Svetha Venkatesh
Bayesian functional optimisation with shape prior
Sep 19, 2018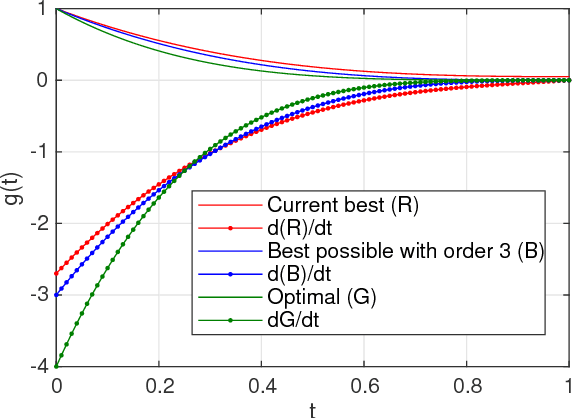
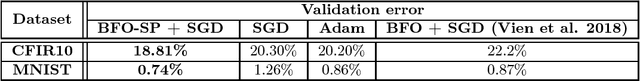
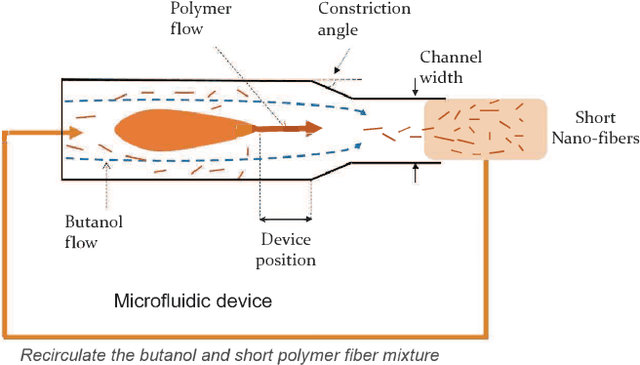
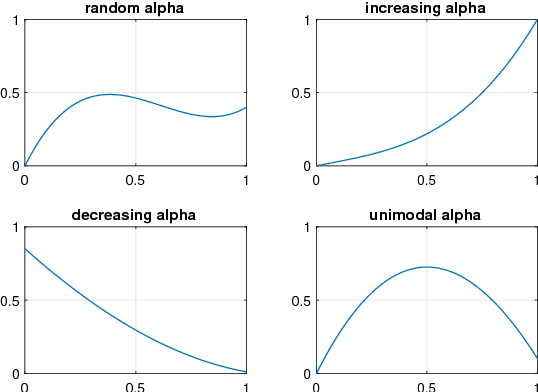
Abstract:Real world experiments are expensive, and thus it is important to reach a target in minimum number of experiments. Experimental processes often involve control variables that changes over time. Such problems can be formulated as a functional optimisation problem. We develop a novel Bayesian optimisation framework for such functional optimisation of expensive black-box processes. We represent the control function using Bernstein polynomial basis and optimise in the coefficient space. We derive the theory and practice required to dynamically adjust the order of the polynomial degree, and show how prior information about shape can be integrated. We demonstrate the effectiveness of our approach for short polymer fibre design and optimising learning rate schedules for deep networks.
On catastrophic forgetting and mode collapse in Generative Adversarial Networks
Sep 12, 2018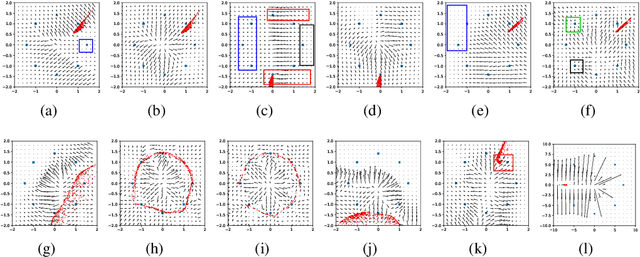
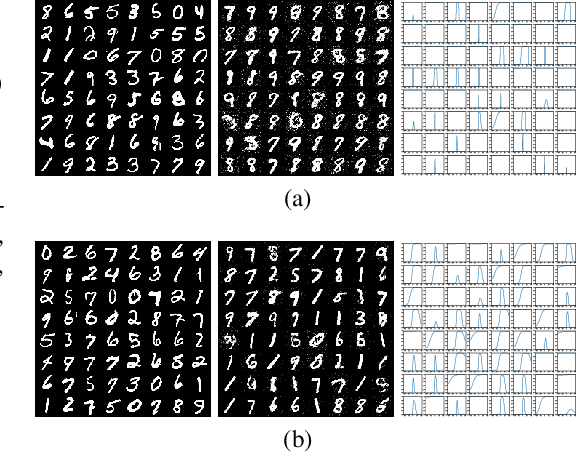
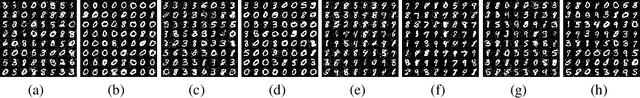
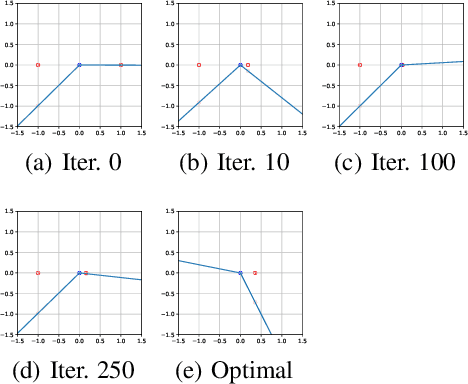


Abstract:Generative Adversarial Networks (GAN) are one of the most prominent tools for learning complicated distributions. However, problems such as mode collapse and catastrophic forgetting, prevent GAN from learning the target distribution. These problems are usually studied independently from each other. In this paper, we show that both problems are present in GAN and their combined effect makes the training of GAN unstable. We also show that methods such as gradient penalties and momentum based optimizers can improve the stability of GAN by effectively preventing these problems from happening. Finally, we study a mechanism for mode collapse to occur and propagate in feedforward neural networks.
Kernel Pre-Training in Feature Space via m-Kernels
May 21, 2018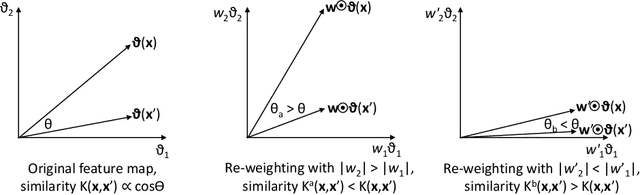
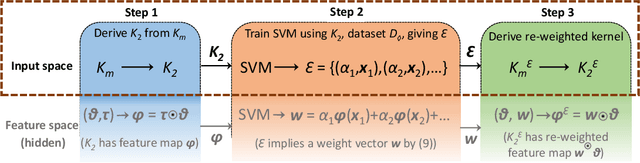


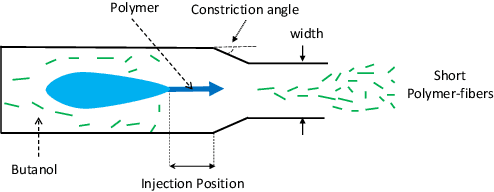
Abstract:This paper presents a novel approach to kernel tuning. The method presented borrows techniques from reproducing kernel Banach space (RKBS) theory and tensor kernels and leverages them to convert (re-weight in feature space) existing kernel functions into new, problem-specific kernels using auxiliary data. The proposed method is applied to accelerating Bayesian optimisation via covariance (kernel) function pre-tuning for short-polymer fibre manufacture and alloy design.
Attentional Multilabel Learning over Graphs: A Message Passing Approach
Apr 11, 2018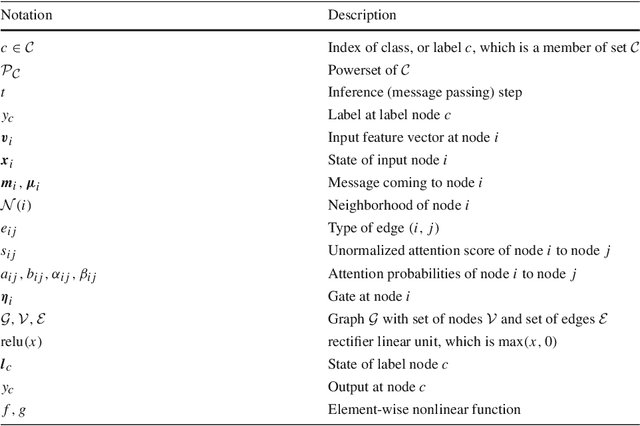
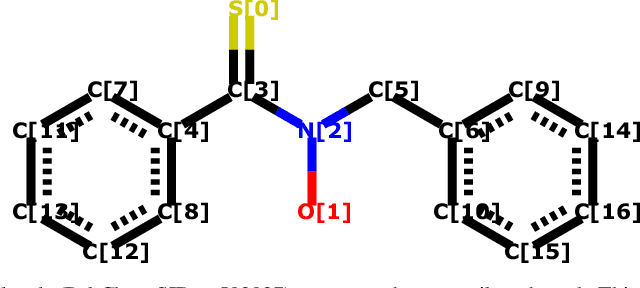
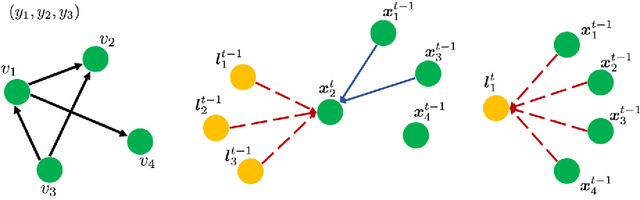
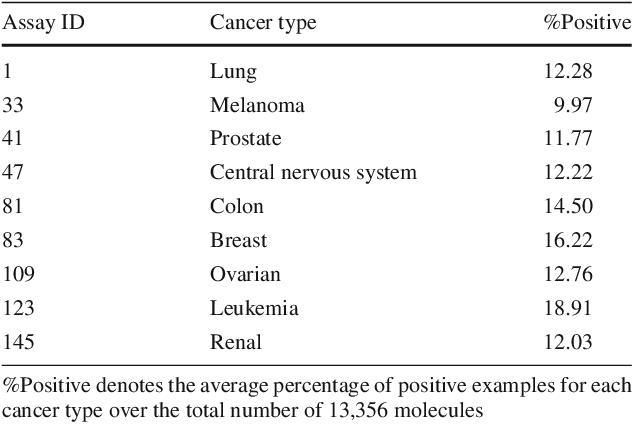
Abstract:We address a largely open problem of multilabel classification over graphs. Unlike traditional vector input, a graph has rich variable-size substructures which are related to the labels in some ways. We believe that uncovering these relations might hold the key to classification performance and explainability. We introduce GAML (Graph Attentional Multi-Label learning), a novel graph neural network that can handle this problem effectively. GAML regards labels as auxiliary nodes and models them in conjunction with the input graph. By applying message passing and attention mechanisms to both the label nodes and the input nodes iteratively, GAML can capture the relations between the labels and the input subgraphs at various resolution scales. Moreover, our model can take advantage of explicit label dependencies. It also scales linearly with the number of labels and graph size thanks to our proposed hierarchical attention. We evaluate GAML on an extensive set of experiments with both graph-structured inputs and classical unstructured inputs. The results show that GAML significantly outperforms other competing methods. Importantly, GAML enables intuitive visualizations for better understanding of the label-substructure relations and explanation of the model behaviors.
Covariance Function Pre-Training with m-Kernels for Accelerated Bayesian Optimisation
Mar 13, 2018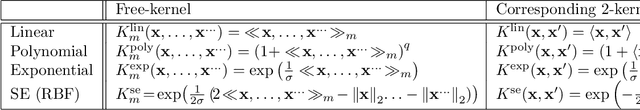
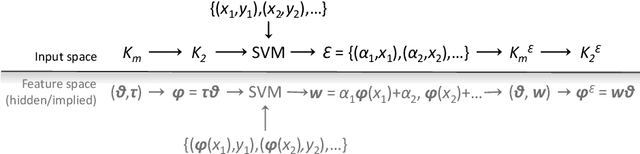
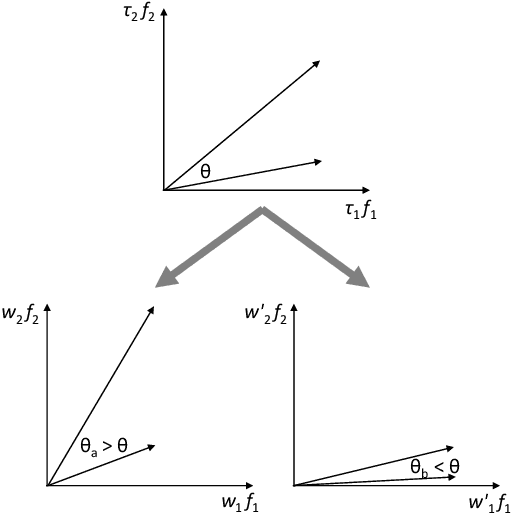

Abstract:The paper presents a novel approach to direct covariance function learning for Bayesian optimisation, with particular emphasis on experimental design problems where an existing corpus of condensed knowledge is present. The method presented borrows techniques from reproducing kernel Banach space theory (specifically m-kernels) and leverages them to convert (or re-weight) existing covariance functions into new, problem-specific covariance functions. The key advantage of this approach is that rather than relying on the user to manually select (with some hyperparameter tuning and experimentation) an appropriate covariance function it constructs the covariance function to specifically match the problem at hand. The technique is demonstrated on two real-world problems - specifically alloy design and short-polymer fibre manufacturing - as well as a selected test function.
Rapid Bayesian optimisation for synthesis of short polymer fiber materials
Feb 16, 2018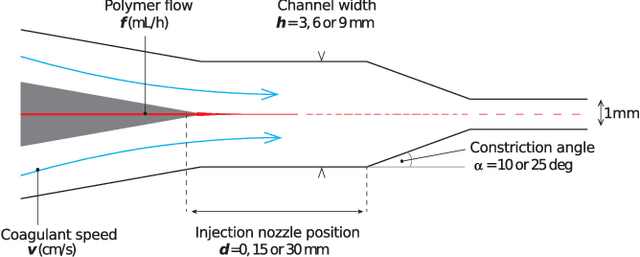
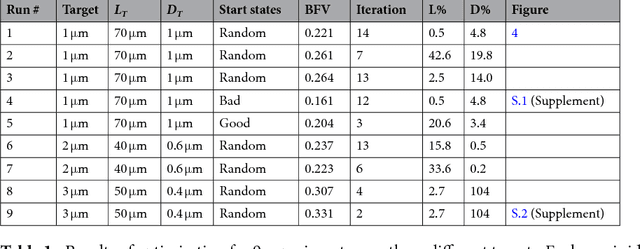
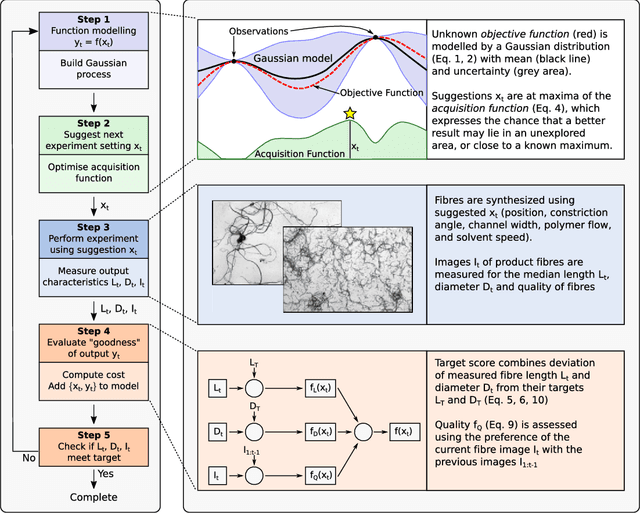

Abstract:The discovery of processes for the synthesis of new materials involves many decisions about process design, operation, and material properties. Experimentation is crucial but as complexity increases, exploration of variables can become impractical using traditional combinatorial approaches. We describe an iterative method which uses machine learning to optimise process development, incorporating multiple qualitative and quantitative objectives. We demonstrate the method with a novel fluid processing platform for synthesis of short polymer fibers, and show how the synthesis process can be efficiently directed to achieve material and process objectives.
High Dimensional Bayesian Optimization Using Dropout
Feb 15, 2018
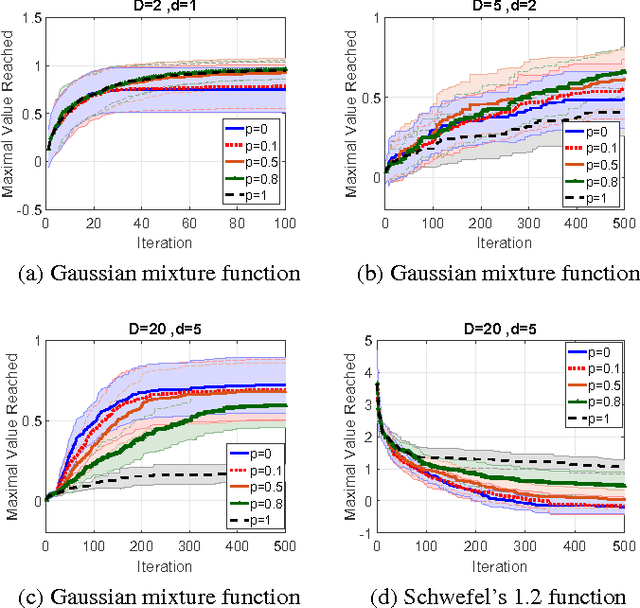
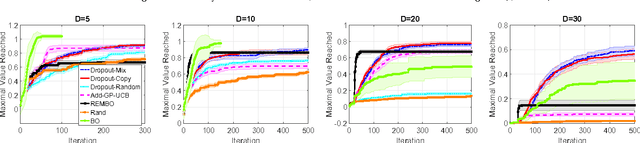
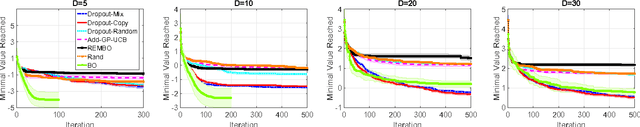
Abstract:Scaling Bayesian optimization to high dimensions is challenging task as the global optimization of high-dimensional acquisition function can be expensive and often infeasible. Existing methods depend either on limited active variables or the additive form of the objective function. We propose a new method for high-dimensional Bayesian optimization, that uses a dropout strategy to optimize only a subset of variables at each iteration. We derive theoretical bounds for the regret and show how it can inform the derivation of our algorithm. We demonstrate the efficacy of our algorithms for optimization on two benchmark functions and two real-world applications- training cascade classifiers and optimizing alloy composition.
Dual Control Memory Augmented Neural Networks for Treatment Recommendations
Feb 11, 2018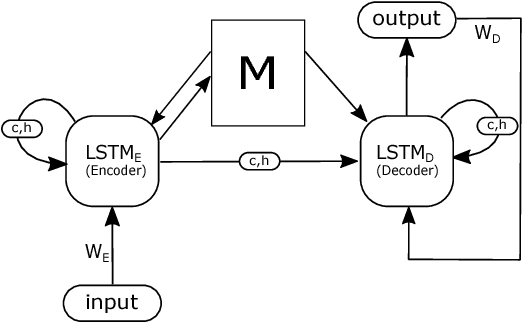
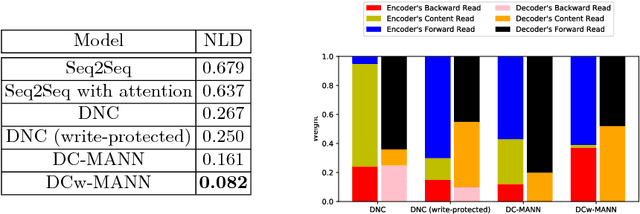
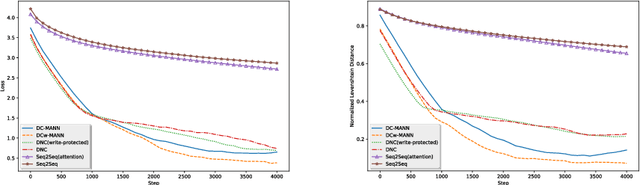
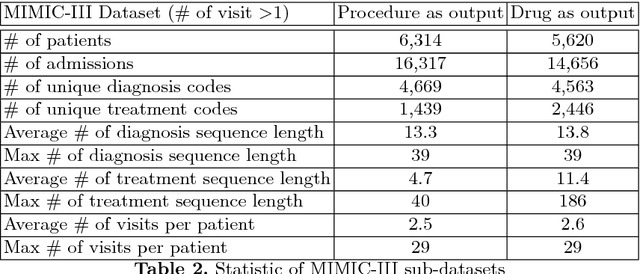
Abstract:Machine-assisted treatment recommendations hold a promise to reduce physician time and decision errors. We formulate the task as a sequence-to-sequence prediction model that takes the entire time-ordered medical history as input, and predicts a sequence of future clinical procedures and medications. It is built on the premise that an effective treatment plan may have long-term dependencies from previous medical history. We approach the problem by using a memory-augmented neural network, in particular, by leveraging the recent differentiable neural computer that consists of a neural controller and an external memory module. But differing from the original model, we use dual controllers, one for encoding the history followed by another for decoding the treatment sequences. In the encoding phase, the memory is updated as new input is read; at the end of this phase, the memory holds not only the medical history but also the information about the current illness. During the decoding phase, the memory is write-protected. The decoding controller generates a treatment sequence, one treatment option at a time. The resulting dual controller write-protected memory-augmented neural network is demonstrated on the MIMIC-III dataset on two tasks: procedure prediction and medication prescription. The results show improved performance over both traditional bag-of-words and sequence-to-sequence methods.
Dual Memory Neural Computer for Asynchronous Two-view Sequential Learning
Feb 11, 2018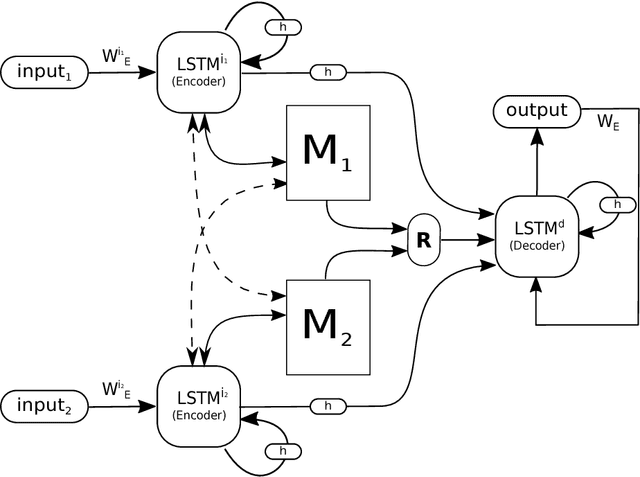
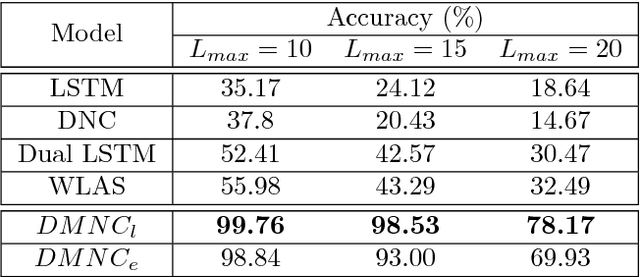
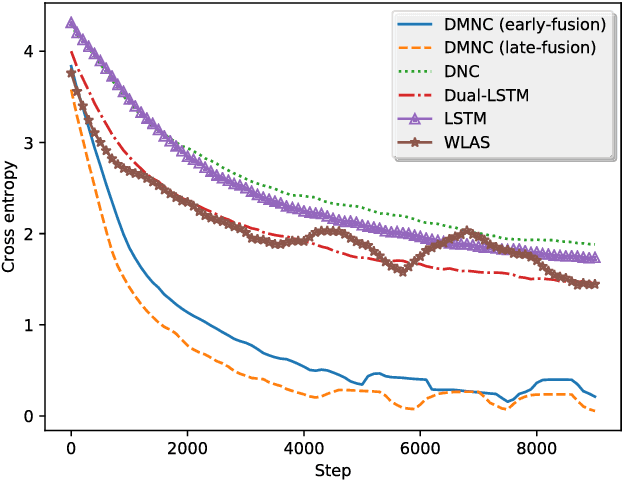
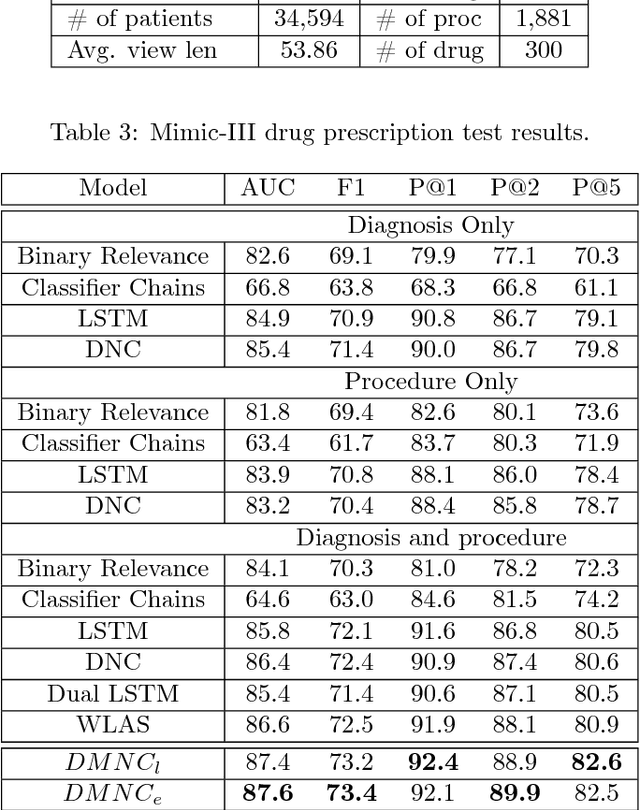
Abstract:One of the core tasks in multi-view learning is to capture relations among views. For sequential data, the relations not only span across views, but also extend throughout the view length to form long-term intra-view and inter-view interactions. In this paper, we present a new memory augmented neural network model that aims to model these complex interactions between two asynchronous sequential views. Our model uses two encoders for reading from and writing to two external memories for encoding input views. The intra-view interactions and the long-term dependencies are captured by the use of memories during this encoding process. There are two modes of memory accessing in our system: late-fusion and early-fusion, corresponding to late and early inter-view interactions. In the late-fusion mode, the two memories are separated, containing only view-specific contents. In the early-fusion mode, the two memories share the same addressing space, allowing cross-memory accessing. In both cases, the knowledge from the memories will be combined by a decoder to make predictions over the output space. The resulting dual memory neural computer is demonstrated on a comprehensive set of experiments, including a synthetic task of summing two sequences and the tasks of drug prescription and disease progression in healthcare. The results demonstrate competitive performance over both traditional algorithms and deep learning methods designed for multi-view problems.
Learning Deep Matrix Representations
Feb 05, 2018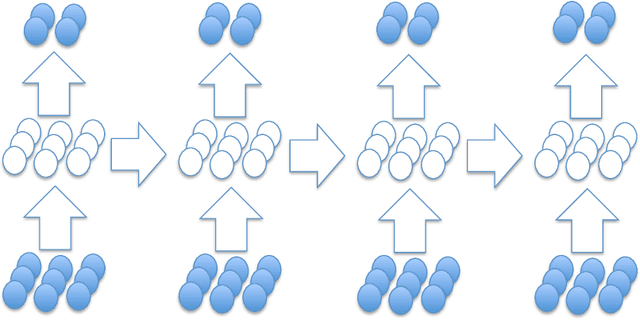
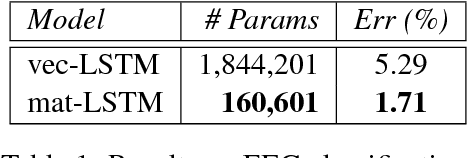
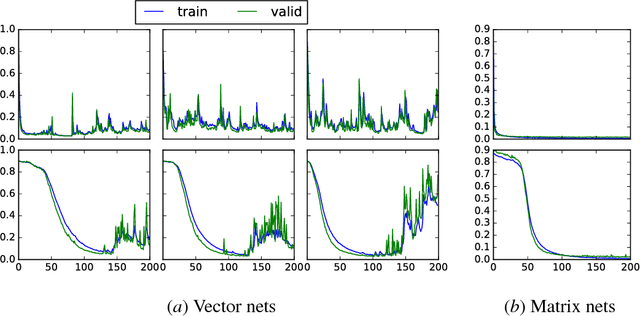
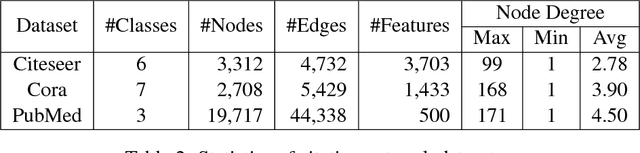
Abstract:We present a new distributed representation in deep neural nets wherein the information is represented in native form as a matrix. This differs from current neural architectures that rely on vector representations. We consider matrices as central to the architecture and they compose the input, hidden and output layers. The model representation is more compact and elegant -- the number of parameters grows only with the largest dimension of the incoming layer rather than the number of hidden units. We derive several new deep networks: (i) feed-forward nets that map an input matrix into an output matrix, (ii) recurrent nets which map a sequence of input matrices into a sequence of output matrices. We also reinterpret existing models for (iii) memory-augmented networks and (iv) graphs using matrix notations. For graphs we demonstrate how the new notations lead to simple but effective extensions with multiple attentions. Extensive experiments on handwritten digits recognition, face reconstruction, sequence to sequence learning, EEG classification, and graph-based node classification demonstrate the efficacy and compactness of the matrix architectures.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge