Takashi Sano
Investigation of Frame Differences as Motion Cues for Video Object Segmentation
Mar 12, 2025



Abstract:Automatic Video Object Segmentation (AVOS) refers to the task of autonomously segmenting target objects in video sequences without relying on human-provided annotations in the first frames. In AVOS, the use of motion information is crucial, with optical flow being a commonly employed method for capturing motion cues. However, the computation of optical flow is resource-intensive, making it unsuitable for real-time applications, especially on edge devices with limited computational resources. In this study, we propose using frame differences as an alternative to optical flow for motion cue extraction. We developed an extended U-Net-like AVOS model that takes a frame on which segmentation is performed and a frame difference as inputs, and outputs an estimated segmentation map. Our experimental results demonstrate that the proposed model achieves performance comparable to the model with optical flow as an input, particularly when applied to videos captured by stationary cameras. Our results suggest the usefulness of employing frame differences as motion cues in cases with limited computational resources.
From Coupled Oscillators to Graph Neural Networks: Reducing Over-smoothing via a Kuramoto Model-based Approach
Nov 06, 2023



Abstract:We propose the Kuramoto Graph Neural Network (KuramotoGNN), a novel class of continuous-depth graph neural networks (GNNs) that employs the Kuramoto model to mitigate the over-smoothing phenomenon, in which node features in GNNs become indistinguishable as the number of layers increases. The Kuramoto model captures the synchronization behavior of non-linear coupled oscillators. Under the view of coupled oscillators, we first show the connection between Kuramoto model and basic GNN and then over-smoothing phenomenon in GNNs can be interpreted as phase synchronization in Kuramoto model. The KuramotoGNN replaces this phase synchronization with frequency synchronization to prevent the node features from converging into each other while allowing the system to reach a stable synchronized state. We experimentally verify the advantages of the KuramotoGNN over the baseline GNNs and existing methods in reducing over-smoothing on various graph deep learning benchmark tasks.
An Analytic Solution to the Inverse Ising Problem in the Tree-reweighted Approximation
May 29, 2018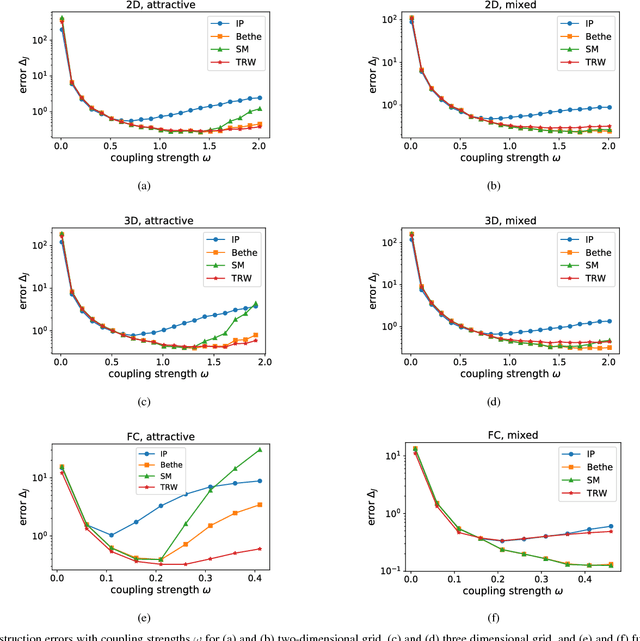
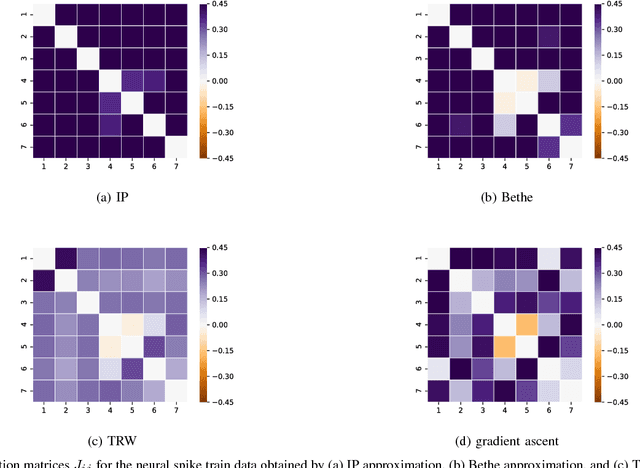
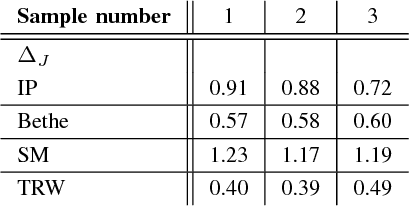
Abstract:Many iterative and non-iterative methods have been developed for inverse problems associated with Ising models. Aiming to derive an accurate non-iterative method for the inverse problems, we employ the tree-reweighted approximation. Using the tree-reweighted approximation, we can optimize the rigorous lower bound of the objective function. By solving the moment-matching and self-consistency conditions analytically, we can derive the interaction matrix as a function of the given data statistics. With this solution, we can obtain the optimal interaction matrix without iterative computation. To evaluate the accuracy of the proposed inverse formula, we compared our results to those obtained by existing inverse formulae derived with other approximations. In an experiment to reconstruct the interaction matrix, we found that the proposed formula returns the best estimates in strongly-attractive regions for various graph structures. We also performed an experiment using real-world biological data. When applied to finding the connectivity of neurons from spike train data, the proposed formula gave the closest result to that obtained by a gradient ascent algorithm, which typically requires thousands of iterations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge