Thijs Vande Vyvere
Unsupervised 3D Brain Anomaly Detection
Oct 09, 2020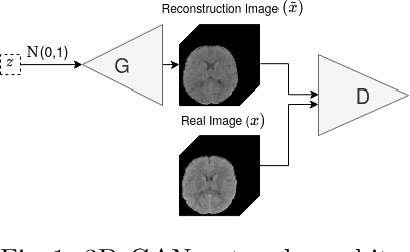
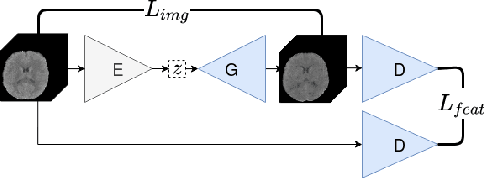
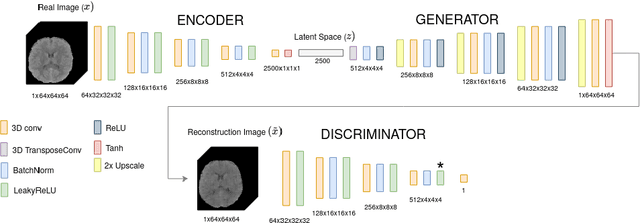
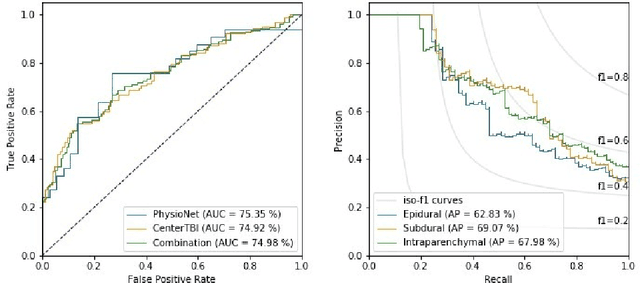
Abstract:Anomaly detection (AD) is the identification of data samples that do not fit a learned data distribution. As such, AD systems can help physicians to determine the presence, severity, and extension of a pathology. Deep generative models, such as Generative Adversarial Networks (GANs), can be exploited to capture anatomical variability. Consequently, any outlier (i.e., sample falling outside of the learned distribution) can be detected as an abnormality in an unsupervised fashion. By using this method, we can not only detect expected or known lesions, but we can even unveil previously unrecognized biomarkers. To the best of our knowledge, this study exemplifies the first AD approach that can efficiently handle volumetric data and detect 3D brain anomalies in one single model. Our proposal is a volumetric and high-detail extension of the 2D f-AnoGAN model obtained by combining a state-of-the-art 3D GAN with refinement training steps. In experiments using non-contrast computed tomography images from traumatic brain injury (TBI) patients, the model detects and localizes TBI abnormalities with an area under the ROC curve of ~75%. Moreover, we test the potential of the method for detecting other anomalies such as low quality images, preprocessing inaccuracies, artifacts, and even the presence of post-operative signs (such as a craniectomy or a brain shunt). The method has potential for rapidly labeling abnormalities in massive imaging datasets, as well as identifying new biomarkers.
A Radiomics Approach to Traumatic Brain Injury Prediction in CT Scans
Nov 14, 2018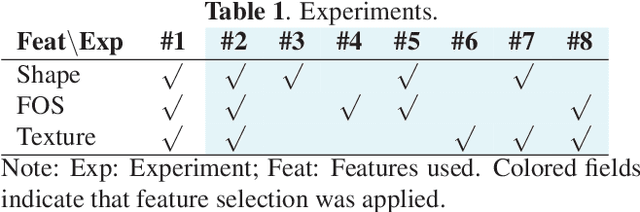
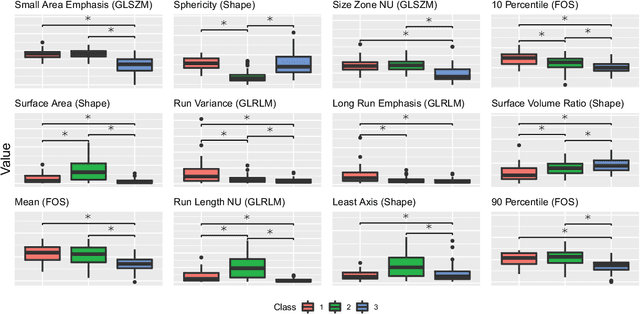
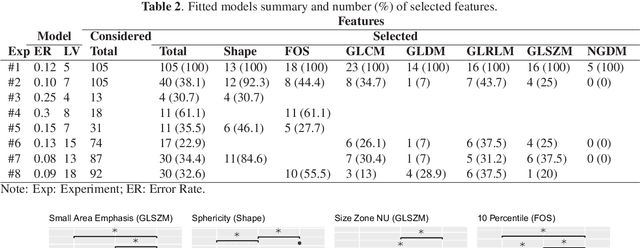
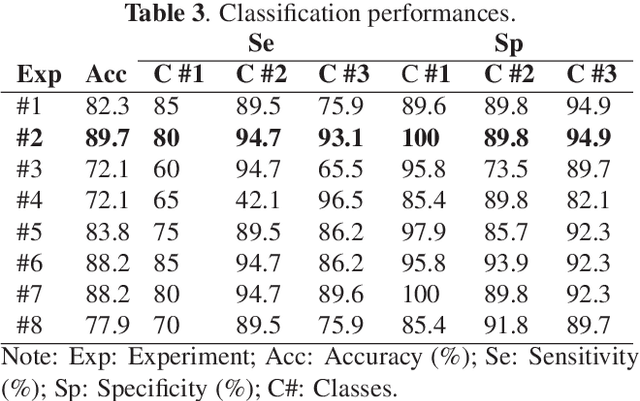
Abstract:Computer Tomography (CT) is the gold standard technique for brain damage evaluation after acute Traumatic Brain Injury (TBI). It allows identification of most lesion types and determines the need of surgical or alternative therapeutic procedures. However, the traditional approach for lesion classification is restricted to visual image inspection. In this work, we characterize and predict TBI lesions by using CT-derived radiomics descriptors. Relevant shape, intensity and texture biomarkers characterizing the different lesions are isolated and a lesion predictive model is built by using Partial Least Squares. On a dataset containing 155 scans (105 train, 50 test) the methodology achieved 89.7 % accuracy over the unseen data. When a model was build using only texture features, a 88.2 % accuracy was obtained. Our results suggest that selected radiomics descriptors could play a key role in brain injury prediction. Besides, the proposed methodology is close to reproduce radiologists decision making. These results open new possibilities for radiomics-inspired brain lesion detection, segmentation and prediction.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge