Svetha Venkatesh
Learning Regularity in Skeleton Trajectories for Anomaly Detection in Videos
Mar 08, 2019
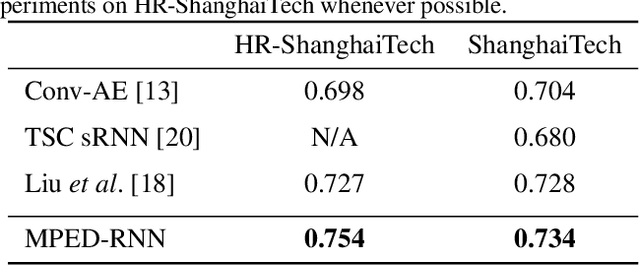
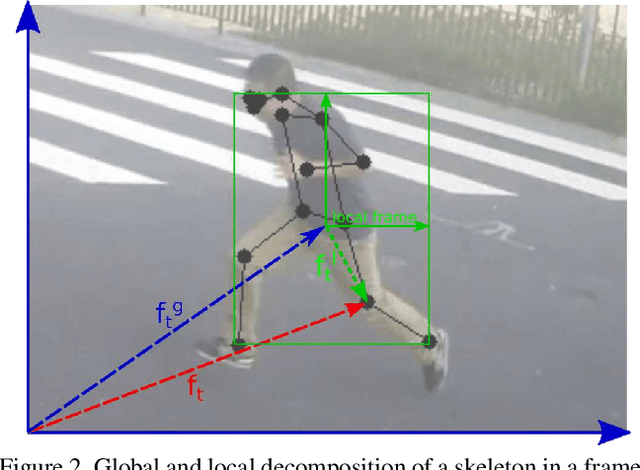
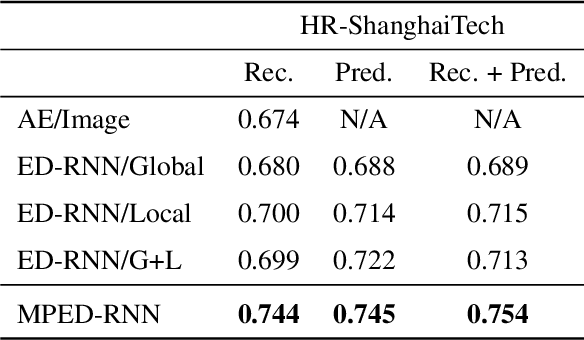
Abstract:Appearance features have been widely used in video anomaly detection even though they contain complex entangled factors. We propose a new method to model the normal patterns of human movements in surveillance video for anomaly detection using dynamic skeleton features. We decompose the skeletal movements into two sub-components: global body movement and local body posture. We model the dynamics and interaction of the coupled features in our novel Message-Passing Encoder-Decoder Recurrent Network. We observed that the decoupled features collaboratively interact in our spatio-temporal model to accurately identify human-related irregular events from surveillance video sequences. Compared to traditional appearance-based models, our method achieves superior outlier detection performance. Our model also offers "open-box" examination and decision explanation made possible by the semantically understandable features and a network architecture supporting interpretability.
Stable Bayesian Optimisation via Direct Stability Quantification
Feb 21, 2019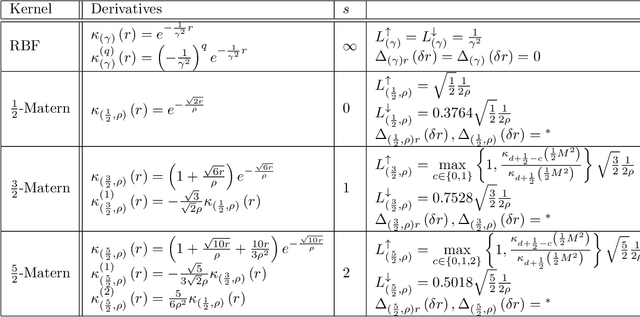
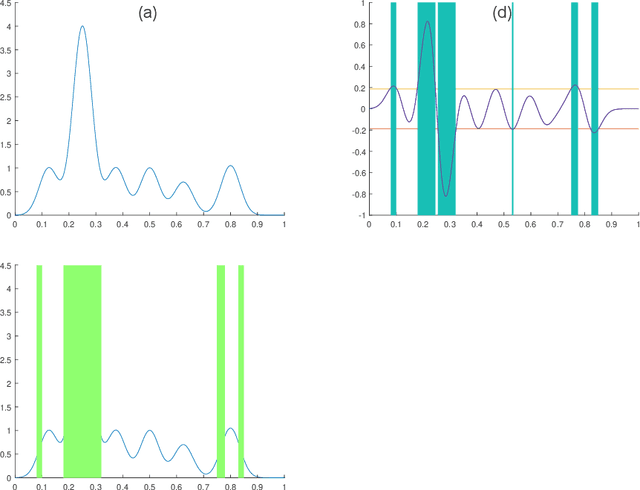
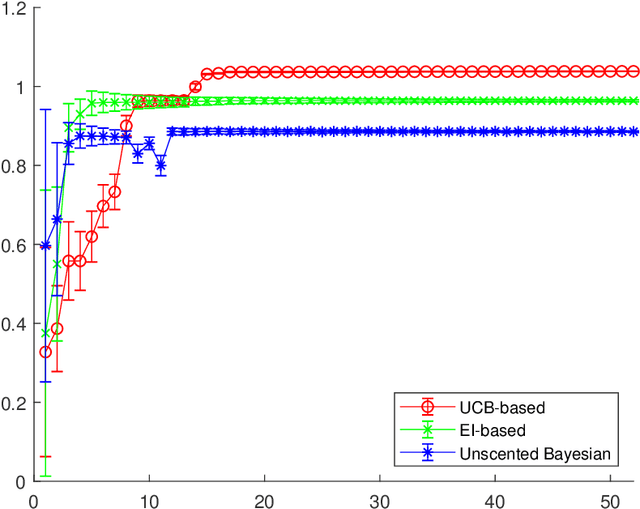
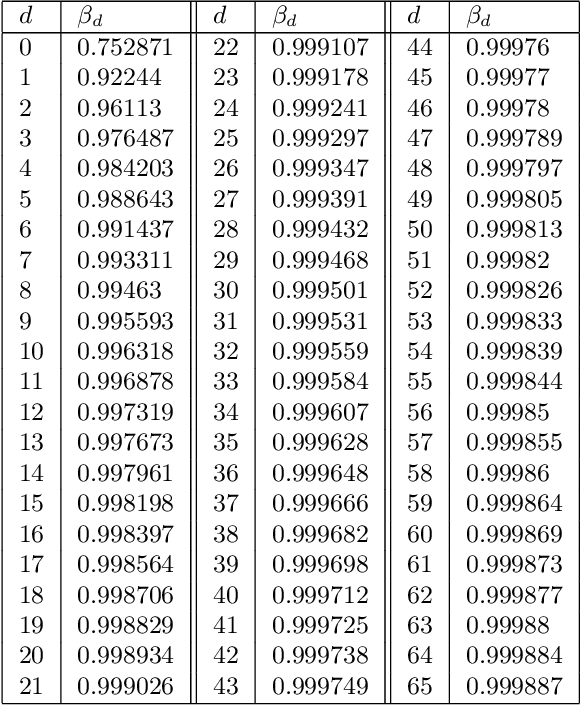
Abstract:In this paper we consider the problem of finding stable maxima of expensive (to evaluate) functions. We are motivated by the optimisation of physical and industrial processes where, for some input ranges, small and unavoidable variations in inputs lead to unacceptably large variation in outputs. Our approach uses multiple gradient Gaussian Process models to estimate the probability that worst-case output variation for specified input perturbation exceeded the desired maxima, and these probabilities are then used to (a) guide the optimisation process toward solutions satisfying our stability criteria and (b) post-filter results to find the best stable solution. We exhibit our algorithm on synthetic and real-world problems and demonstrate that it is able to effectively find stable maxima.
Multi-objective Bayesian optimisation with preferences over objectives
Feb 12, 2019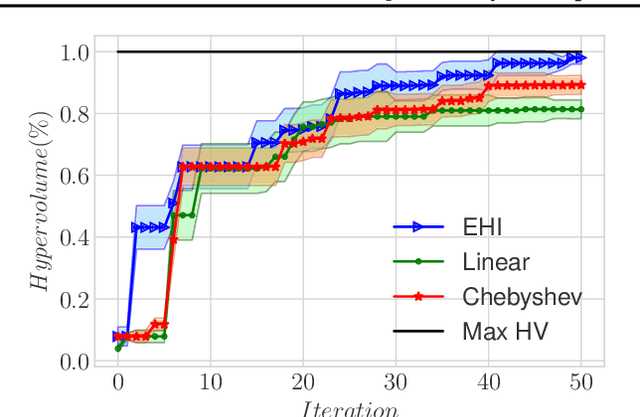
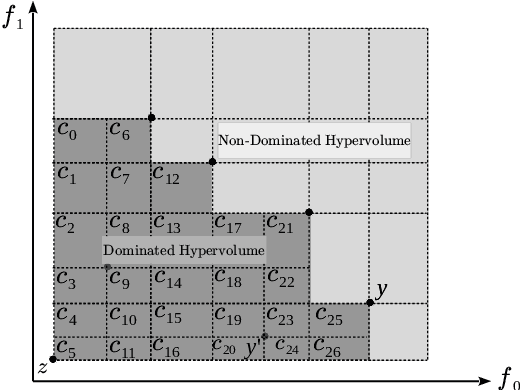
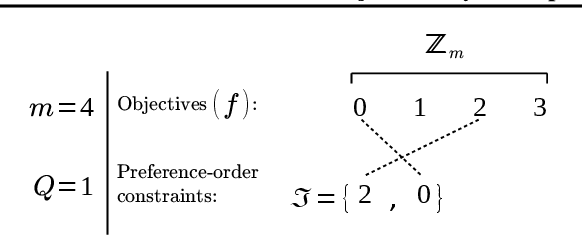


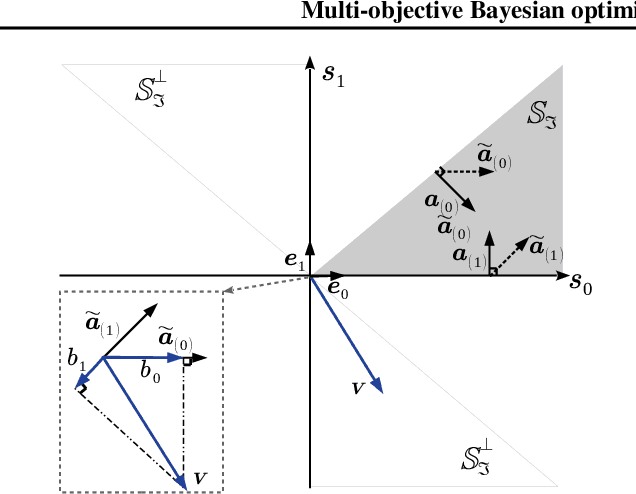
Abstract:We present a Bayesian multi-objective optimisation algorithm that allows the user to express preference-order constraints on the objectives of the type `objective A is more important than objective B'. Rather than attempting to find a representative subset of the complete Pareto front, our algorithm searches for and returns only those Pareto-optimal points that satisfy these constraints. We formulate a new acquisition function based on expected improvement in dominated hypervolume (EHI) to ensure that the subset of Pareto front satisfying the constraints is thoroughly explored. The hypervolume calculation only includes those points that satisfy the preference-order constraints, where the probability of a point satisfying the constraints is calculated from a gradient Gaussian Process model. We demonstrate our algorithm on both synthetic and real-world problems.
Improving Generalization and Stability of Generative Adversarial Networks
Feb 11, 2019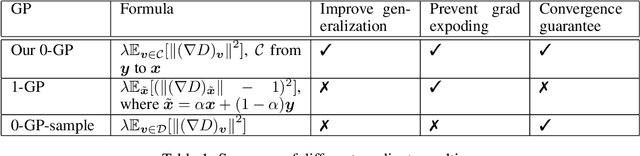
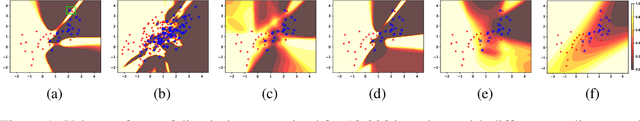
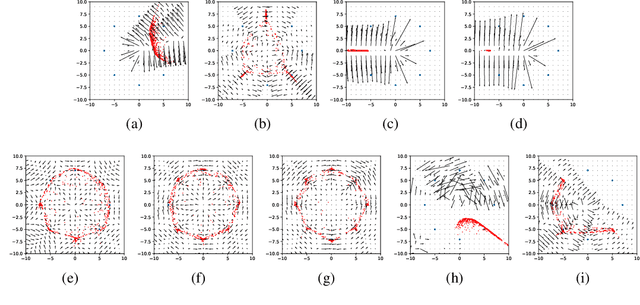

Abstract:Generative Adversarial Networks (GANs) are one of the most popular tools for learning complex high dimensional distributions. However, generalization properties of GANs have not been well understood. In this paper, we analyze the generalization of GANs in practical settings. We show that discriminators trained on discrete datasets with the original GAN loss have poor generalization capability and do not approximate the theoretically optimal discriminator. We propose a zero-centered gradient penalty for improving the generalization of the discriminator by pushing it toward the optimal discriminator. The penalty guarantees the generalization and convergence of GANs. Experiments on synthetic and large scale datasets verify our theoretical analysis.
Fast Hyperparameter Tuning using Bayesian Optimization with Directional Derivatives
Feb 06, 2019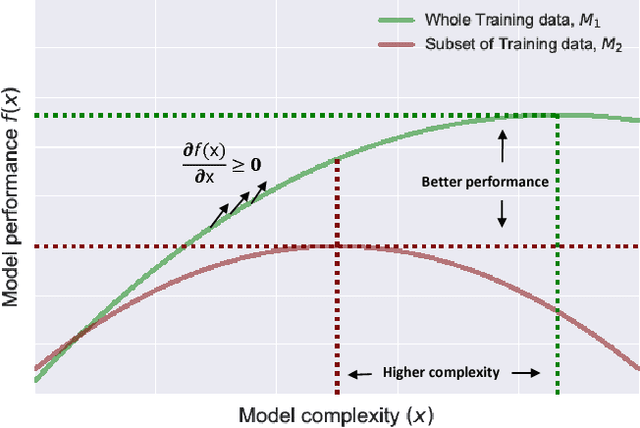

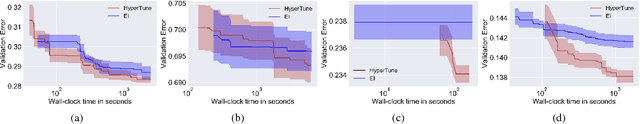

Abstract:In this paper we develop a Bayesian optimization based hyperparameter tuning framework inspired by statistical learning theory for classifiers. We utilize two key facts from PAC learning theory; the generalization bound will be higher for a small subset of data compared to the whole, and the highest accuracy for a small subset of data can be achieved with a simple model. We initially tune the hyperparameters on a small subset of training data using Bayesian optimization. While tuning the hyperparameters on the whole training data, we leverage the insights from the learning theory to seek more complex models. We realize this by using directional derivative signs strategically placed in the hyperparameter search space to seek a more complex model than the one obtained with small data. We demonstrate the performance of our method on the tasks of tuning the hyperparameters of several machine learning algorithms.
Learning to Remember More with Less Memorization
Jan 05, 2019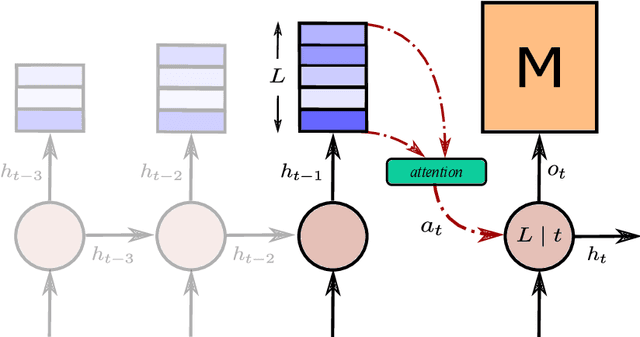
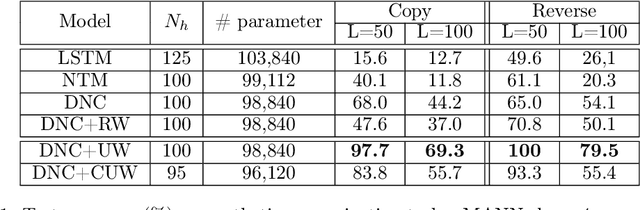
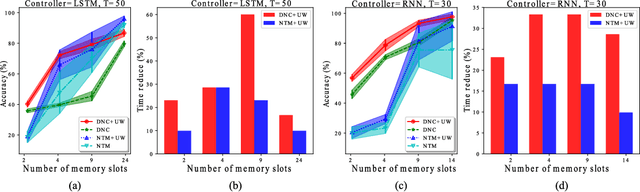
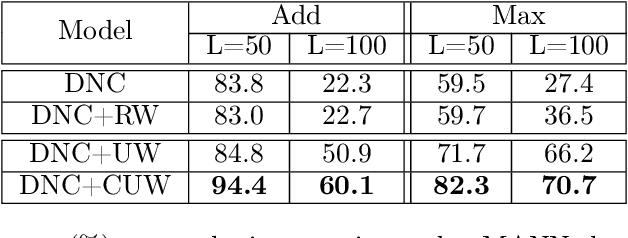
Abstract:Memory-augmented neural networks consisting of a neural controller and an external memory have shown potentials in long-term sequential learning. Current RAM-like memory models maintain memory accessing every timesteps, thus they do not effectively leverage the short-term memory held in the controller. We hypothesize that this scheme of writing is suboptimal in memory utilization and introduces redundant computation. To validate our hypothesis, we derive a theoretical bound on the amount of information stored in a RAM-like system and formulate an optimization problem that maximizes the bound. The proposed solution dubbed Uniform Writing is proved to be optimal under the assumption of equal timestep contributions. To relax this assumption, we introduce modifications to the original solution, resulting in a solution termed Cached Uniform Writing. This method aims to balance between maximizing memorization and forgetting via overwriting mechanisms. Through an extensive set of experiments, we empirically demonstrate the advantages of our solutions over other recurrent architectures, claiming the state-of-the-arts in various sequential modeling tasks.
Graph Transformation Policy Network for Chemical Reaction Prediction
Dec 22, 2018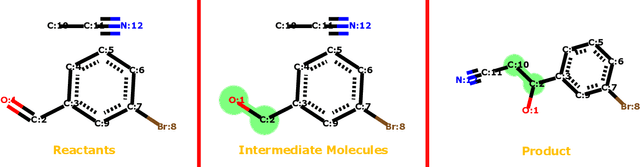
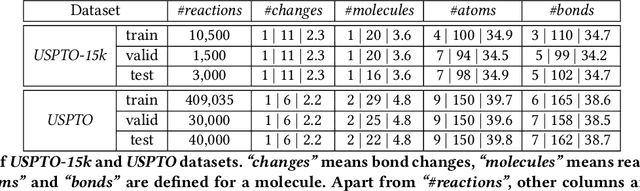
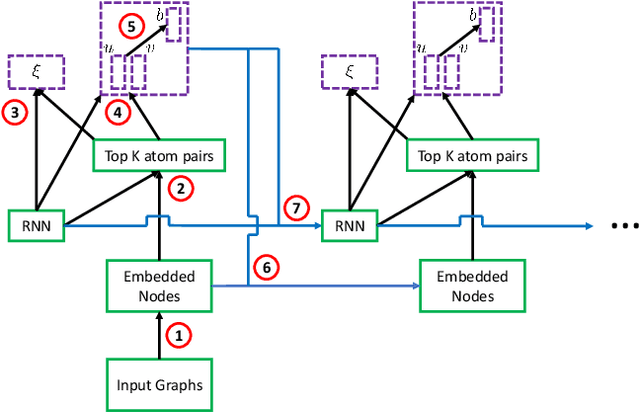
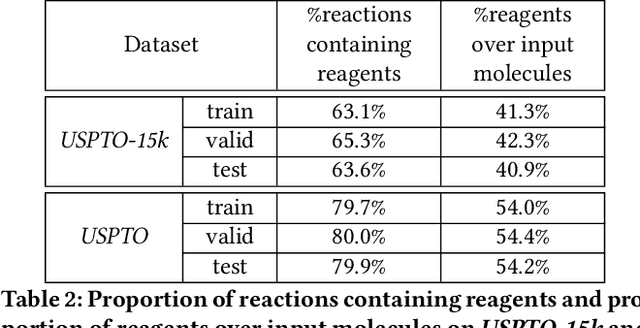
Abstract:We address a fundamental problem in chemistry known as chemical reaction product prediction. Our main insight is that the input reactant and reagent molecules can be jointly represented as a graph, and the process of generating product molecules from reactant molecules can be formulated as a sequence of graph transformations. To this end, we propose Graph Transformation Policy Network (GTPN) -- a novel generic method that combines the strengths of graph neural networks and reinforcement learning to learn the reactions directly from data with minimal chemical knowledge. Compared to previous methods, GTPN has some appealing properties such as: end-to-end learning, and making no assumption about the length or the order of graph transformations. In order to guide model search through the complex discrete space of sets of bond changes effectively, we extend the standard policy gradient loss by adding useful constraints. Evaluation results show that GTPN improves the top-1 accuracy over the current state-of-the-art method by about 3% on the large USPTO dataset. Our model's performances and prediction errors are also analyzed carefully in the paper.
Practical Batch Bayesian Optimization for Less Expensive Functions
Nov 05, 2018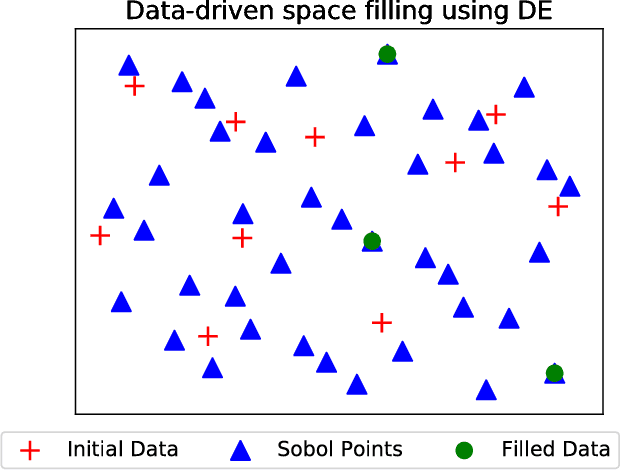
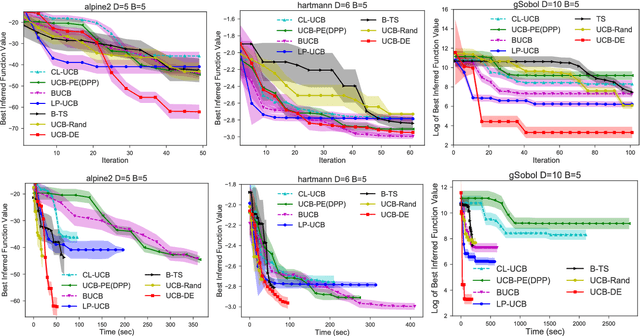
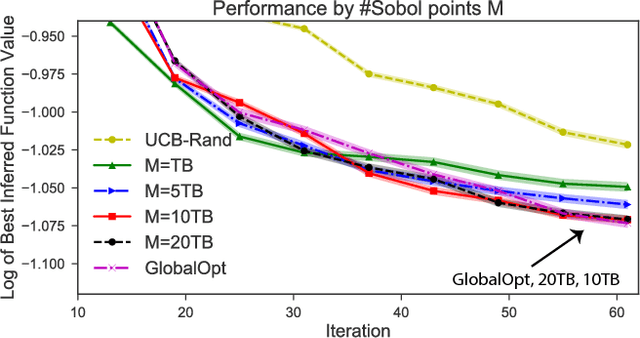
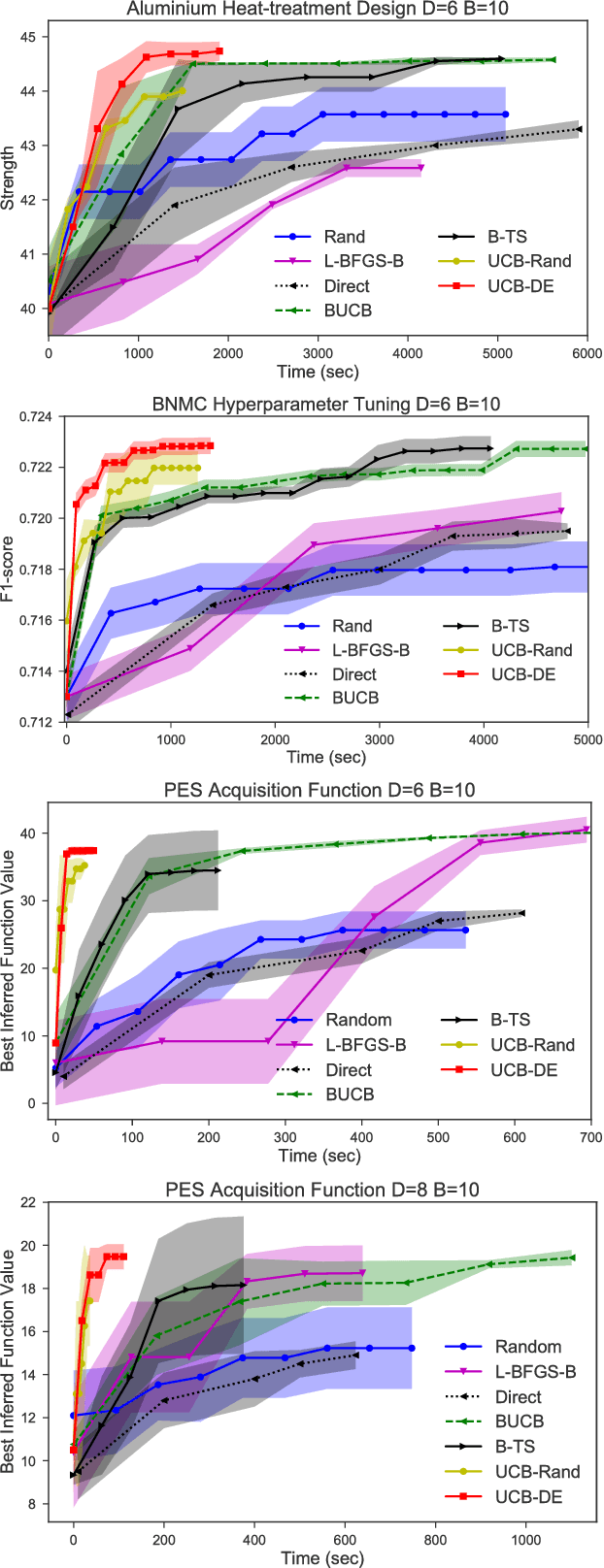
Abstract:Bayesian optimization (BO) and its batch extensions are successful for optimizing expensive black-box functions. However, these traditional BO approaches are not yet ideal for optimizing less expensive functions when the computational cost of BO can dominate the cost of evaluating the blackbox function. Examples of these less expensive functions are cheap machine learning models, inexpensive physical experiment through simulators, and acquisition function optimization in Bayesian optimization. In this paper, we consider a batch BO setting for situations where function evaluations are less expensive. Our model is based on a new exploration strategy using geometric distance that provides an alternative way for exploration, selecting a point far from the observed locations. Using that intuition, we propose to use Sobol sequence to guide exploration that will get rid of running multiple global optimization steps as used in previous works. Based on the proposed distance exploration, we present an efficient batch BO approach. We demonstrate that our approach outperforms other baselines and global optimization methods when the function evaluations are less expensive.
Relational dynamic memory networks
Oct 30, 2018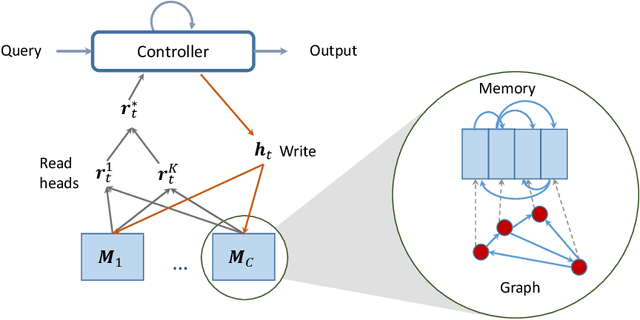
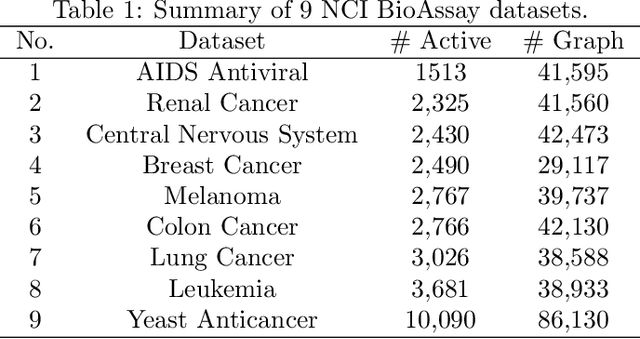
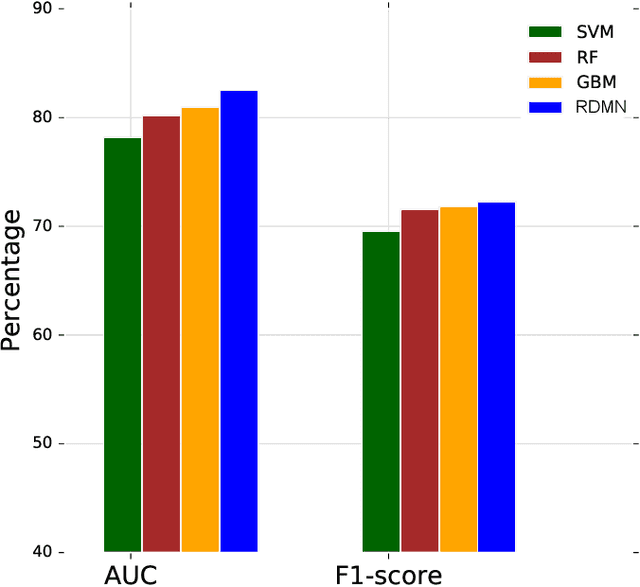
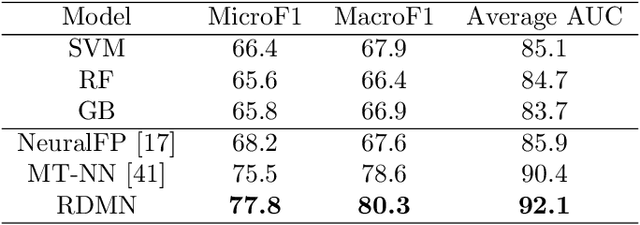
Abstract:Neural networks excel in detecting regular patterns but are less successful in representing and manipulating complex data structures, possibly due to the lack of an external memory. This has led to the recent development of a new line of architectures known as Memory-Augmented Neural Networks (MANNs), each of which consists of a neural network that interacts with an external memory matrix. However, this RAM-like memory matrix is unstructured and thus does not naturally encode structured objects. Here we design a new MANN dubbed Relational Dynamic Memory Network (RMDN) to bridge the gap. Like existing MANNs, RMDN has a neural controller but its memory is structured as multi-relational graphs. RMDN uses the memory to represent and manipulate graph-structured data in response to query; and as a neural network, RMDN is trainable from labeled data. Thus RMDN learns to answer queries about a set of graph-structured objects without explicit programming. We evaluate the capability of RMDN on several important prediction problems, including software vulnerability, molecular bioactivity and chemical-chemical interaction. Results demonstrate the efficacy of the proposed model.
Variational Memory Encoder-Decoder
Oct 20, 2018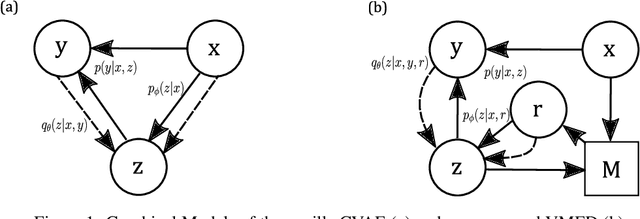
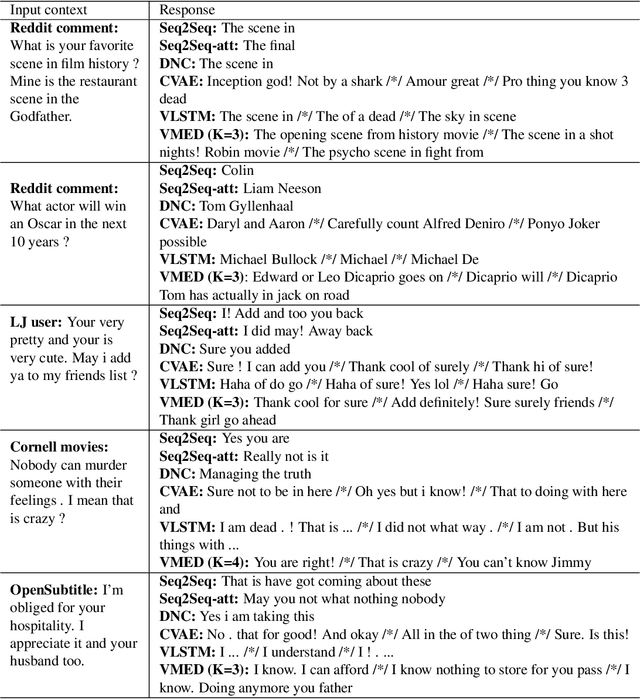
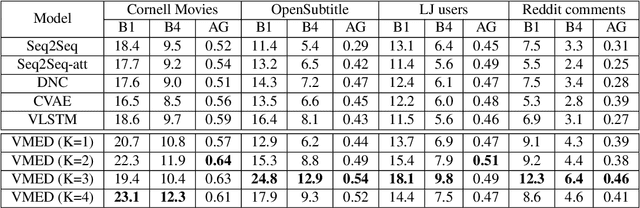
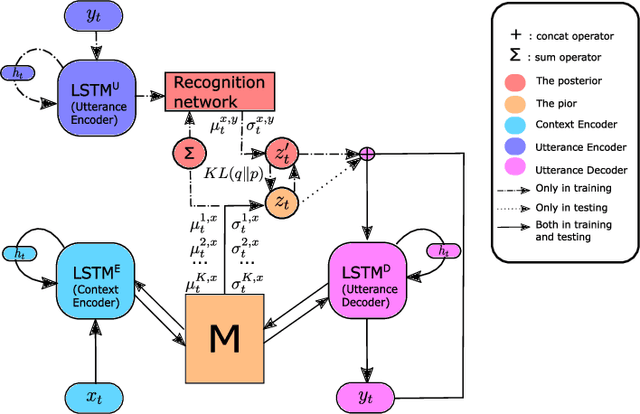

Abstract:Introducing variability while maintaining coherence is a core task in learning to generate utterances in conversation. Standard neural encoder-decoder models and their extensions using conditional variational autoencoder often result in either trivial or digressive responses. To overcome this, we explore a novel approach that injects variability into neural encoder-decoder via the use of external memory as a mixture model, namely Variational Memory Encoder-Decoder (VMED). By associating each memory read with a mode in the latent mixture distribution at each timestep, our model can capture the variability observed in sequential data such as natural conversations. We empirically compare the proposed model against other recent approaches on various conversational datasets. The results show that VMED consistently achieves significant improvement over others in both metric-based and qualitative evaluations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge