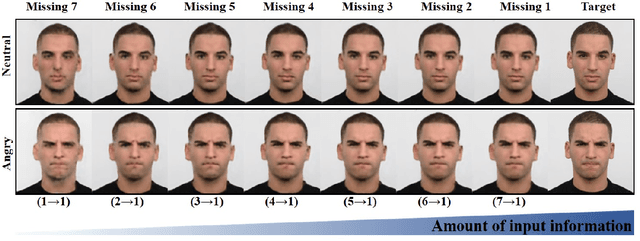


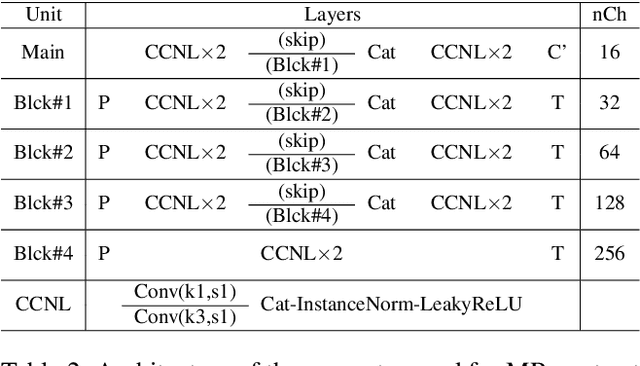
Abstract:In many applications requiring multiple inputs to obtain a desired output, if any of the input data is missing, it often introduces large amounts of bias. Although many techniques have been developed for imputing missing data, the image imputation is still difficult due to complicated nature of natural images. To address this problem, here we proposed a novel framework for missing image data imputation, called Collaborative Generative Adversarial Network (CollaGAN). CollaGAN converts an image imputation problem to a multi-domain images-to-image translation task so that a single generator and discriminator network can successfully estimate the missing data using the remaining clean data set. We demonstrate that CollaGAN produces the images with a higher visual quality compared to the existing competing approaches in various image imputation tasks.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge