William Klunk
Multi-Domain Learning by Meta-Learning: Taking Optimal Steps in Multi-Domain Loss Landscapes by Inner-Loop Learning
Feb 25, 2021
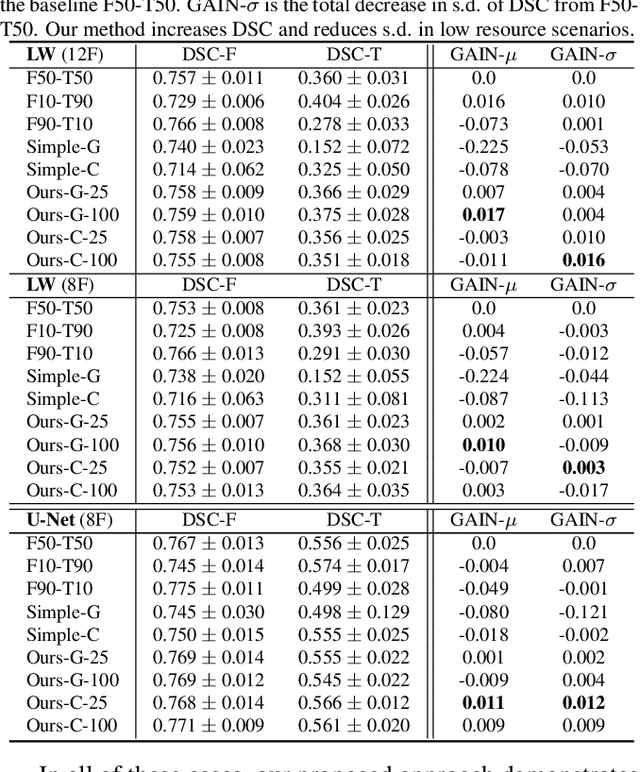
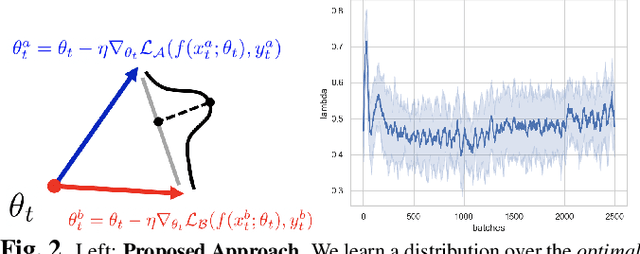
Abstract:We consider a model-agnostic solution to the problem of Multi-Domain Learning (MDL) for multi-modal applications. Many existing MDL techniques are model-dependent solutions which explicitly require nontrivial architectural changes to construct domain-specific modules. Thus, properly applying these MDL techniques for new problems with well-established models, e.g. U-Net for semantic segmentation, may demand various low-level implementation efforts. In this paper, given emerging multi-modal data (e.g., various structural neuroimaging modalities), we aim to enable MDL purely algorithmically so that widely used neural networks can trivially achieve MDL in a model-independent manner. To this end, we consider a weighted loss function and extend it to an effective procedure by employing techniques from the recently active area of learning-to-learn (meta-learning). Specifically, we take inner-loop gradient steps to dynamically estimate posterior distributions over the hyperparameters of our loss function. Thus, our method is model-agnostic, requiring no additional model parameters and no network architecture changes; instead, only a few efficient algorithmic modifications are needed to improve performance in MDL. We demonstrate our solution to a fitting problem in medical imaging, specifically, in the automatic segmentation of white matter hyperintensity (WMH). We look at two neuroimaging modalities (T1-MR and FLAIR) with complementary information fitting for our problem.
Robust White Matter Hyperintensity Segmentation on Unseen Domain
Feb 17, 2021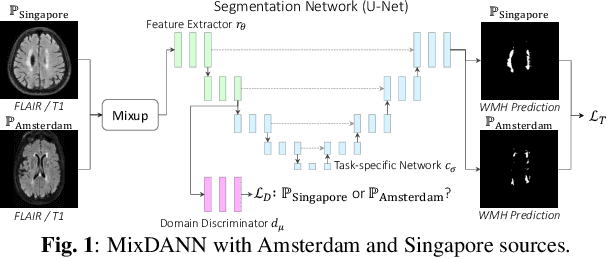
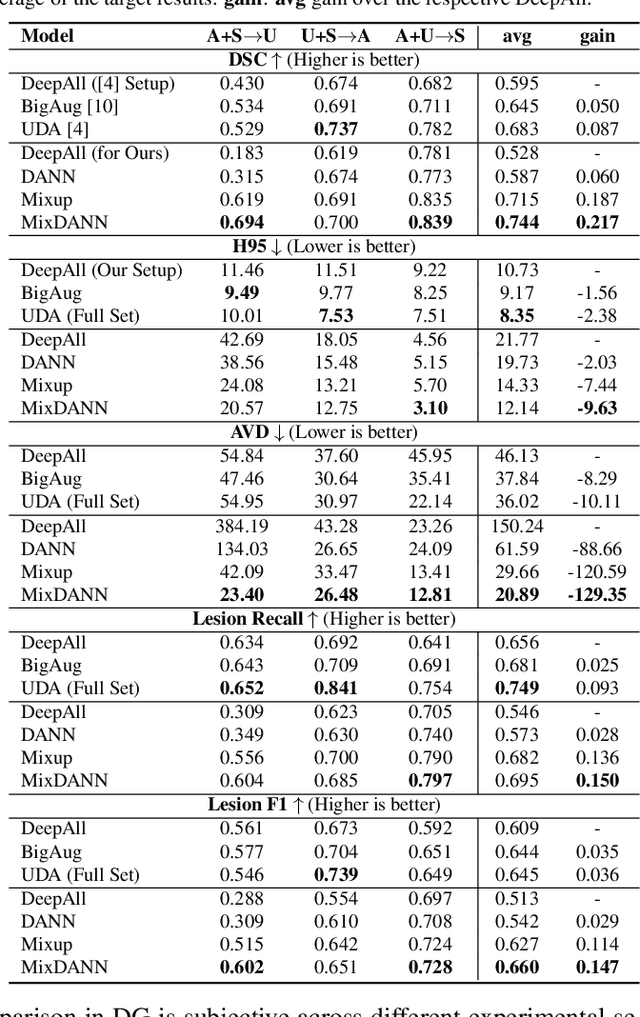
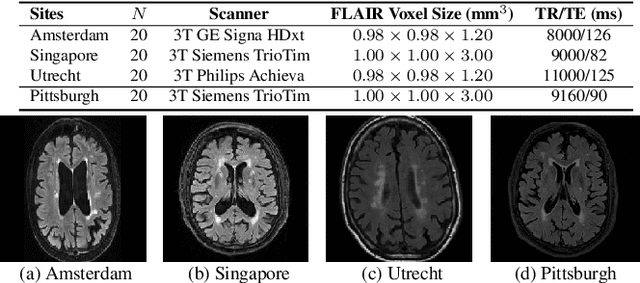
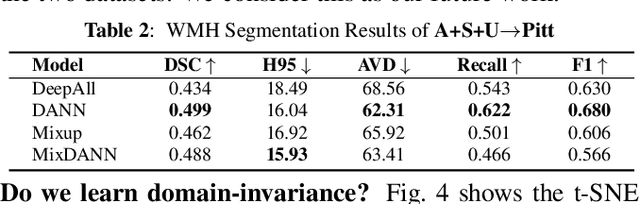
Abstract:Typical machine learning frameworks heavily rely on an underlying assumption that training and test data follow the same distribution. In medical imaging which increasingly begun acquiring datasets from multiple sites or scanners, this identical distribution assumption often fails to hold due to systematic variability induced by site or scanner dependent factors. Therefore, we cannot simply expect a model trained on a given dataset to consistently work well, or generalize, on a dataset from another distribution. In this work, we address this problem, investigating the application of machine learning models to unseen medical imaging data. Specifically, we consider the challenging case of Domain Generalization (DG) where we train a model without any knowledge about the testing distribution. That is, we train on samples from a set of distributions (sources) and test on samples from a new, unseen distribution (target). We focus on the task of white matter hyperintensity (WMH) prediction using the multi-site WMH Segmentation Challenge dataset and our local in-house dataset. We identify how two mechanically distinct DG approaches, namely domain adversarial learning and mix-up, have theoretical synergy. Then, we show drastic improvements of WMH prediction on an unseen target domain.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge