Teemu Roos
LEMUR: Learned Multi-Vector Retrieval
Jan 29, 2026Abstract:Multi-vector representations generated by late interaction models, such as ColBERT, enable superior retrieval quality compared to single-vector representations in information retrieval applications. In multi-vector retrieval systems, both queries and documents are encoded using one embedding for each token, and similarity between queries and documents is measured by the MaxSim similarity measure. However, the improved recall of multi-vector retrieval comes at the expense of significantly increased latency. This necessitates designing efficient approximate nearest neighbor search (ANNS) algorithms for multi-vector search. In this work, we introduce LEMUR, a simple-yet-efficient framework for multi-vector similarity search. LEMUR consists of two consecutive problem reductions: We first formulate multi-vector similarity search as a supervised learning problem that can be solved using a one-hidden-layer neural network. Second, we reduce inference under this model to single-vector similarity search in its latent space, which enables the use of existing single-vector ANNS methods for speeding up retrieval. In addition to performance evaluation on ColBERTv2 embeddings, we evaluate LEMUR on embeddings generated by modern multi-vector text models and multi-vector visual document retrieval models. LEMUR is an order of magnitude faster than earlier multi-vector similarity search methods.
Deterministic and probabilistic neural surrogates of global hybrid-Vlasov simulations
Jan 21, 2026Abstract:Hybrid-Vlasov simulations resolve ion-kinetic effects for modeling the solar wind-magnetosphere interaction, but even 5D (2D + 3V) simulations are computationally expensive. We show that graph-based machine learning emulators can learn the spatiotemporal evolution of electromagnetic fields and lower order moments of ion velocity distribution in the near-Earth space environment from four 5D Vlasiator runs performed with identical steady solar wind conditions. The initial ion number density is systematically varied, while the grid spacing is held constant, to scan the ratio of the characteristic ion skin depth to the numerical grid size. Using a graph neural network architecture operating on the 2D spatial simulation grid comprising 670k cells, we demonstrate that both a deterministic forecasting model (Graph-FM) and a probabilistic ensemble forecasting model (Graph-EFM) based on a latent variable formulation are capable of producing accurate predictions of future plasma states. A divergence penalty is incorporated during training to encourage divergence-freeness in the magnetic fields and improve physical consistency. For the probabilistic model, a continuous ranked probability score objective is added to improve the calibration of the ensemble forecasts. When trained, the emulators achieve more than two orders of magnitude speedup in generating the next time step relative to the original simulation on a single GPU compared to 100 CPUs for the Vlasiator runs, while closely matching physical magnetospheric response of the different runs. These results demonstrate that machine learning offers a way to make hybrid-Vlasov simulation tractable for real-time use while providing forecast uncertainty.
VIBE: Vector Index Benchmark for Embeddings
May 23, 2025Abstract:Approximate nearest neighbor (ANN) search is a performance-critical component of many machine learning pipelines. Rigorous benchmarking is essential for evaluating the performance of vector indexes for ANN search. However, the datasets of the existing benchmarks are no longer representative of the current applications of ANN search. Hence, there is an urgent need for an up-to-date set of benchmarks. To this end, we introduce Vector Index Benchmark for Embeddings (VIBE), an open source project for benchmarking ANN algorithms. VIBE contains a pipeline for creating benchmark datasets using dense embedding models characteristic of modern applications, such as retrieval-augmented generation (RAG). To replicate real-world workloads, we also include out-of-distribution (OOD) datasets where the queries and the corpus are drawn from different distributions. We use VIBE to conduct a comprehensive evaluation of SOTA vector indexes, benchmarking 21 implementations on 12 in-distribution and 6 out-of-distribution datasets.
LoRANN: Low-Rank Matrix Factorization for Approximate Nearest Neighbor Search
Oct 24, 2024



Abstract:Approximate nearest neighbor (ANN) search is a key component in many modern machine learning pipelines; recent use cases include retrieval-augmented generation (RAG) and vector databases. Clustering-based ANN algorithms, that use score computation methods based on product quantization (PQ), are often used in industrial-scale applications due to their scalability and suitability for distributed and disk-based implementations. However, they have slower query times than the leading graph-based ANN algorithms. In this work, we propose a new supervised score computation method based on the observation that inner product approximation is a multivariate (multi-output) regression problem that can be solved efficiently by reduced-rank regression. Our experiments show that on modern high-dimensional data sets, the proposed reduced-rank regression (RRR) method is superior to PQ in both query latency and memory usage. We also introduce LoRANN, a clustering-based ANN library that leverages the proposed score computation method. LoRANN is competitive with the leading graph-based algorithms and outperforms the state-of-the-art GPU ANN methods on high-dimensional data sets.
Regional Ocean Forecasting with Hierarchical Graph Neural Networks
Oct 15, 2024



Abstract:Accurate ocean forecasting systems are vital for understanding marine dynamics, which play a crucial role in environmental management and climate adaptation strategies. Traditional numerical solvers, while effective, are computationally expensive and time-consuming. Recent advancements in machine learning have revolutionized weather forecasting, offering fast and energy-efficient alternatives. Building on these advancements, we introduce SeaCast, a neural network designed for high-resolution, medium-range ocean forecasting. SeaCast employs a graph-based framework to effectively handle the complex geometry of ocean grids and integrates external forcing data tailored to the regional ocean context. Our approach is validated through experiments at a high spatial resolution using the operational numerical model of the Mediterranean Sea provided by the Copernicus Marine Service, along with both numerical and data-driven atmospheric forcings.
Quotient Normalized Maximum Likelihood Criterion for Learning Bayesian Network Structures
Aug 27, 2024Abstract:We introduce an information theoretic criterion for Bayesian network structure learning which we call quotient normalized maximum likelihood (qNML). In contrast to the closely related factorized normalized maximum likelihood criterion, qNML satisfies the property of score equivalence. It is also decomposable and completely free of adjustable hyperparameters. For practical computations, we identify a remarkably accurate approximation proposed earlier by Szpankowski and Weinberger. Experiments on both simulated and real data demonstrate that the new criterion leads to parsimonious models with good predictive accuracy.
* Accepted to AISTATS 2018
Modeling 3D Infant Kinetics Using Adaptive Graph Convolutional Networks
Feb 22, 2024



Abstract:Reliable methods for the neurodevelopmental assessment of infants are essential for early detection of medical issues that may need prompt interventions. Spontaneous motor activity, or `kinetics', is shown to provide a powerful surrogate measure of upcoming neurodevelopment. However, its assessment is by and large qualitative and subjective, focusing on visually identified, age-specific gestures. Here, we follow an alternative approach, predicting infants' neurodevelopmental maturation based on data-driven evaluation of individual motor patterns. We utilize 3D video recordings of infants processed with pose-estimation to extract spatio-temporal series of anatomical landmarks, and apply adaptive graph convolutional networks to predict the actual age. We show that our data-driven approach achieves improvement over traditional machine learning baselines based on manually engineered features.
Graph Representation of the Magnetic Field Topology in High-Fidelity Plasma Simulations for Machine Learning Applications
Jul 26, 2023Abstract:Topological analysis of the magnetic field in simulated plasmas allows the study of various physical phenomena in a wide range of settings. One such application is magnetic reconnection, a phenomenon related to the dynamics of the magnetic field topology, which is difficult to detect and characterize in three dimensions. We propose a scalable pipeline for topological data analysis and spatiotemporal graph representation of three-dimensional magnetic vector fields. We demonstrate our methods on simulations of the Earth's magnetosphere produced by Vlasiator, a supercomputer-scale Vlasov theory-based simulation for near-Earth space. The purpose of this work is to challenge the machine learning community to explore graph-based machine learning approaches to address a largely open scientific problem with wide-ranging potential impact.
Transfer Learning with Ensembles of Deep Neural Networks for Skin Cancer Classification in Imbalanced Data Sets
Mar 30, 2021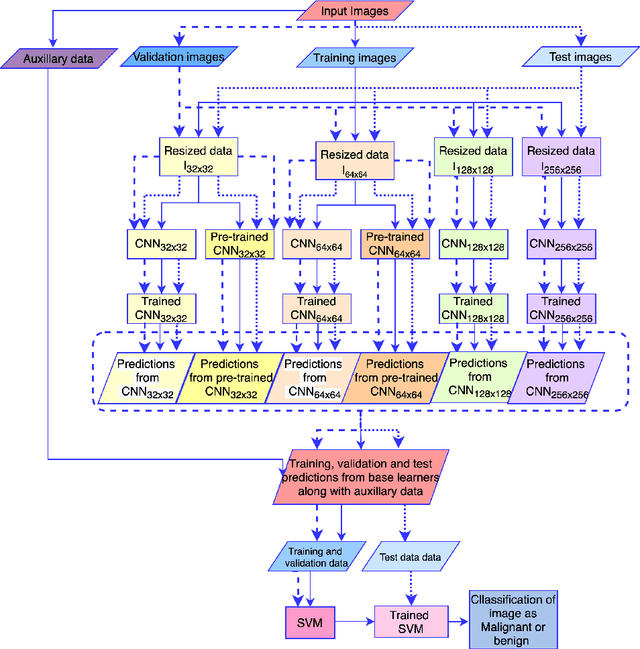
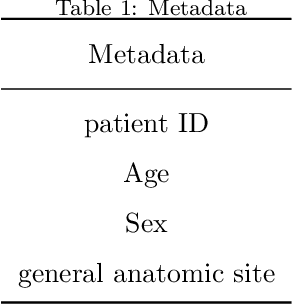
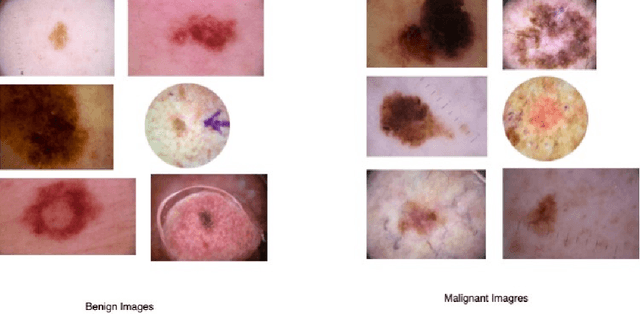
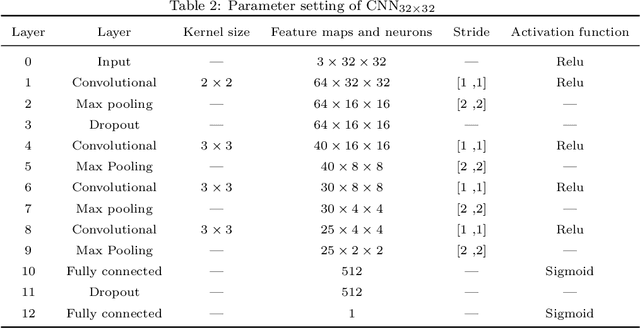
Abstract:Early diagnosis plays a key role in prevention and treatment of skin cancer.Several machine learning techniques for accurate classification of skin cancer from medical images have been reported. Many of these techniques are based on pre-trained convolutional neural networks (CNNs), which enable training the models based on limited amounts of training data. However, the classification accuracy of these models still tends to be severely limited by the scarcity of representative images from malignant tumours. We propose a novel ensemble-based CNN architecture where multiple CNN models, some of which are pre-trained and some are trained only on the data at hand, along with patient information (meta-data) are combined using a meta-learner. The proposed approach improves the model's ability to handle scarce, imbalanced data. We demonstrate the benefits of the proposed technique using a dataset with 33126 dermoscopic images from 2000 patients.We evaluate the performance of the proposed technique in terms of the F1-measure, area under the ROC curve (AUC-ROC), and area under the PR curve (AUC-PR), and compare it with that of seven different benchmark methods, including two recent CNN-based techniques. The proposed technique achieves superior performance in terms of all the evaluation metrics (F1-measure $0.53$, AUC-PR $0.58$, AUC-ROC $0.97$).
Gradient-Based Training and Pruning of Radial Basis Function Networks with an Application in Materials Physics
Apr 06, 2020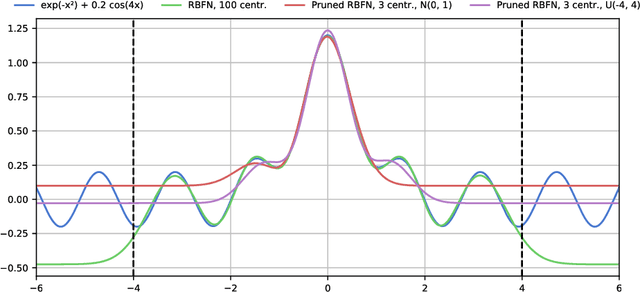
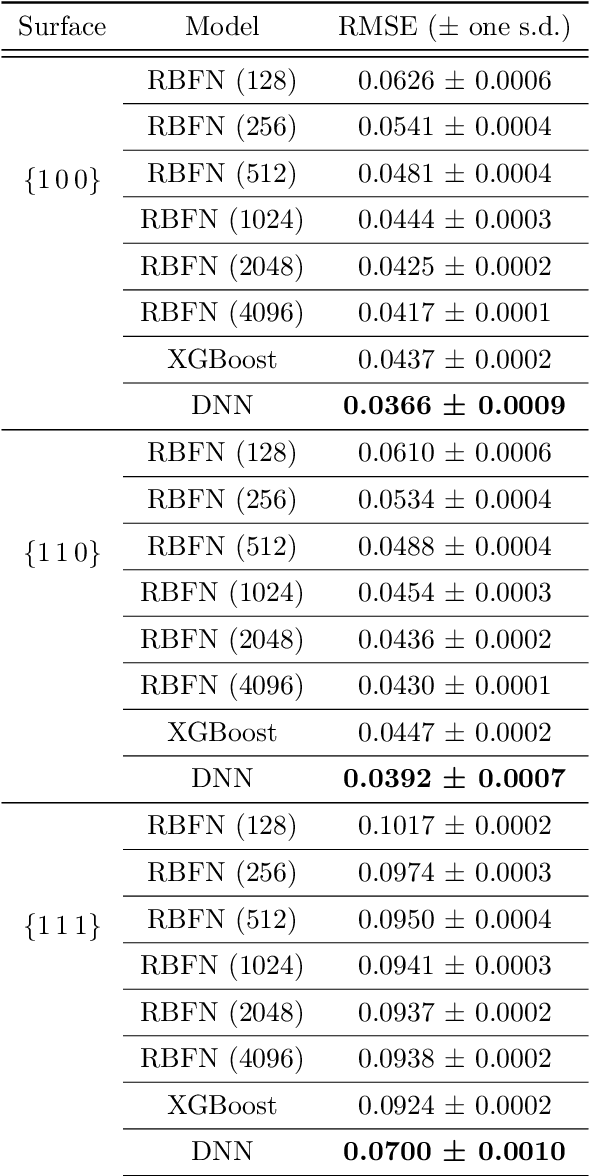

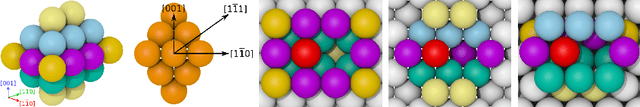
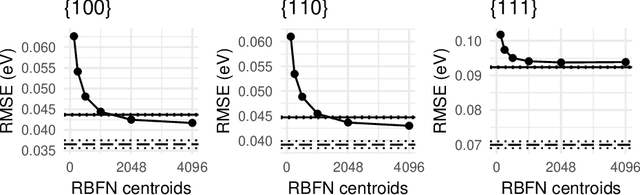
Abstract:Many applications, especially in physics and other sciences, call for easily interpretable and robust machine learning techniques. We propose a fully gradient-based technique for training radial basis function networks with an efficient and scalable open-source implementation. We derive novel closed-form optimization criteria for pruning the models for continuous as well as binary data which arise in a challenging real-world material physics problem. The pruned models are optimized to provide compact and interpretable versions of larger models based on informed assumptions about the data distribution. Visualizations of the pruned models provide insight into the atomic configurations that determine atom-level migration processes in solid matter; these results may inform future research on designing more suitable descriptors for use with machine learning algorithms.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge