Wanda Kellum
Using Ensemble Models in the Histological Examination of Tissue Abnormalities
May 15, 2015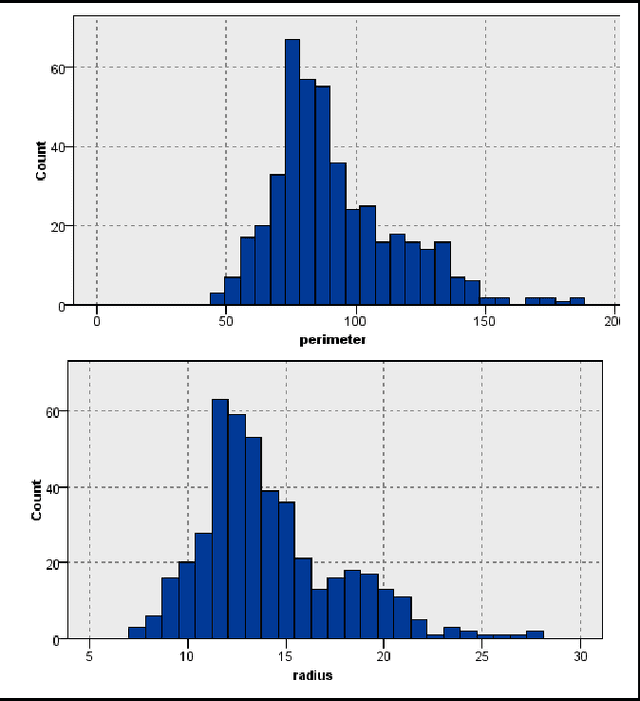
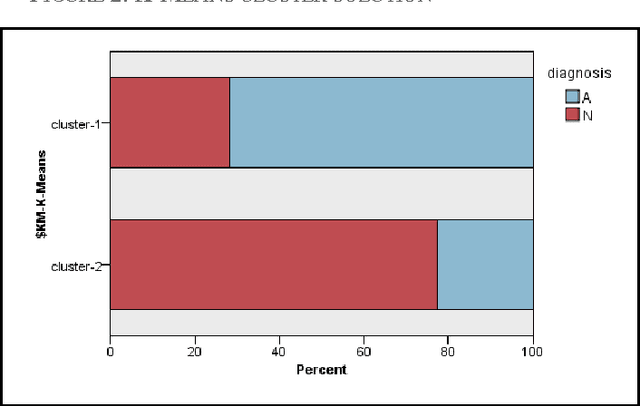
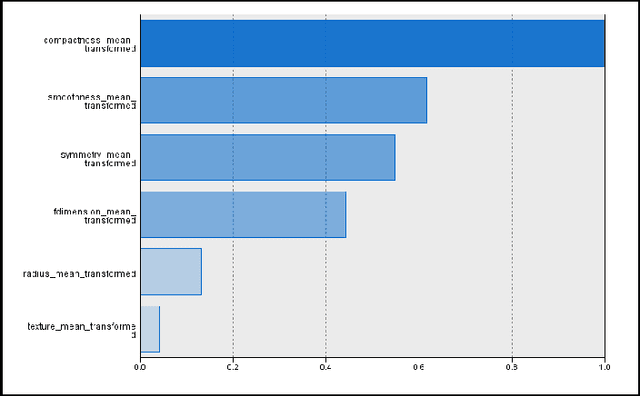
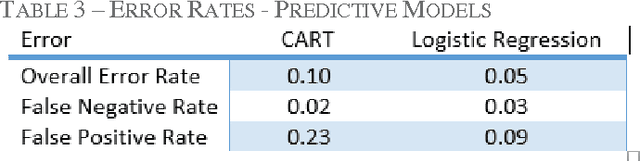
Abstract:Classification models for the automatic detection of abnormalities on histological samples do exists, with an active debate on the cost associated with false negative diagnosis (underdiagnosis) and false positive diagnosis (overdiagnosis). Current models tend to underdiagnose, failing to recognize a potentially fatal disease. The objective of this study is to investigate the possibility of automatically identifying abnormalities in tissue samples through the use of an ensemble model on data generated by histological examination and to minimize the number of false negative cases.
A Multivariate Biomarker for Parkinson's Disease
May 15, 2015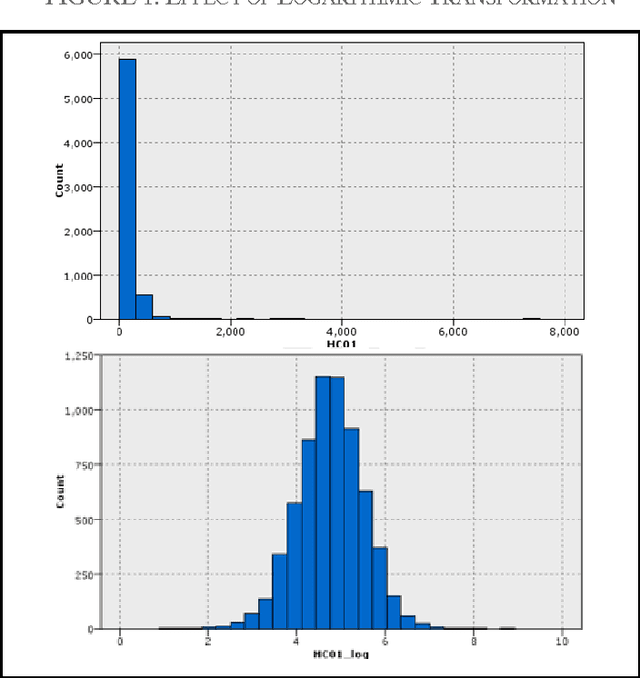
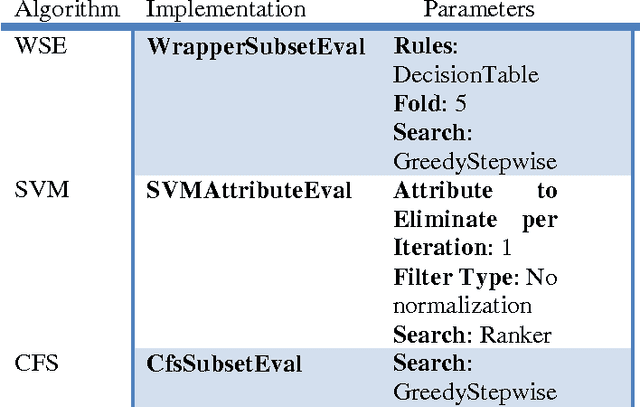
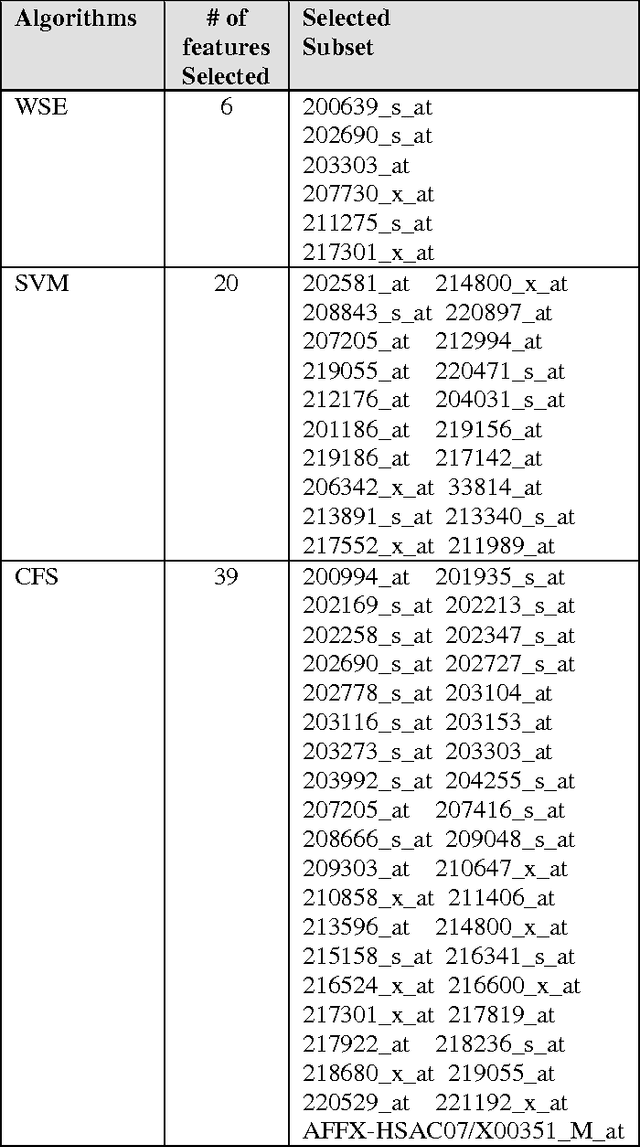
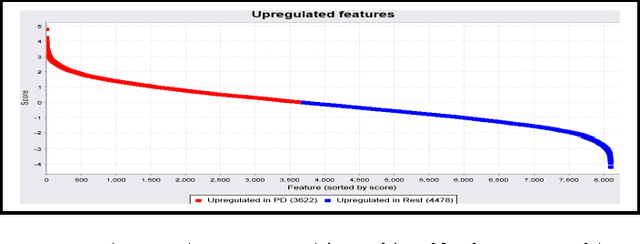

Abstract:In this study, we executed a genomic analysis with the objective of selecting a set of genes (possibly small) that would help in the detection and classification of samples from patients affected by Parkinson Disease. We performed a complete data analysis and during the exploratory phase, we selected a list of differentially expressed genes. Despite their association with the diseased state, we could not use them as a biomarker tool. Therefore, our research was extended to include a multivariate analysis approach resulting in the identification and selection of a group of 20 genes that showed a clear potential in detecting and correctly classify Parkinson Disease samples even in the presence of other neurodegenerative disorders.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge