Theodore J. Perkins
Fluid-Agent Reinforcement Learning
Feb 16, 2026Abstract:The primary focus of multi-agent reinforcement learning (MARL) has been to study interactions among a fixed number of agents embedded in an environment. However, in the real world, the number of agents is neither fixed nor known a priori. Moreover, an agent can decide to create other agents (for example, a cell may divide, or a company may spin off a division). In this paper, we propose a framework that allows agents to create other agents; we call this a fluid-agent environment. We present game-theoretic solution concepts for fluid-agent games and empirically evaluate the performance of several MARL algorithms within this framework. Our experiments include fluid variants of established benchmarks such as Predator-Prey and Level-Based Foraging, where agents can dynamically spawn, as well as a new environment we introduce that highlights how fluidity can unlock novel solution strategies beyond those observed in fixed-population settings. We demonstrate that this framework yields agent teams that adjust their size dynamically to match environmental demands.
Is graph-based feature selection of genes better than random?
Nov 19, 2019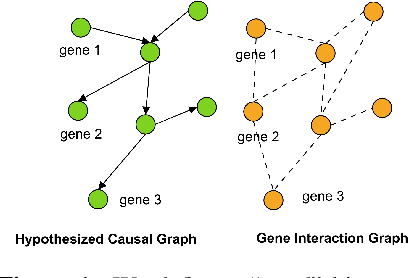
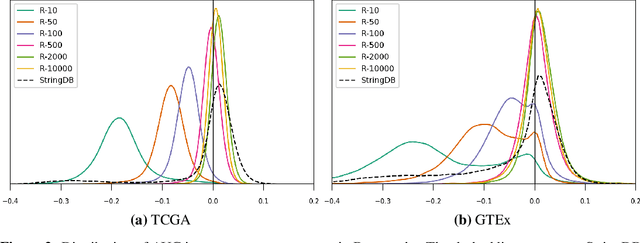
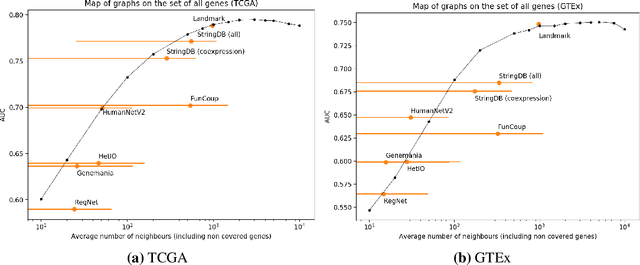
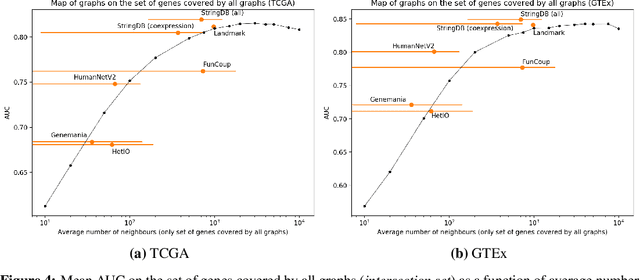
Abstract:Gene interaction graphs aim to capture various relationships between genes and represent decades of biology research. When trying to make predictions from genomic data, those graphs could be used to overcome the curse of dimensionality by making machine learning models sparser and more consistent with biological common knowledge. In this work, we focus on assessing whether those graphs capture dependencies seen in gene expression data better than random. We formulate a condition that graphs should satisfy to provide a good prior knowledge and propose to test it using a `Single Gene Inference' (SGI) task. We compare random graphs with seven major gene interaction graphs published by different research groups, aiming to measure the true benefit of using biologically relevant graphs in this context. Our analysis finds that dependencies can be captured almost as well at random which suggests that, in terms of gene expression levels, the relevant information about the state of the cell is spread across many genes.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge