Tue Herlau
Probability trees and the value of a single intervention
May 18, 2022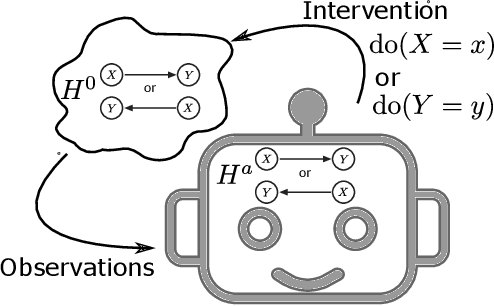
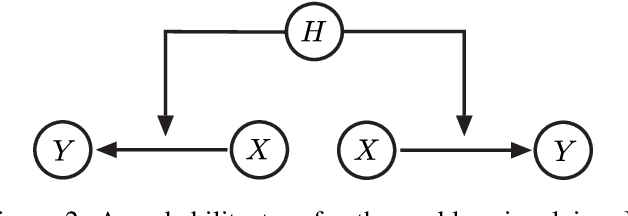
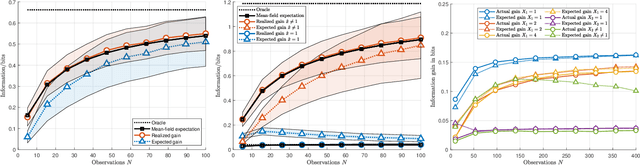
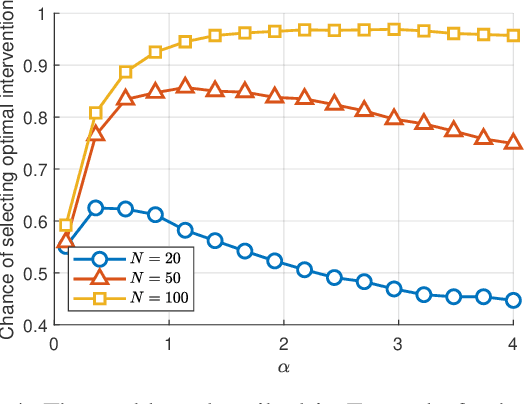
Abstract:The most fundamental problem in statistical causality is determining causal relationships from limited data. Probability trees, which combine prior causal structures with Bayesian updates, have been suggested as a possible solution. In this work, we quantify the information gain from a single intervention and show that both the anticipated information gain, prior to making an intervention, and the expected gain from an intervention have simple expressions. This results in an active-learning method that simply selects the intervention with the highest anticipated gain, which we illustrate through several examples. Our work demonstrates how probability trees, and Bayesian estimation of their parameters, offer a simple yet viable approach to fast causal induction.
Moral reinforcement learning using actual causation
May 17, 2022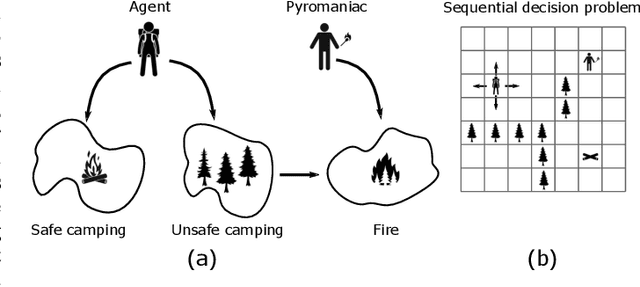
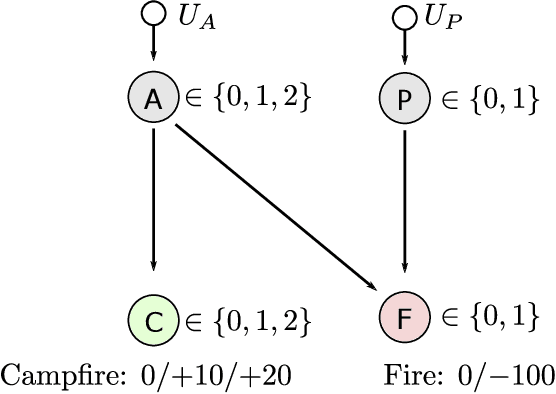
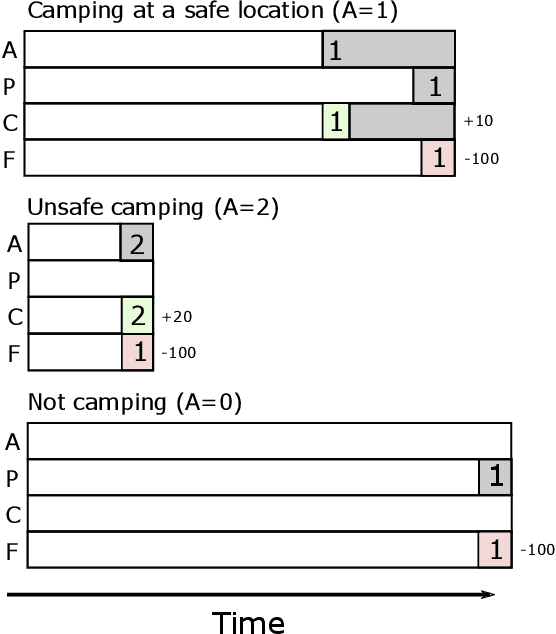
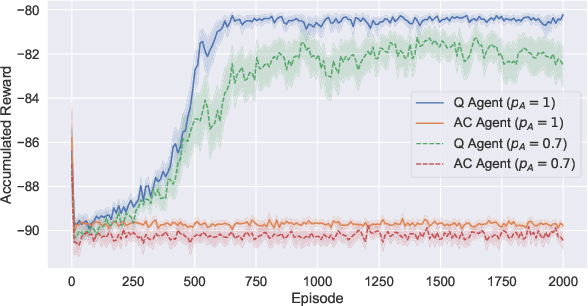
Abstract:Reinforcement learning systems will to a greater and greater extent make decisions that significantly impact the well-being of humans, and it is therefore essential that these systems make decisions that conform to our expectations of morally good behavior. The morally good is often defined in causal terms, as in whether one's actions have in fact caused a particular outcome, and whether the outcome could have been anticipated. We propose an online reinforcement learning method that learns a policy under the constraint that the agent should not be the cause of harm. This is accomplished by defining cause using the theory of actual causation and assigning blame to the agent when its actions are the actual cause of an undesirable outcome. We conduct experiments on a toy ethical dilemma in which a natural choice of reward function leads to clearly undesirable behavior, but our method learns a policy that avoids being the cause of harmful behavior, demonstrating the soundness of our approach. Allowing an agent to learn while observing causal moral distinctions such as blame, opens the possibility to learning policies that better conform to our moral judgments.
Active learning of causal probability trees
May 17, 2022
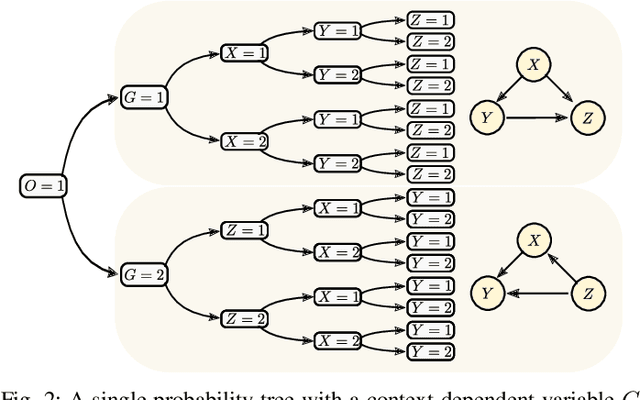
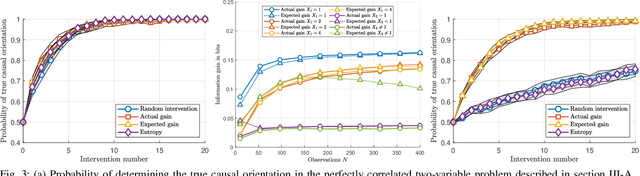
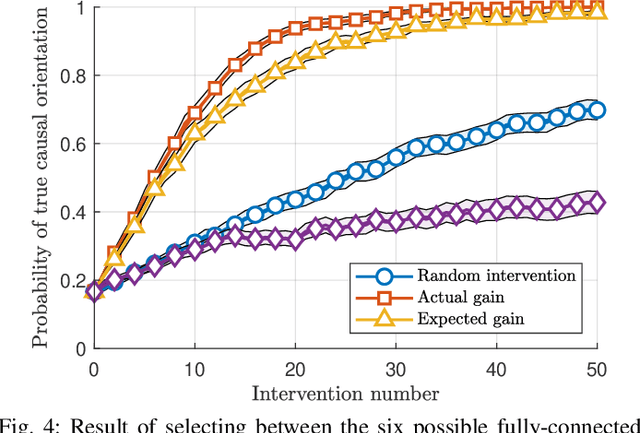
Abstract:The past two decades have seen a growing interest in combining causal information, commonly represented using causal graphs, with machine learning models. Probability trees provide a simple yet powerful alternative representation of causal information. They enable both computation of intervention and counterfactuals, and are strictly more general, since they allow context-dependent causal dependencies. Here we present a Bayesian method for learning probability trees from a combination of interventional and observational data. The method quantifies the expected information gain from an intervention, and selects the interventions with the largest gain. We demonstrate the efficiency of the method on simulated and real data. An effective method for learning probability trees on a limited interventional budget will greatly expand their applicability.
Causal variables from reinforcement learning using generalized Bellman equations
Oct 29, 2020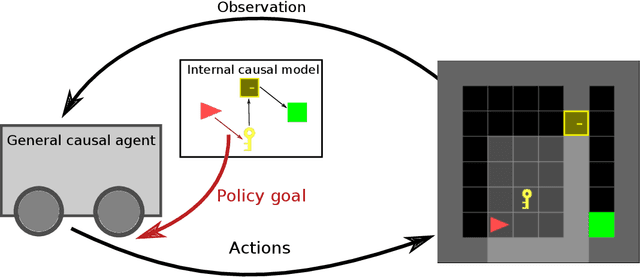

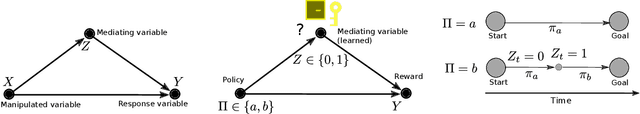
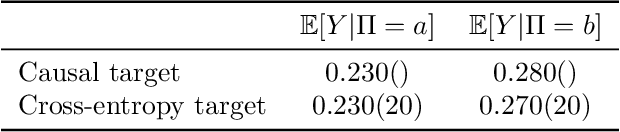
Abstract:Many open problems in machine learning are intrinsically related to causality, however, the use of causal analysis in machine learning is still in its early stage. Within a general reinforcement learning setting, we consider the problem of building a general reinforcement learning agent which uses experience to construct a causal graph of the environment, and use this graph to inform its policy. Our approach has three characteristics: First, we learn a simple, coarse-grained causal graph, in which the variables reflect states at many time instances, and the interventions happen at the level of policies, rather than individual actions. Secondly, we use mediation analysis to obtain an optimization target. By minimizing this target, we define the causal variables. Thirdly, our approach relies on estimating conditional expectations rather the familiar expected return from reinforcement learning, and we therefore apply a generalization of Bellman's equations. We show the method can learn a plausible causal graph in a grid-world environment, and the agent obtains an improvement in performance when using the causally informed policy. To our knowledge, this is the first attempt to apply causal analysis in a reinforcement learning setting without strict restrictions on the number of states. We have observed that mediation analysis provides a promising avenue for transforming the problem of causal acquisition into one of cost-function minimization, but importantly one which involves estimating conditional expectations. This is a new challenge, and we think that causal reinforcement learning will involve development methods suited for online estimation of such conditional expectations. Finally, a benefit of our approach is the use of very simple causal models, which are arguably a more natural model of human causal understanding.
Completely random measures for modelling block-structured networks
Dec 04, 2015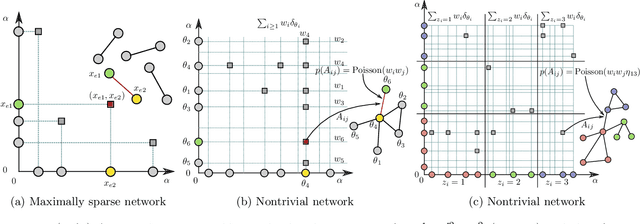
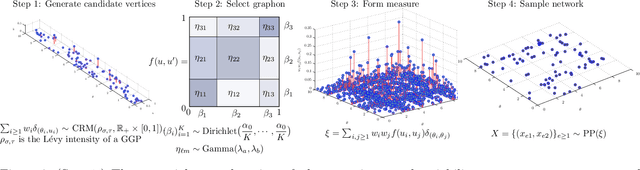
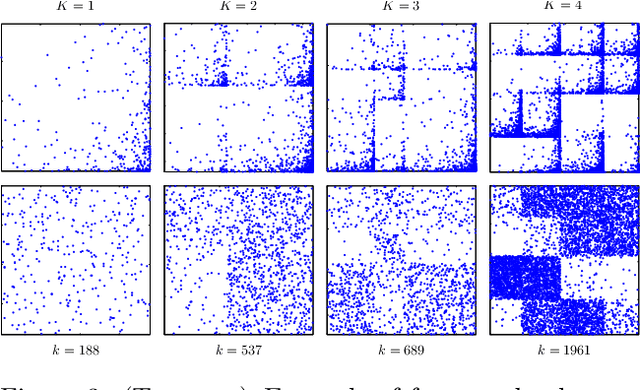
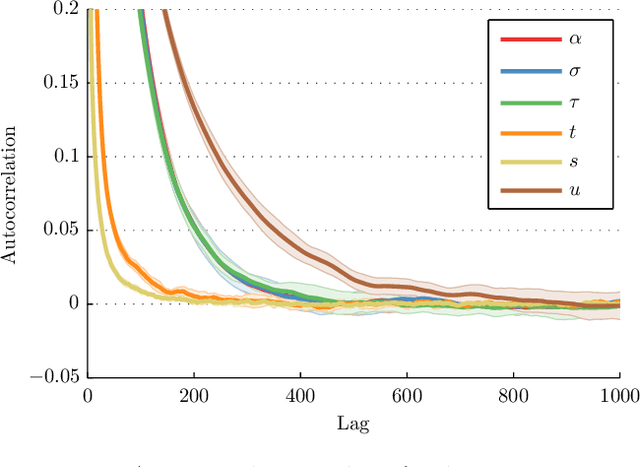
Abstract:Many statistical methods for network data parameterize the edge-probability by attributing latent traits to the vertices such as block structure and assume exchangeability in the sense of the Aldous-Hoover representation theorem. Empirical studies of networks indicate that many real-world networks have a power-law distribution of the vertices which in turn implies the number of edges scale slower than quadratically in the number of vertices. These assumptions are fundamentally irreconcilable as the Aldous-Hoover theorem implies quadratic scaling of the number of edges. Recently Caron and Fox (2014) proposed the use of a different notion of exchangeability due to Kallenberg (2009) and obtained a network model which admits power-law behaviour while retaining desirable statistical properties, however this model does not capture latent vertex traits such as block-structure. In this work we re-introduce the use of block-structure for network models obeying Kallenberg's notion of exchangeability and thereby obtain a model which admits the inference of block-structure and edge inhomogeneity. We derive a simple expression for the likelihood and an efficient sampling method. The obtained model is not significantly more difficult to implement than existing approaches to block-modelling and performs well on real network datasets.
Bayesian Dropout
Aug 12, 2015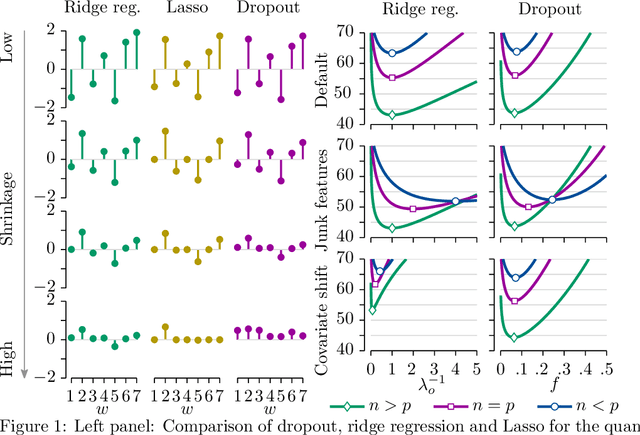
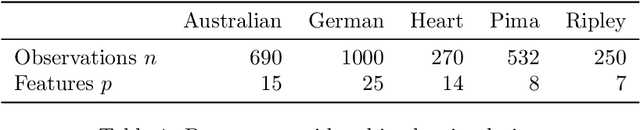
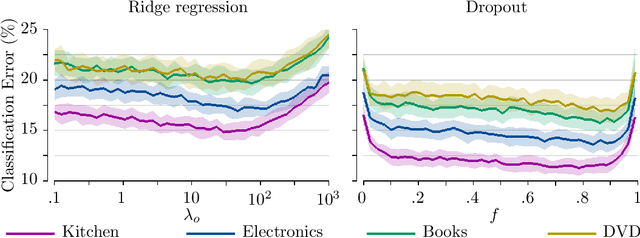
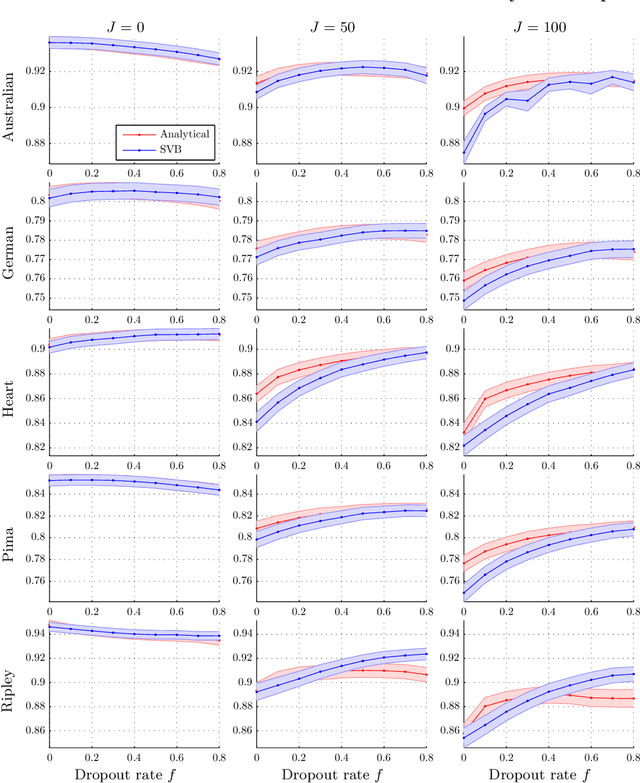
Abstract:Dropout has recently emerged as a powerful and simple method for training neural networks preventing co-adaptation by stochastically omitting neurons. Dropout is currently not grounded in explicit modelling assumptions which so far has precluded its adoption in Bayesian modelling. Using Bayesian entropic reasoning we show that dropout can be interpreted as optimal inference under constraints. We demonstrate this on an analytically tractable regression model providing a Bayesian interpretation of its mechanism for regularizing and preventing co-adaptation as well as its connection to other Bayesian techniques. We also discuss two general approximate techniques for applying Bayesian dropout for general models, one based on an analytical approximation and the other on stochastic variational techniques. These techniques are then applied to a Baysian logistic regression problem and are shown to improve performance as the model become more misspecified. Our framework roots dropout as a theoretically justified and practical tool for statistical modelling allowing Bayesians to tap into the benefits of dropout training.
Efficient inference of overlapping communities in complex networks
Nov 28, 2014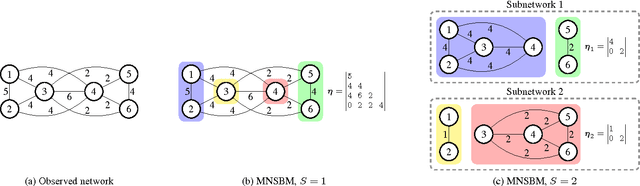
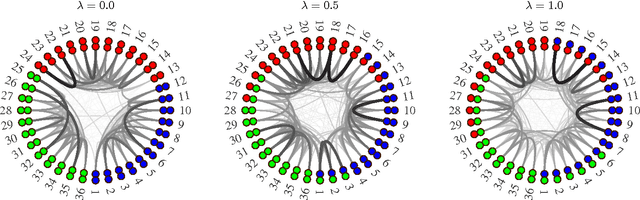
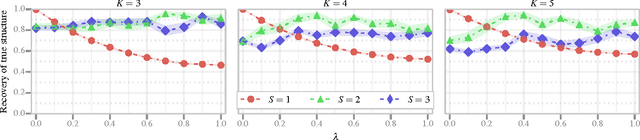
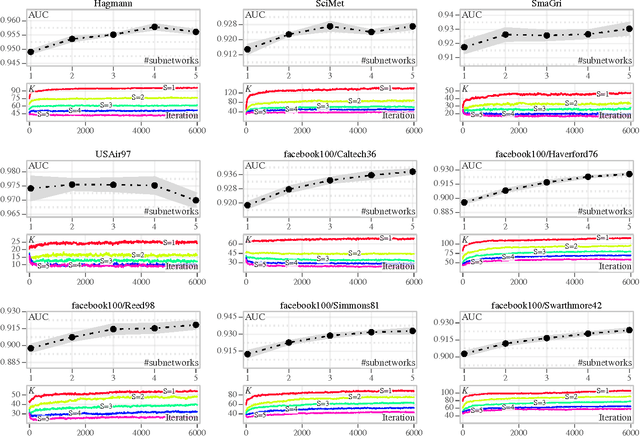
Abstract:We discuss two views on extending existing methods for complex network modeling which we dub the communities first and the networks first view, respectively. Inspired by the networks first view that we attribute to White, Boorman, and Breiger (1976)[1], we formulate the multiple-networks stochastic blockmodel (MNSBM), which seeks to separate the observed network into subnetworks of different types and where the problem of inferring structure in each subnetwork becomes easier. We show how this model is specified in a generative Bayesian framework where parameters can be inferred efficiently using Gibbs sampling. The result is an effective multiple-membership model without the drawbacks of introducing complex definitions of "groups" and how they interact. We demonstrate results on the recovery of planted structure in synthetic networks and show very encouraging results on link prediction performances using multiple-networks models on a number of real-world network data sets.
Adaptive Reconfiguration Moves for Dirichlet Mixtures
May 31, 2014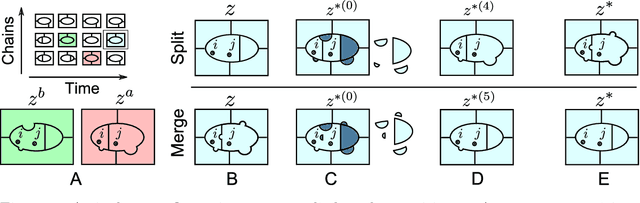
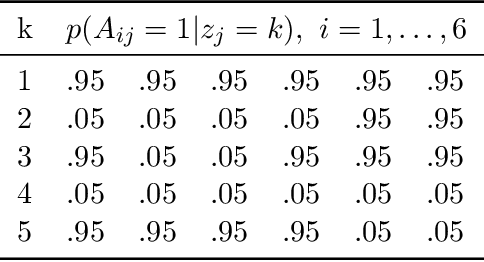

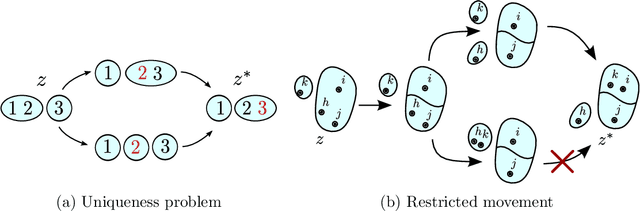
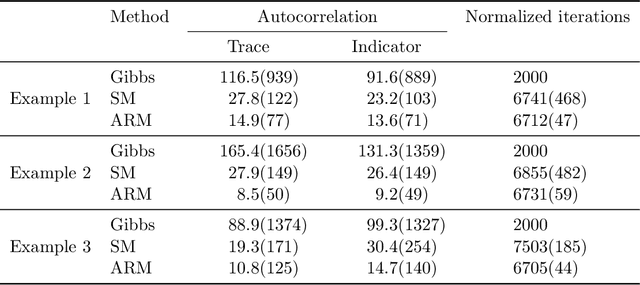
Abstract:Bayesian mixture models are widely applied for unsupervised learning and exploratory data analysis. Markov chain Monte Carlo based on Gibbs sampling and split-merge moves are widely used for inference in these models. However, both methods are restricted to limited types of transitions and suffer from torpid mixing and low accept rates even for problems of modest size. We propose a method that considers a broader range of transitions that are close to equilibrium by exploiting multiple chains in parallel and using the past states adaptively to inform the proposal distribution. The method significantly improves on Gibbs and split-merge sampling as quantified using convergence diagnostics and acceptance rates. Adaptive MCMC methods which use past states to inform the proposal distribution has given rise to many ingenious sampling schemes for continuous problems and the present work can be seen as an important first step in bringing these benefits to partition-based problems
The Infinite Degree Corrected Stochastic Block Model
May 30, 2014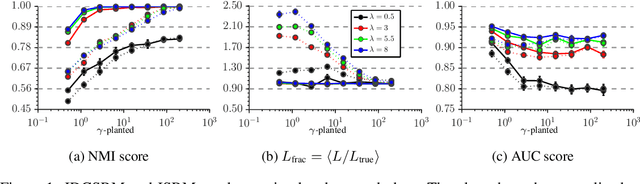
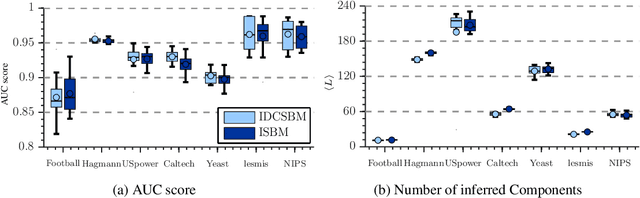
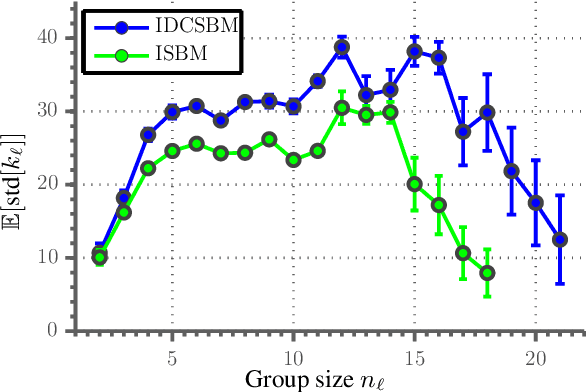
Abstract:In Stochastic blockmodels, which are among the most prominent statistical models for cluster analysis of complex networks, clusters are defined as groups of nodes with statistically similar link probabilities within and between groups. A recent extension by Karrer and Newman incorporates a node degree correction to model degree heterogeneity within each group. Although this demonstrably leads to better performance on several networks it is not obvious whether modelling node degree is always appropriate or necessary. We formulate the degree corrected stochastic blockmodel as a non-parametric Bayesian model, incorporating a parameter to control the amount of degree correction which can then be inferred from data. Additionally, our formulation yields principled ways of inferring the number of groups as well as predicting missing links in the network which can be used to quantify the model's predictive performance. On synthetic data we demonstrate that including the degree correction yields better performance both on recovering the true group structure and predicting missing links when degree heterogeneity is present, whereas performance is on par for data with no degree heterogeneity within clusters. On seven real networks (with no ground truth group structure available) we show that predictive performance is about equal whether or not degree correction is included; however, for some networks significantly fewer clusters are discovered when correcting for degree indicating that the data can be more compactly explained by clusters of heterogenous degree nodes.
* Originally presented at the Complex Networks workshop NIPS 2013
Nonparametric Bayesian models of hierarchical structure in complex networks
Nov 21, 2013
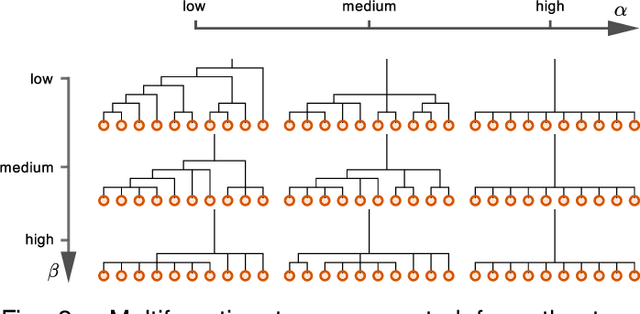
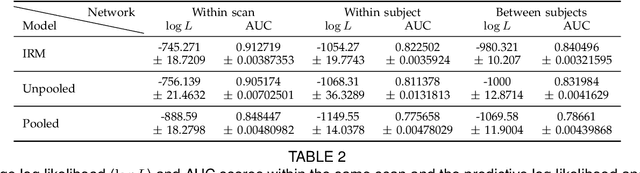
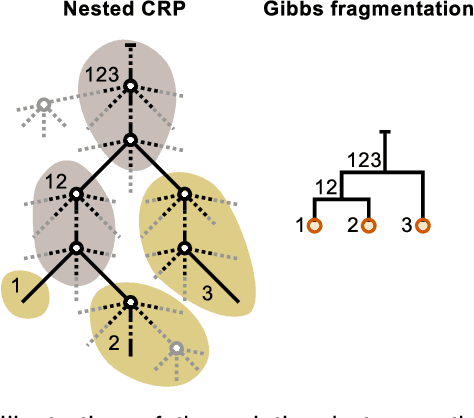
Abstract:Analyzing and understanding the structure of complex relational data is important in many applications including analysis of the connectivity in the human brain. Such networks can have prominent patterns on different scales, calling for a hierarchically structured model. We propose two non-parametric Bayesian hierarchical network models based on Gibbs fragmentation tree priors, and demonstrate their ability to capture nested patterns in simulated networks. On real networks we demonstrate detection of hierarchical structure and show predictive performance on par with the state of the art. We envision that our methods can be employed in exploratory analysis of large scale complex networks for example to model human brain connectivity.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge