Svetha Venkatesh
Resset: A Recurrent Model for Sequence of Sets with Applications to Electronic Medical Records
Feb 03, 2018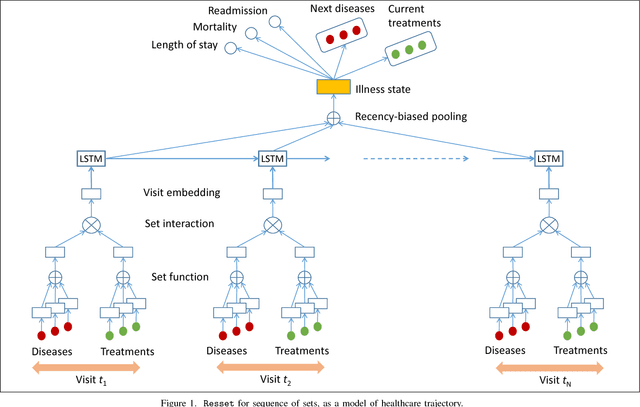
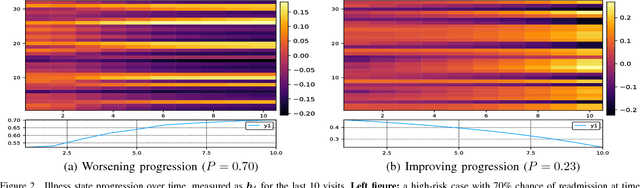

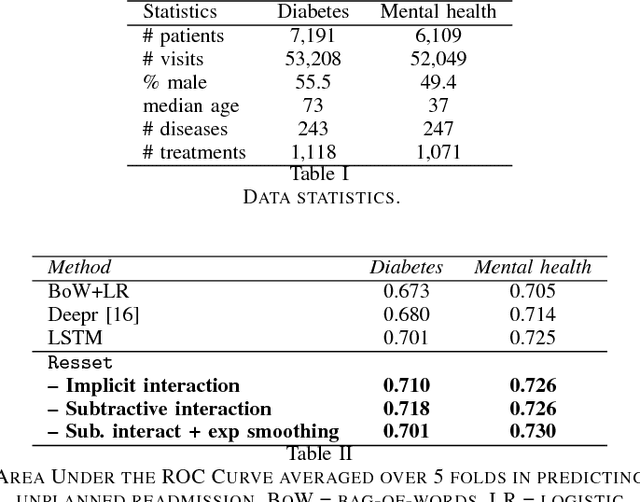
Abstract:Modern healthcare is ripe for disruption by AI. A game changer would be automatic understanding the latent processes from electronic medical records, which are being collected for billions of people worldwide. However, these healthcare processes are complicated by the interaction between at least three dynamic components: the illness which involves multiple diseases, the care which involves multiple treatments, and the recording practice which is biased and erroneous. Existing methods are inadequate in capturing the dynamic structure of care. We propose Resset, an end-to-end recurrent model that reads medical record and predicts future risk. The model adopts the algebraic view in that discrete medical objects are embedded into continuous vectors lying in the same space. We formulate the problem as modeling sequences of sets, a novel setting that have rarely, if not, been addressed. Within Resset, the bag of diseases recorded at each clinic visit is modeled as function of sets. The same hold for the bag of treatments. The interaction between the disease bag and the treatment bag at a visit is modeled in several, one of which as residual of diseases minus the treatments. Finally, the health trajectory, which is a sequence of visits, is modeled using a recurrent neural network. We report results on over a hundred thousand hospital visits by patients suffered from two costly chronic diseases -- diabetes and mental health. Resset shows promises in multiple predictive tasks such as readmission prediction, treatments recommendation and diseases progression.
Graph Memory Networks for Molecular Activity Prediction
Jan 27, 2018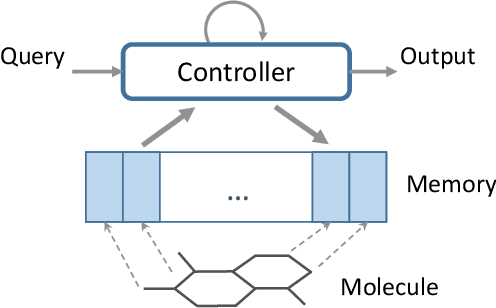
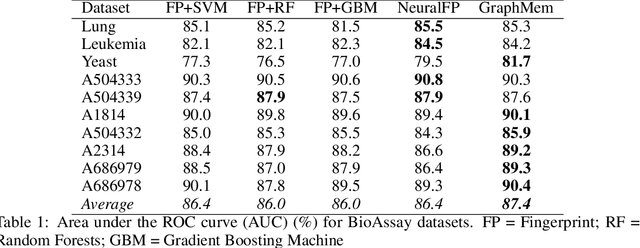
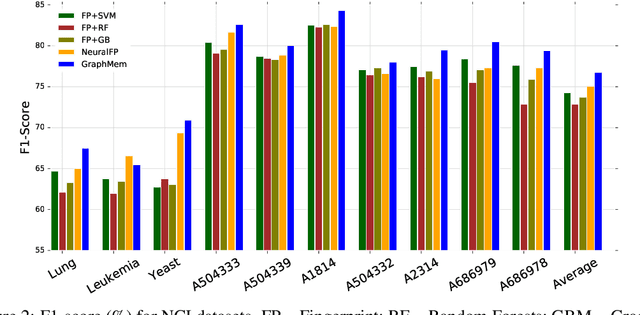
Abstract:Molecular activity prediction is critical in drug design. Machine learning techniques such as kernel methods and random forests have been successful for this task. These models require fixed-size feature vectors as input while the molecules are variable in size and structure. As a result, fixed-size fingerprint representation is poor in handling substructures for large molecules. In addition, molecular activity tests, or a so-called BioAssays, are relatively small in the number of tested molecules due to its complexity. Here we approach the problem through deep neural networks as they are flexible in modeling structured data such as grids, sequences and graphs. We train multiple BioAssays using a multi-task learning framework, which combines information from multiple sources to improve the performance of prediction, especially on small datasets. We propose Graph Memory Network (GraphMem), a memory-augmented neural network to model the graph structure in molecules. GraphMem consists of a recurrent controller coupled with an external memory whose cells dynamically interact and change through a multi-hop reasoning process. Applied to the molecules, the dynamic interactions enable an iterative refinement of the representation of molecular graphs with multiple bond types. GraphMem is capable of jointly training on multiple datasets by using a specific-task query fed to the controller as an input. We demonstrate the effectiveness of the proposed model for separately and jointly training on more than 100K measurements, spanning across 9 BioAssay activity tests.
Knowledge Graph Embedding with Multiple Relation Projections
Jan 26, 2018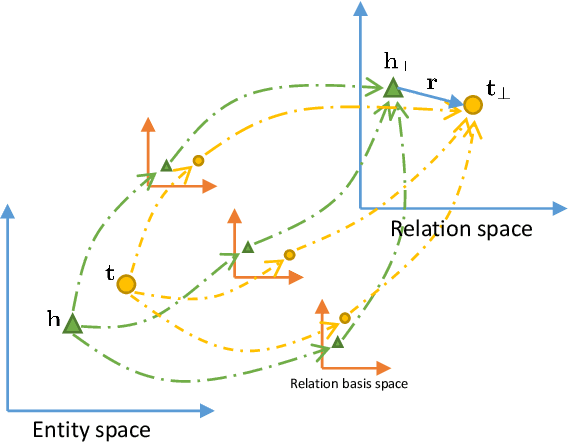
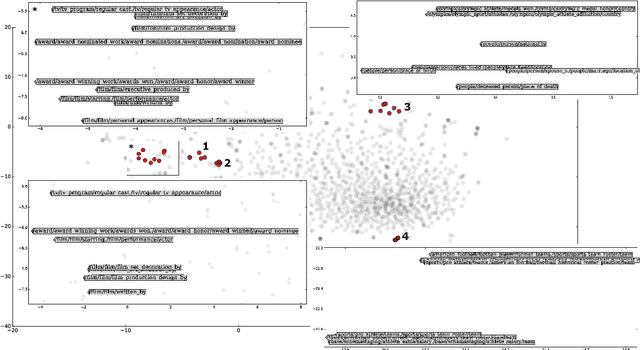
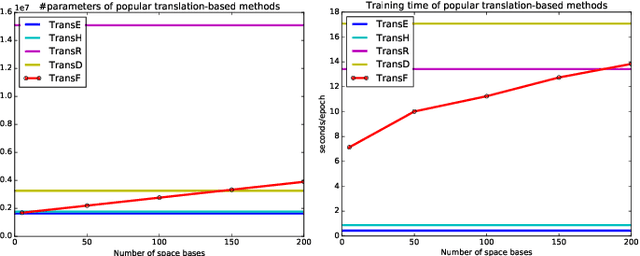
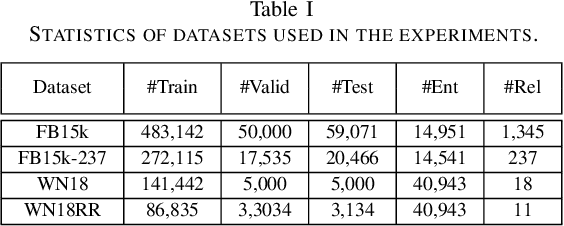
Abstract:Knowledge graphs contain rich relational structures of the world, and thus complement data-driven machine learning in heterogeneous data. One of the most effective methods in representing knowledge graphs is to embed symbolic relations and entities into continuous spaces, where relations are approximately linear translation between projected images of entities in the relation space. However, state-of-the-art relation projection methods such as TransR, TransD or TransSparse do not model the correlation between relations, and thus are not scalable to complex knowledge graphs with thousands of relations, both in computational demand and in statistical robustness. To this end we introduce TransF, a novel translation-based method which mitigates the burden of relation projection by explicitly modeling the basis subspaces of projection matrices. As a result, TransF is far more light weight than the existing projection methods, and is robust when facing a high number of relations. Experimental results on the canonical link prediction task show that our proposed model outperforms competing rivals by a large margin and achieves state-of-the-art performance. Especially, TransF improves by 9%/5% in the head/tail entity prediction task for N-to-1/1-to-N relations over the best performing translation-based method.
Finding Algebraic Structure of Care in Time: A Deep Learning Approach
Nov 21, 2017
Abstract:Understanding the latent processes from Electronic Medical Records could be a game changer in modern healthcare. However, the processes are complex due to the interaction between at least three dynamic components: the illness, the care and the recording practice. Existing methods are inadequate in capturing the dynamic structure of care. We propose an end-to-end model that reads medical record and predicts future risk. The model adopts the algebraic view in that discrete medical objects are embedded into continuous vectors lying in the same space. The bag of disease and comorbidities recorded at each hospital visit are modeled as function of sets. The same holds for the bag of treatments. The interaction between diseases and treatments at a visit is modeled as the residual of the diseases minus the treatments. Finally, the health trajectory, which is a sequence of visits, is modeled using a recurrent neural network. We report preliminary results on chronic diseases - diabetes and mental health - for predicting unplanned readmission.
Nonnegative Restricted Boltzmann Machines for Parts-based Representations Discovery and Predictive Model Stabilization
Aug 18, 2017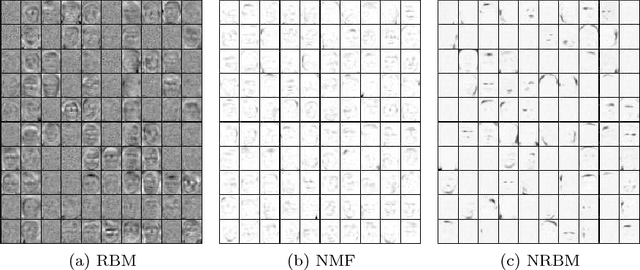
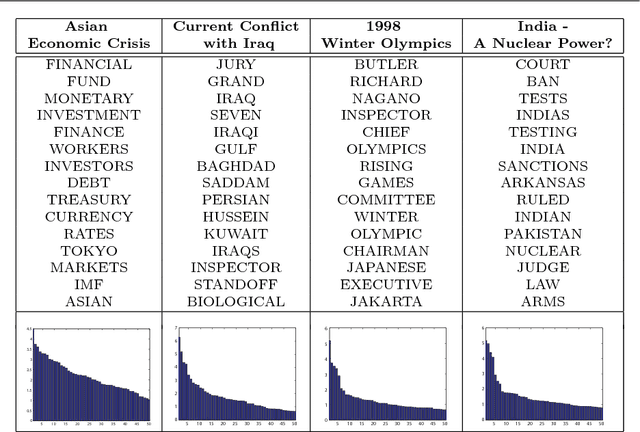
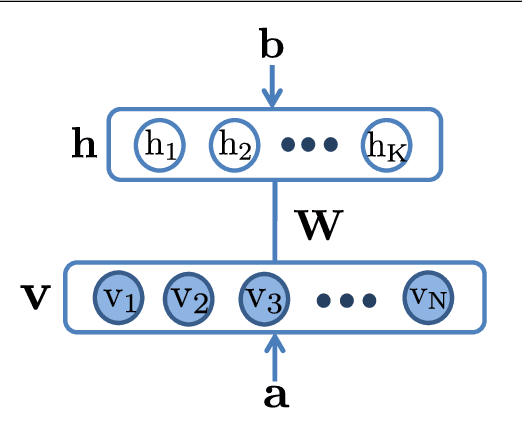
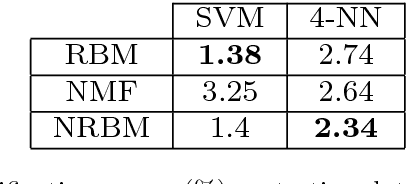
Abstract:The success of any machine learning system depends critically on effective representations of data. In many cases, it is desirable that a representation scheme uncovers the parts-based, additive nature of the data. Of current representation learning schemes, restricted Boltzmann machines (RBMs) have proved to be highly effective in unsupervised settings. However, when it comes to parts-based discovery, RBMs do not usually produce satisfactory results. We enhance such capacity of RBMs by introducing nonnegativity into the model weights, resulting in a variant called nonnegative restricted Boltzmann machine (NRBM). The NRBM produces not only controllable decomposition of data into interpretable parts but also offers a way to estimate the intrinsic nonlinear dimensionality of data, and helps to stabilize linear predictive models. We demonstrate the capacity of our model on applications such as handwritten digit recognition, face recognition, document classification and patient readmission prognosis. The decomposition quality on images is comparable with or better than what produced by the nonnegative matrix factorization (NMF), and the thematic features uncovered from text are qualitatively interpretable in a similar manner to that of the latent Dirichlet allocation (LDA). The stability performance of feature selection on medical data is better than RBM and competitive with NMF. The learned features, when used for classification, are more discriminative than those discovered by both NMF and LDA and comparable with those by RBM.
Statistical Latent Space Approach for Mixed Data Modelling and Applications
Aug 18, 2017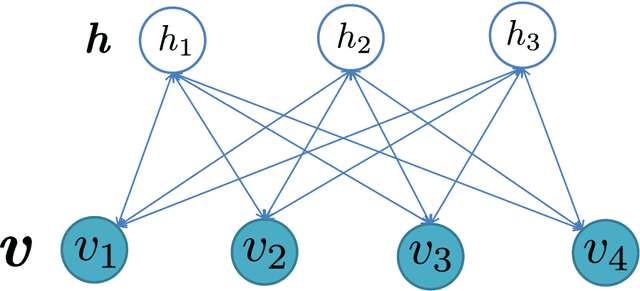
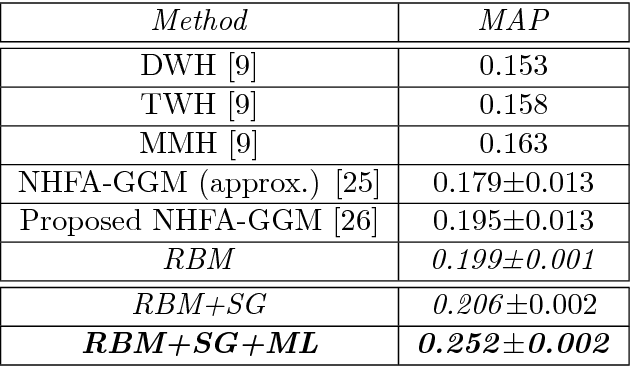
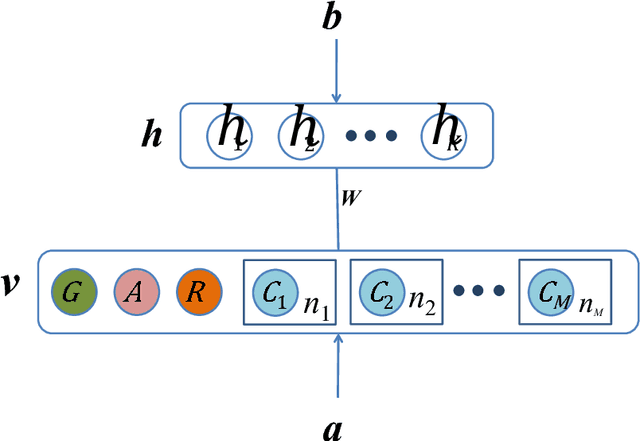
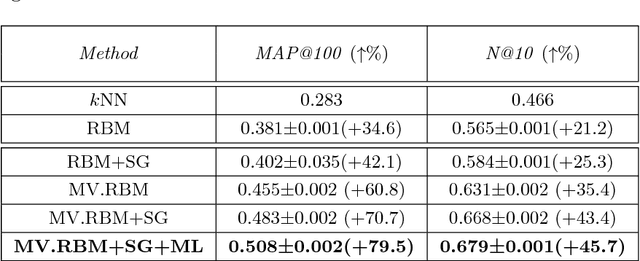
Abstract:The analysis of mixed data has been raising challenges in statistics and machine learning. One of two most prominent challenges is to develop new statistical techniques and methodologies to effectively handle mixed data by making the data less heterogeneous with minimum loss of information. The other challenge is that such methods must be able to apply in large-scale tasks when dealing with huge amount of mixed data. To tackle these challenges, we introduce parameter sharing and balancing extensions to our recent model, the mixed-variate restricted Boltzmann machine (MV.RBM) which can transform heterogeneous data into homogeneous representation. We also integrate structured sparsity and distance metric learning into RBM-based models. Our proposed methods are applied in various applications including latent patient profile modelling in medical data analysis and representation learning for image retrieval. The experimental results demonstrate the models perform better than baseline methods in medical data and outperform state-of-the-art rivals in image dataset.
Energy-based Models for Video Anomaly Detection
Aug 17, 2017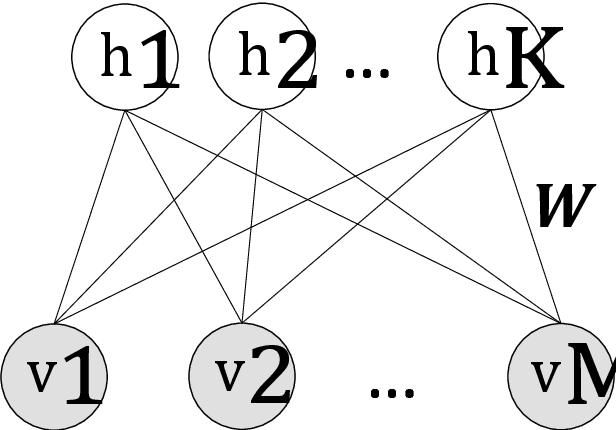
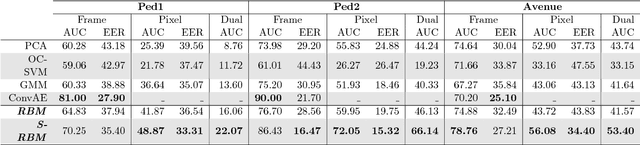
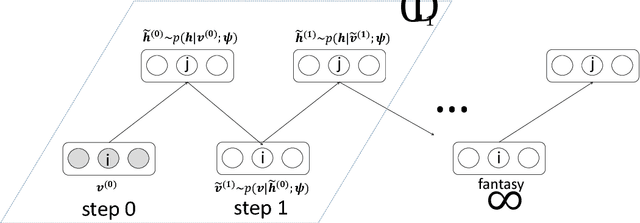
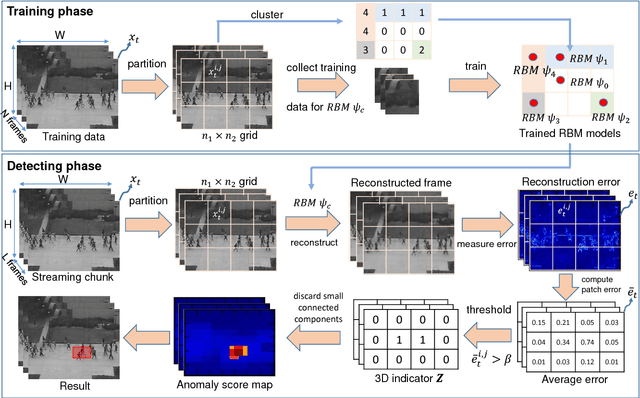
Abstract:Automated detection of abnormalities in data has been studied in research area in recent years because of its diverse applications in practice including video surveillance, industrial damage detection and network intrusion detection. However, building an effective anomaly detection system is a non-trivial task since it requires to tackle challenging issues of the shortage of annotated data, inability of defining anomaly objects explicitly and the expensive cost of feature engineering procedure. Unlike existing appoaches which only partially solve these problems, we develop a unique framework to cope the problems above simultaneously. Instead of hanlding with ambiguous definition of anomaly objects, we propose to work with regular patterns whose unlabeled data is abundant and usually easy to collect in practice. This allows our system to be trained completely in an unsupervised procedure and liberate us from the need for costly data annotation. By learning generative model that capture the normality distribution in data, we can isolate abnormal data points that result in low normality scores (high abnormality scores). Moreover, by leverage on the power of generative networks, i.e. energy-based models, we are also able to learn the feature representation automatically rather than replying on hand-crafted features that have been dominating anomaly detection research over many decades. We demonstrate our proposal on the specific application of video anomaly detection and the experimental results indicate that our method performs better than baselines and are comparable with state-of-the-art methods in many benchmark video anomaly detection datasets.
Graph Classification via Deep Learning with Virtual Nodes
Aug 14, 2017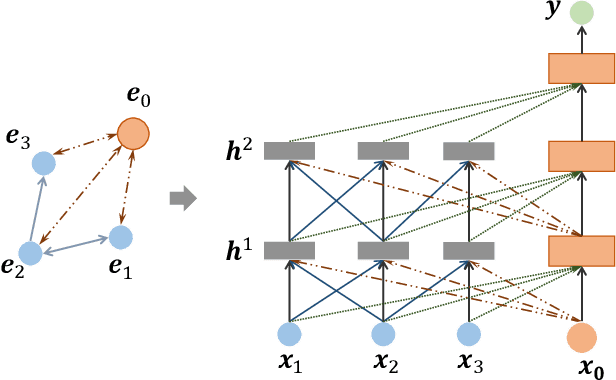

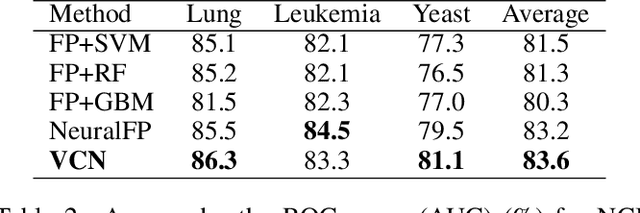
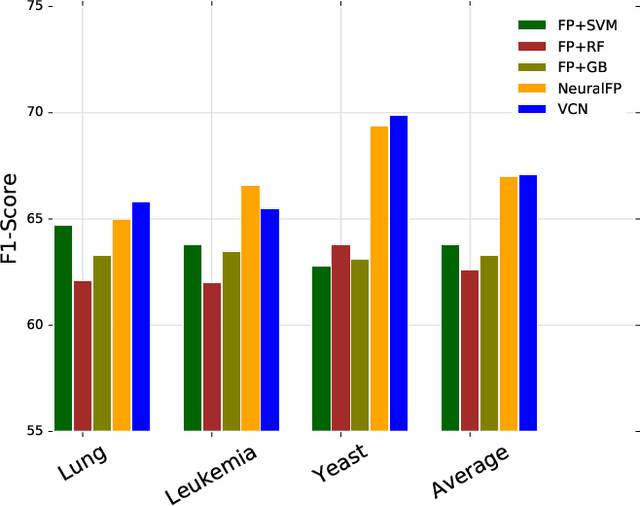
Abstract:Learning representation for graph classification turns a variable-size graph into a fixed-size vector (or matrix). Such a representation works nicely with algebraic manipulations. Here we introduce a simple method to augment an attributed graph with a virtual node that is bidirectionally connected to all existing nodes. The virtual node represents the latent aspects of the graph, which are not immediately available from the attributes and local connectivity structures. The expanded graph is then put through any node representation method. The representation of the virtual node is then the representation of the entire graph. In this paper, we use the recently introduced Column Network for the expanded graph, resulting in a new end-to-end graph classification model dubbed Virtual Column Network (VCN). The model is validated on two tasks: (i) predicting bio-activity of chemical compounds, and (ii) finding software vulnerability from source code. Results demonstrate that VCN is competitive against well-established rivals.
Deep Learning to Attend to Risk in ICU
Jul 17, 2017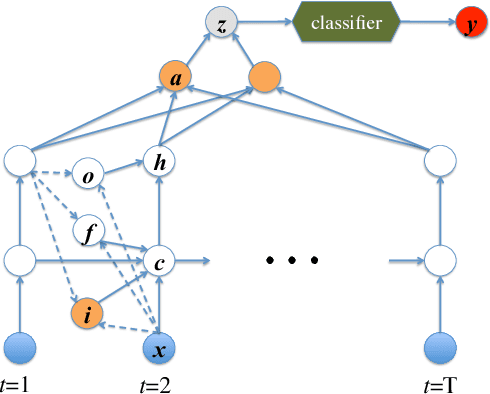
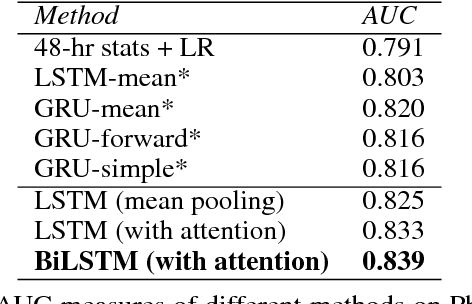
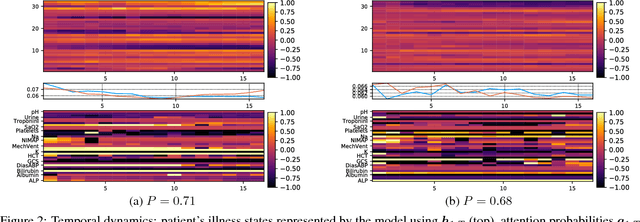
Abstract:Modeling physiological time-series in ICU is of high clinical importance. However, data collected within ICU are irregular in time and often contain missing measurements. Since absence of a measure would signify its lack of importance, the missingness is indeed informative and might reflect the decision making by the clinician. Here we propose a deep learning architecture that can effectively handle these challenges for predicting ICU mortality outcomes. The model is based on Long Short-Term Memory, and has layered attention mechanisms. At the sensing layer, the model decides whether to observe and incorporate parts of the current measurements. At the reasoning layer, evidences across time steps are weighted and combined. The model is evaluated on the PhysioNet 2012 dataset showing competitive and interpretable results.
Budgeted Batch Bayesian Optimization With Unknown Batch Sizes
Apr 15, 2017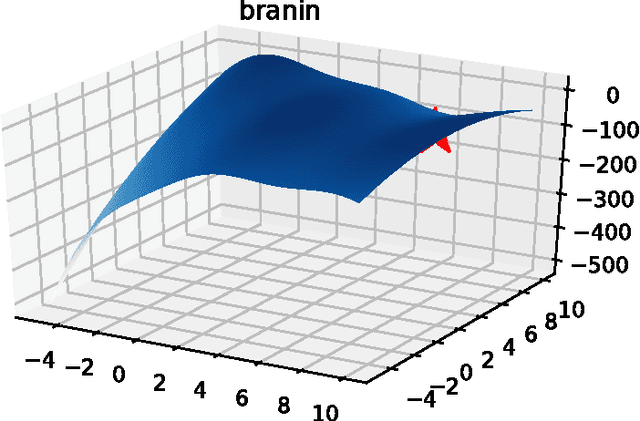
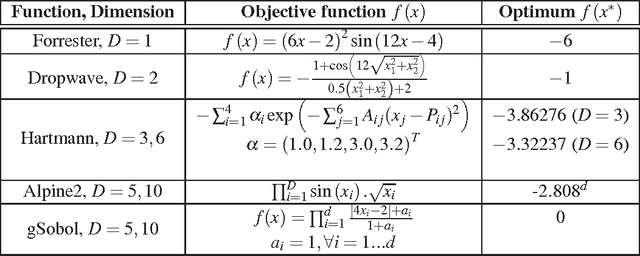

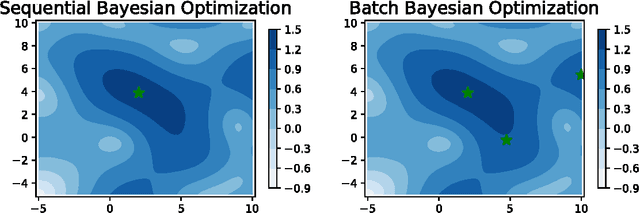
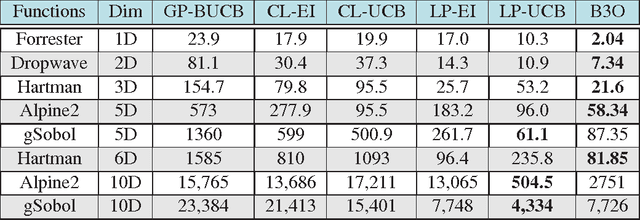
Abstract:Parameter settings profoundly impact the performance of machine learning algorithms and laboratory experiments. The classical grid search or trial-error methods are exponentially expensive in large parameter spaces, and Bayesian optimization (BO) offers an elegant alternative for global optimization of black box functions. In situations where the black box function can be evaluated at multiple points simultaneously, batch Bayesian optimization is used. Current batch BO approaches are restrictive in that they fix the number of evaluations per batch, and this can be wasteful when the number of specified evaluations is larger than the number of real maxima in the underlying acquisition function. We present the Budgeted Batch Bayesian Optimization (B3O) for hyper-parameter tuning and experimental design - we identify the appropriate batch size for each iteration in an elegant way. To set the batch size flexible, we use the infinite Gaussian mixture model (IGMM) for automatically identifying the number of peaks in the underlying acquisition functions. We solve the intractability of estimating the IGMM directly from the acquisition function by formulating the batch generalized slice sampling to efficiently draw samples from the acquisition function. We perform extensive experiments for both synthetic functions and two real world applications - machine learning hyper-parameter tuning and experimental design for alloy hardening. We show empirically that the proposed B3O outperforms the existing fixed batch BO approaches in finding the optimum whilst requiring a fewer number of evaluations, thus saving cost and time.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge