Tomas Vicar
Weakly supervised deep learning-based intracranial hemorrhage localization
May 03, 2021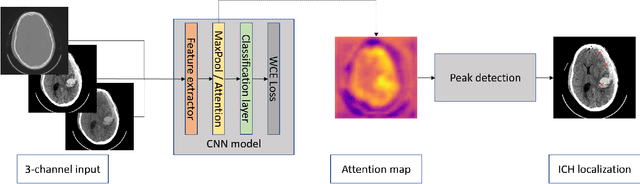
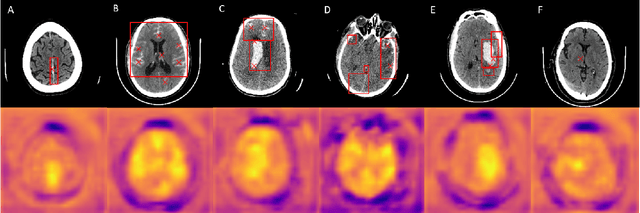
Abstract:Intracranial hemorrhage is a life-threatening disease, which requires fast medical intervention. Owing to the duration of data annotation, head CT images are usually available only with slice-level labeling. This paper presents a weakly supervised method of precise hemorrhage localization in axial slices using only position-free labels, which is based on multiple instance learning. An algorithm is introduced that generates hemorrhage likelihood maps and finds the coordinates of bleeding. The Dice coefficient of 58.08 % is achieved on data from a publicly available dataset.
A Tool for Automatic Estimation of Patient Position in Spinal CT Data
Jun 27, 2020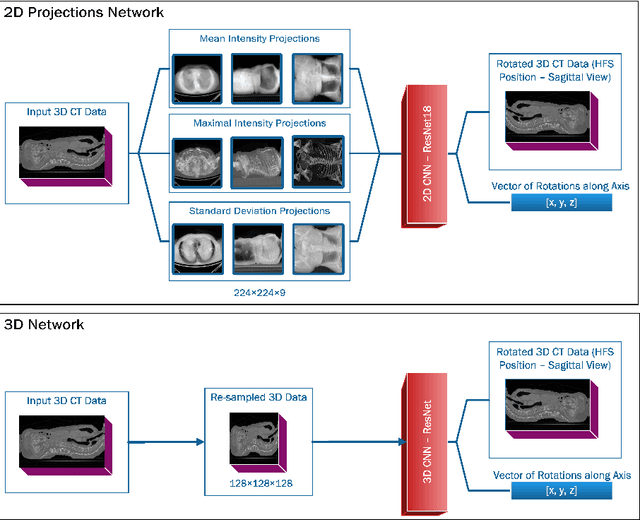
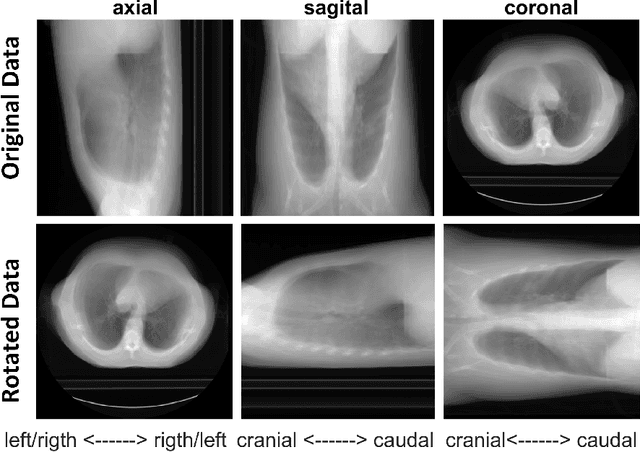
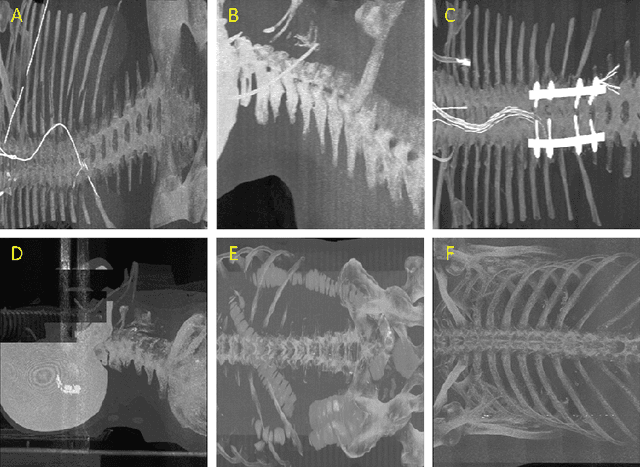
Abstract:Much of the recently available research and challenge data lack the meta-data containing any information about the patient position. This paper presents a tool for automatic rotation of CT data into a standardized (HFS) patient position. The proposed method is based on the prediction of rotation angle with CNN, and it achieved nearly perfect results with an accuracy of 99.55 %. We provide implementations with easy to use an example for both Matlab and Python (PyTorch), which can be used, for example, for automatic rotation correction of VerSe2020 challenge data.
Probabilistic Noise2Void: Unsupervised Content-Aware Denoising
Jun 04, 2019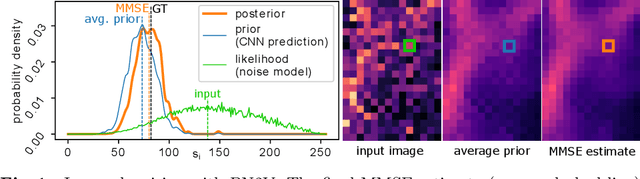
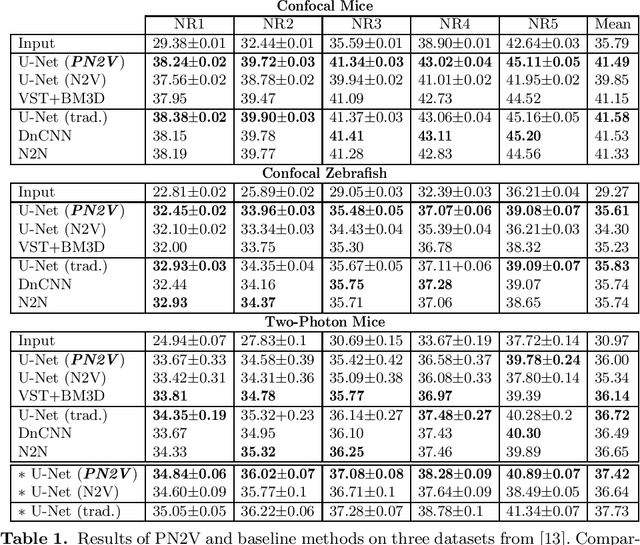

Abstract:Today, Convolutional Neural Networks (CNNs) are the leading method for image denoising. They are traditionally trained on pairs of images, which are often hard to obtain for practical applications. This motivates self-supervised training methods such as Noise2Void~(N2V) that operate on single noisy images. Self-supervised methods are, unfortunately, not competitive with models trained on image pairs. Here, we present 'Probabilistic Noise2Void' (PN2V), a method to train CNNs to predict per-pixel intensity distributions. Combining these with a suitable description of the noise, we obtain a complete probabilistic model for the noisy observations and true signal in every pixel. We evaluate PN2V on publicly available microscopy datasets, under a broad range of noise regimes, and achieve competitive results with respect to supervised state-of-the-art methods.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge