Victor Chardès
Learning biophysical models of gene regulation with probability flow matching
Apr 27, 2026Abstract:Cellular differentiation is governed by gene regulatory networks, the high-dimensional stochastic biochemical systems that determine the transcriptional landscape and mediate cellular responses to signals and perturbations. Although single-cell RNA sequencing provides quantitative snapshots of the transcriptome, current methods for inferring gene-regulatory dynamics often lack mechanistic interpretability and fail to generalize to unseen conditions. Here we introduce Probability Flow Matching (PFM), a scalable framework for learning biophysically consistent stochastic processes directly from time-resolved single-cell measurements. Applying PFM to three hematopoiesis datasets, we show that models with similar interpolation accuracy can encode fundamentally different dynamics, with only biophysically consistent formulations accurately capturing mechanisms of lineage transitions, fate specification, and gene perturbation responses. We further demonstrate that PFM accommodates unbalanced populations, enabling simultaneous inference of cellular proliferation and death dynamics. Together, these results establish PFM as a flexible, scalable framework for integrating mechanistic modeling with single-cell omics.
Random Matrix Theory-guided sparse PCA for single-cell RNA-seq data
Sep 18, 2025Abstract:Single-cell RNA-seq provides detailed molecular snapshots of individual cells but is notoriously noisy. Variability stems from biological differences, PCR amplification bias, limited sequencing depth, and low capture efficiency, making it challenging to adapt computational pipelines to heterogeneous datasets or evolving technologies. As a result, most studies still rely on principal component analysis (PCA) for dimensionality reduction, valued for its interpretability and robustness. Here, we improve upon PCA with a Random Matrix Theory (RMT)-based approach that guides the inference of sparse principal components using existing sparse PCA algorithms. We first introduce a novel biwhitening method, inspired by the Sinkhorn-Knopp algorithm, that simultaneously stabilizes variance across genes and cells. This enables the use of an RMT-based criterion to automatically select the sparsity level, rendering sparse PCA nearly parameter-free. Our mathematically grounded approach retains the interpretability of PCA while enabling robust, hands-off inference of sparse principal components. Across seven single-cell RNA-seq technologies and four sparse PCA algorithms, we show that this method systematically improves the reconstruction of the principal subspace and consistently outperforms PCA-, autoencoder-, and diffusion-based methods in cell-type classification tasks.
Inferring stochastic dynamics with growth from cross-sectional data
May 19, 2025Abstract:Time-resolved single-cell omics data offers high-throughput, genome-wide measurements of cellular states, which are instrumental to reverse-engineer the processes underpinning cell fate. Such technologies are inherently destructive, allowing only cross-sectional measurements of the underlying stochastic dynamical system. Furthermore, cells may divide or die in addition to changing their molecular state. Collectively these present a major challenge to inferring realistic biophysical models. We present a novel approach, \emph{unbalanced} probability flow inference, that addresses this challenge for biological processes modelled as stochastic dynamics with growth. By leveraging a Lagrangian formulation of the Fokker-Planck equation, our method accurately disentangles drift from intrinsic noise and growth. We showcase the applicability of our approach through evaluation on a range of simulated and real single-cell RNA-seq datasets. Comparing to several existing methods, we find our method achieves higher accuracy while enjoying a simple two-step training scheme.
Inferring biological processes with intrinsic noise from cross-sectional data
Oct 10, 2024



Abstract:Inferring dynamical models from data continues to be a significant challenge in computational biology, especially given the stochastic nature of many biological processes. We explore a common scenario in omics, where statistically independent cross-sectional samples are available at a few time points, and the goal is to infer the underlying diffusion process that generated the data. Existing inference approaches often simplify or ignore noise intrinsic to the system, compromising accuracy for the sake of optimization ease. We circumvent this compromise by inferring the phase-space probability flow that shares the same time-dependent marginal distributions as the underlying stochastic process. Our approach, probability flow inference (PFI), disentangles force from intrinsic stochasticity while retaining the algorithmic ease of ODE inference. Analytically, we prove that for Ornstein-Uhlenbeck processes the regularized PFI formalism yields a unique solution in the limit of well-sampled distributions. In practical applications, we show that PFI enables accurate parameter and force estimation in high-dimensional stochastic reaction networks, and that it allows inference of cell differentiation dynamics with molecular noise, outperforming state-of-the-art approaches.
Stochastic force inference via density estimation
Oct 03, 2023



Abstract:Inferring dynamical models from low-resolution temporal data continues to be a significant challenge in biophysics, especially within transcriptomics, where separating molecular programs from noise remains an important open problem. We explore a common scenario in which we have access to an adequate amount of cross-sectional samples at a few time-points, and assume that our samples are generated from a latent diffusion process. We propose an approach that relies on the probability flow associated with an underlying diffusion process to infer an autonomous, nonlinear force field interpolating between the distributions. Given a prior on the noise model, we employ score-matching to differentiate the force field from the intrinsic noise. Using relevant biophysical examples, we demonstrate that our approach can extract non-conservative forces from non-stationary data, that it learns equilibrium dynamics when applied to steady-state data, and that it can do so with both additive and multiplicative noise models.
The Connection between Discrete- and Continuous-Time Descriptions of Gaussian Continuous Processes
Jan 20, 2021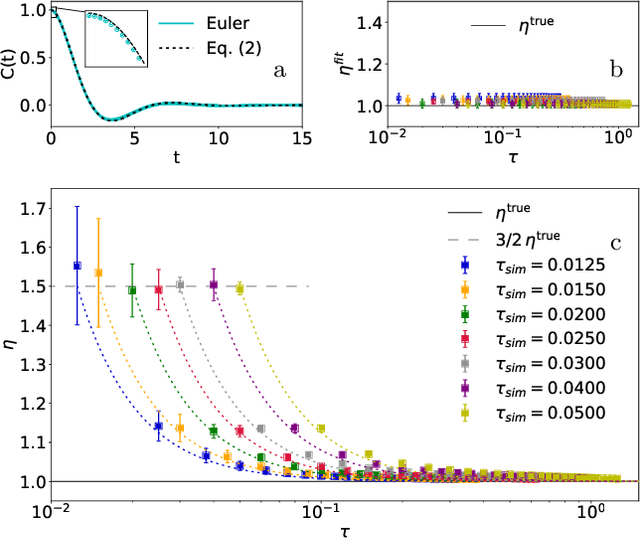
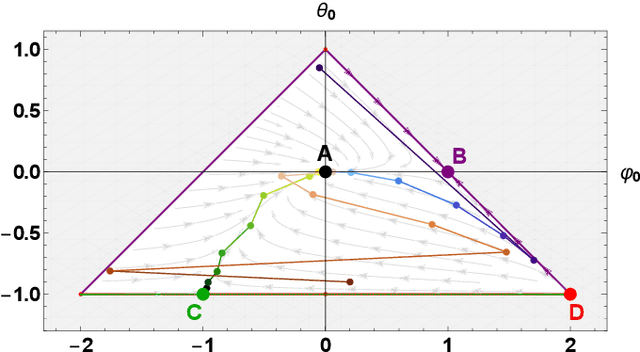

Abstract:Learning the continuous equations of motion from discrete observations is a common task in all areas of physics. However, not any discretization of a Gaussian continuous-time stochastic process can be adopted in parametric inference. We show that discretizations yielding consistent estimators have the property of `invariance under coarse-graining', and correspond to fixed points of a renormalization group map on the space of autoregressive moving average (ARMA) models (for linear processes). This result explains why combining differencing schemes for derivatives reconstruction and local-in-time inference approaches does not work for time series analysis of second or higher order stochastic differential equations, even if the corresponding integration schemes may be acceptably good for numerical simulations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge