Suchi Saria
Computer Science, Statistics, and Health Policy, Johns Hopkins University, Baltimore, MD, USA, ML, AI and Healthcare Lab, Bayesian Health, New York, NY, USA
Beyond Low Earth Orbit: Biomonitoring, Artificial Intelligence, and Precision Space Health
Dec 22, 2021
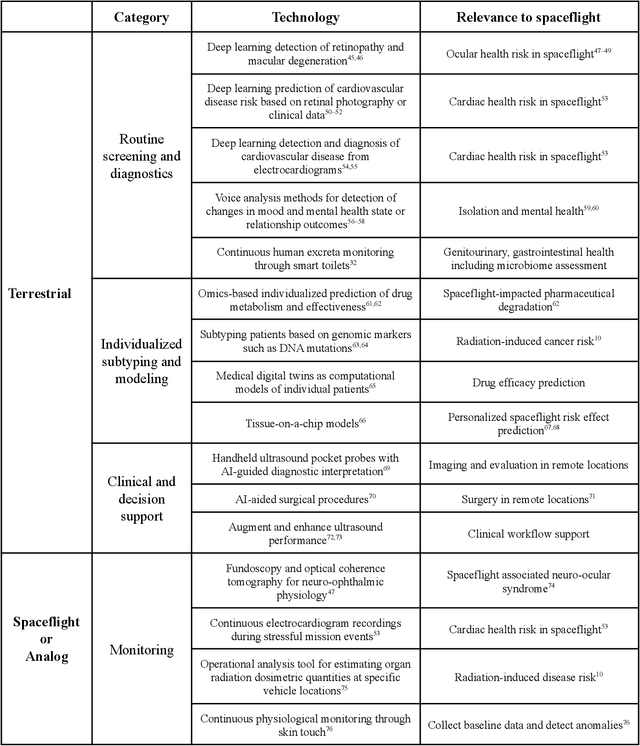
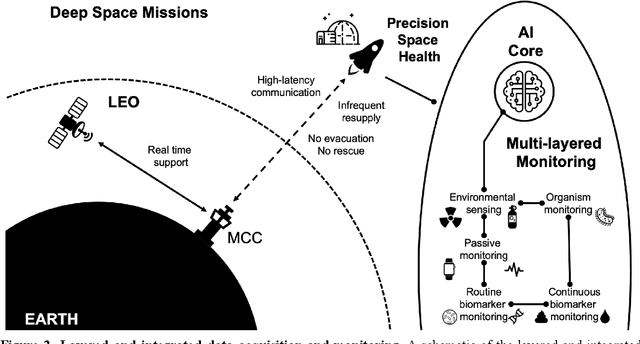
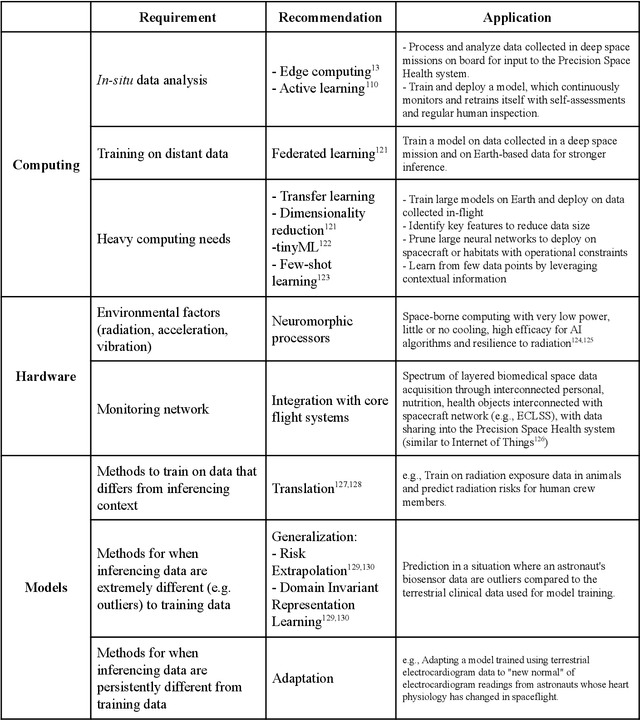
Abstract:Human space exploration beyond low Earth orbit will involve missions of significant distance and duration. To effectively mitigate myriad space health hazards, paradigm shifts in data and space health systems are necessary to enable Earth-independence, rather than Earth-reliance. Promising developments in the fields of artificial intelligence and machine learning for biology and health can address these needs. We propose an appropriately autonomous and intelligent Precision Space Health system that will monitor, aggregate, and assess biomedical statuses; analyze and predict personalized adverse health outcomes; adapt and respond to newly accumulated data; and provide preventive, actionable, and timely insights to individual deep space crew members and iterative decision support to their crew medical officer. Here we present a summary of recommendations from a workshop organized by the National Aeronautics and Space Administration, on future applications of artificial intelligence in space biology and health. In the next decade, biomonitoring technology, biomarker science, spacecraft hardware, intelligent software, and streamlined data management must mature and be woven together into a Precision Space Health system to enable humanity to thrive in deep space.
Beyond Low Earth Orbit: Biological Research, Artificial Intelligence, and Self-Driving Labs
Dec 22, 2021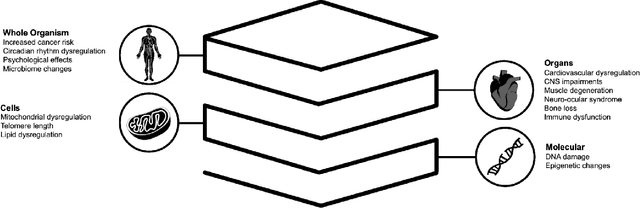
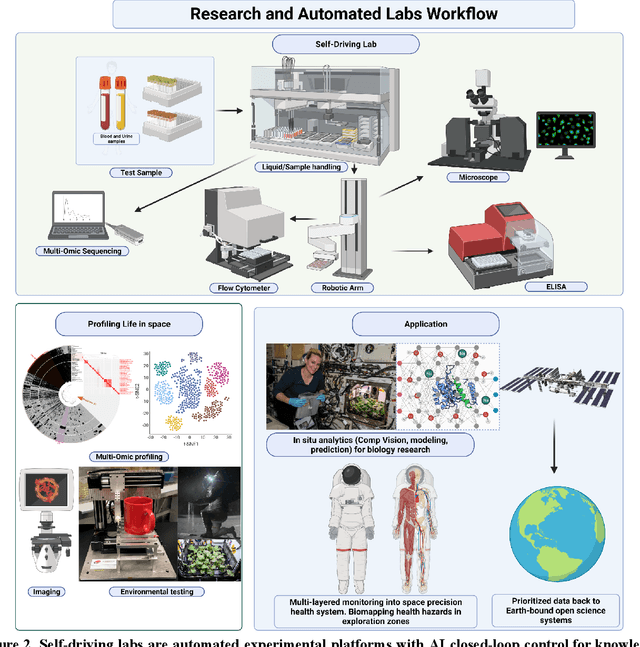
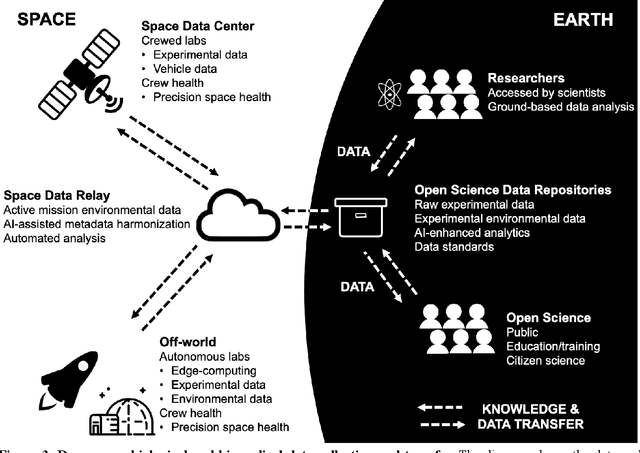
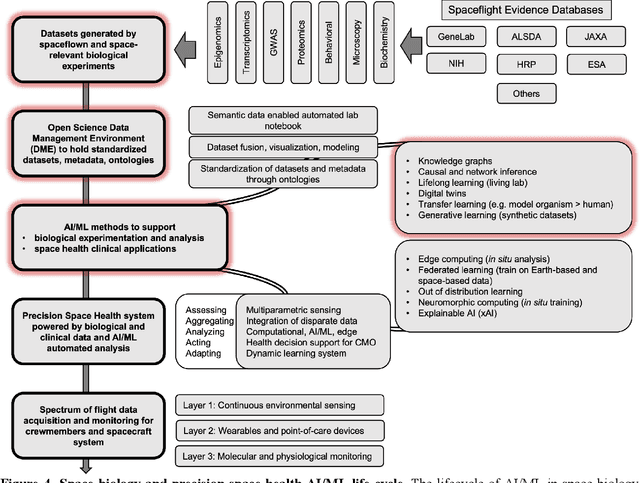
Abstract:Space biology research aims to understand fundamental effects of spaceflight on organisms, develop foundational knowledge to support deep space exploration, and ultimately bioengineer spacecraft and habitats to stabilize the ecosystem of plants, crops, microbes, animals, and humans for sustained multi-planetary life. To advance these aims, the field leverages experiments, platforms, data, and model organisms from both spaceborne and ground-analog studies. As research is extended beyond low Earth orbit, experiments and platforms must be maximally autonomous, light, agile, and intelligent to expedite knowledge discovery. Here we present a summary of recommendations from a workshop organized by the National Aeronautics and Space Administration on artificial intelligence, machine learning, and modeling applications which offer key solutions toward these space biology challenges. In the next decade, the synthesis of artificial intelligence into the field of space biology will deepen the biological understanding of spaceflight effects, facilitate predictive modeling and analytics, support maximally autonomous and reproducible experiments, and efficiently manage spaceborne data and metadata, all with the goal to enable life to thrive in deep space.
Partial Identifiability in Discrete Data With Measurement Error
Dec 23, 2020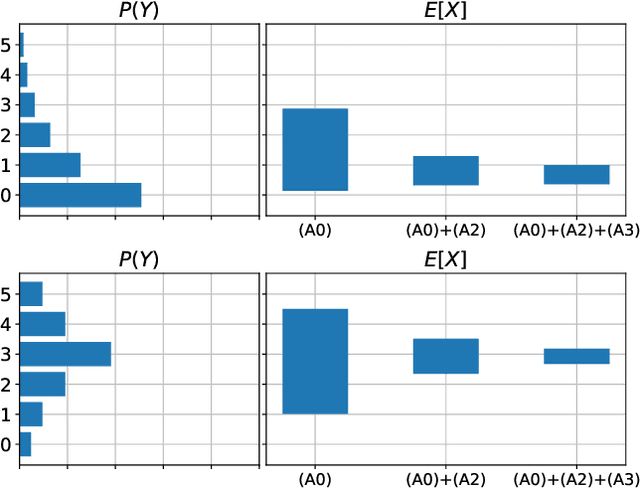
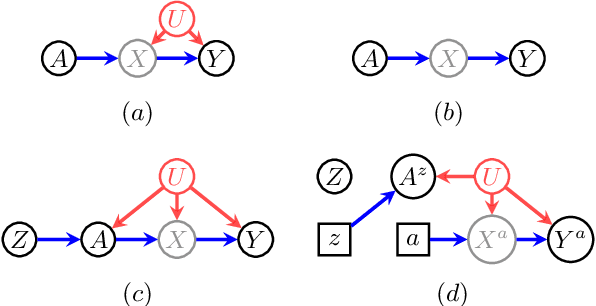
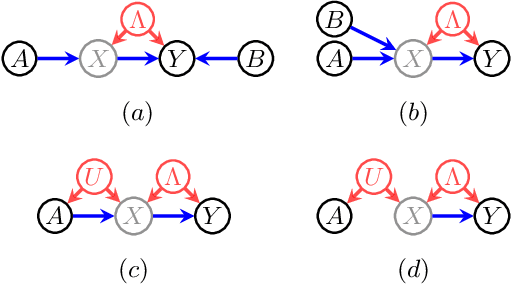
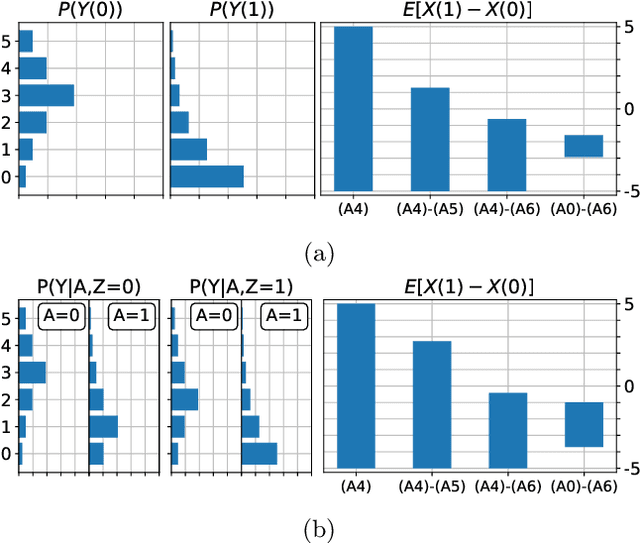
Abstract:When data contains measurement errors, it is necessary to make assumptions relating the observed, erroneous data to the unobserved true phenomena of interest. These assumptions should be justifiable on substantive grounds, but are often motivated by mathematical convenience, for the sake of exactly identifying the target of inference. We adopt the view that it is preferable to present bounds under justifiable assumptions than to pursue exact identification under dubious ones. To that end, we demonstrate how a broad class of modeling assumptions involving discrete variables, including common measurement error and conditional independence assumptions, can be expressed as linear constraints on the parameters of the model. We then use linear programming techniques to produce sharp bounds for factual and counterfactual distributions under measurement error in such models. We additionally propose a procedure for obtaining outer bounds on non-linear models. Our method yields sharp bounds in a number of important settings -- such as the instrumental variable scenario with measurement error -- for which no bounds were previously known.
Evaluating Model Robustness to Dataset Shift
Oct 28, 2020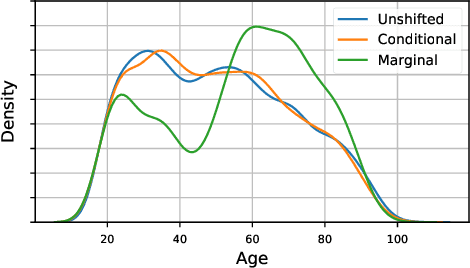
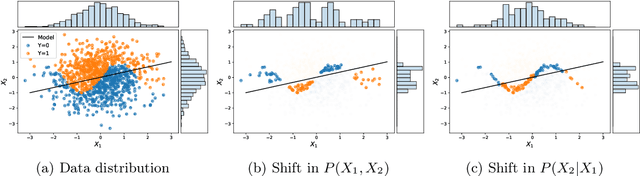
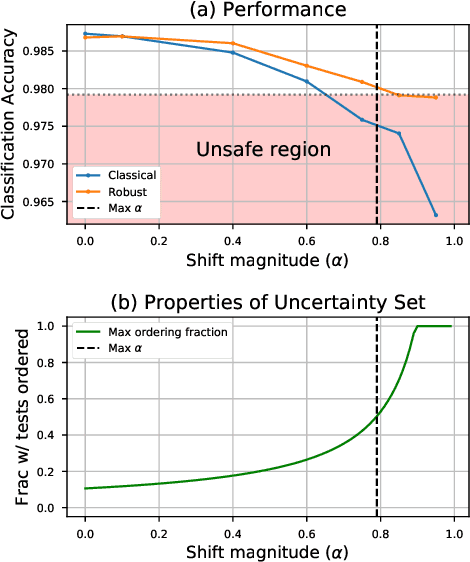
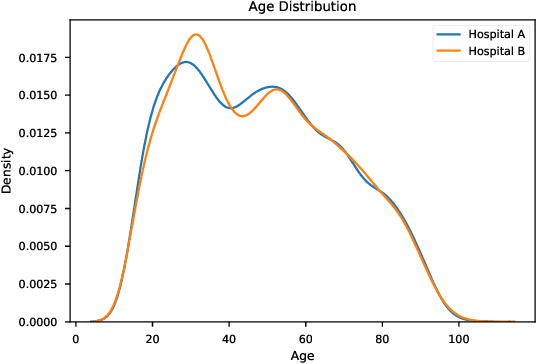
Abstract:As the use of machine learning in safety-critical domains becomes widespread, the importance of evaluating their safety has increased. An important aspect of this is evaluating how robust a model is to changes in setting or population, which typically requires applying the model to multiple, independent datasets. Since the cost of collecting such datasets is often prohibitive, in this paper, we propose a framework for evaluating this type of robustness using a single, fixed evaluation dataset. We use the original evaluation data to define an uncertainty set of possible evaluation distributions and estimate the algorithm's performance on the "worst-case" distribution within this set. Specifically, we consider distribution shifts defined by conditional distributions, allowing some distributions to shift while keeping other portions of the data distribution fixed. This results in finer-grained control over the considered shifts and more plausible worst-case distributions than previous approaches based on covariate shifts. To address the challenges associated with estimation in complex, high-dimensional distributions, we derive a "debiased" estimator which maintains $\sqrt{N}$-consistency even when machine learning methods with slower convergence rates are used to estimate the nuisance parameters. In experiments on a real medical risk prediction task, we show that this estimator can be used to evaluate robustness and accounts for realistic shifts that cannot be expressed as covariate shift. The proposed framework provides a means for practitioners to proactively evaluate the safety of their models using a single validation dataset.
I-SPEC: An End-to-End Framework for Learning Transportable, Shift-Stable Models
Feb 20, 2020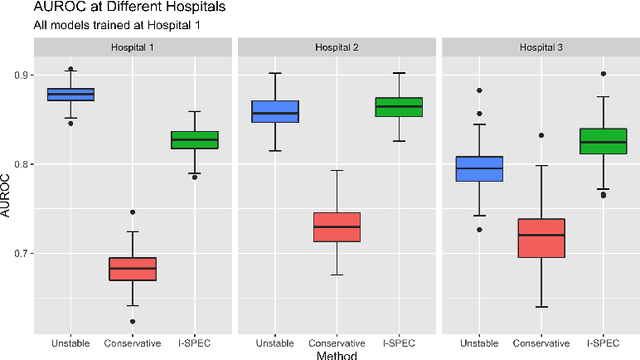
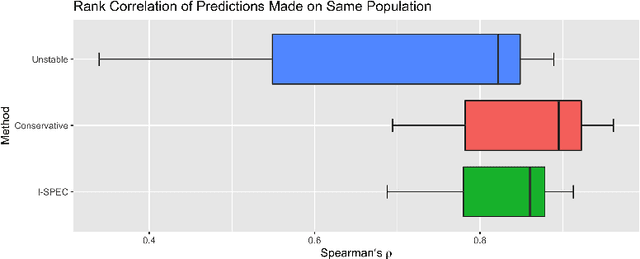
Abstract:Shifts in environment between development and deployment cause classical supervised learning to produce models that fail to generalize well to new target distributions. Recently, many solutions which find invariant predictive distributions have been developed. Among these, graph-based approaches do not require data from the target environment and can capture more stable information than alternative methods which find stable feature sets. However, these approaches assume that the data generating process is known in the form of a full causal graph, which is generally not the case. In this paper, we propose I-SPEC, an end-to-end framework that addresses this shortcoming by using data to learn a partial ancestral graph (PAG). Using the PAG we develop an algorithm that determines an interventional distribution that is stable to the declared shifts; this subsumes existing approaches which find stable feature sets that are less accurate. We apply I-SPEC to a mortality prediction problem to show it can learn a model that is robust to shifts without needing upfront knowledge of the full causal DAG.
The Hierarchy of Stable Distributions and Operators to Trade Off Stability and Performance
May 28, 2019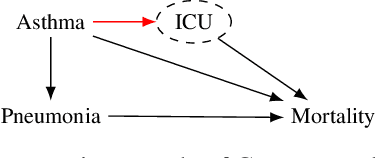
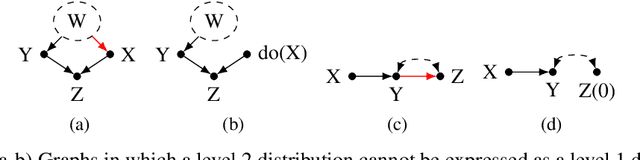
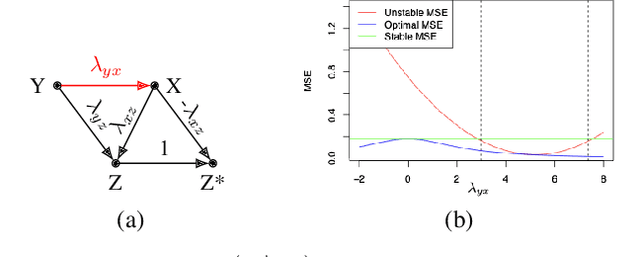
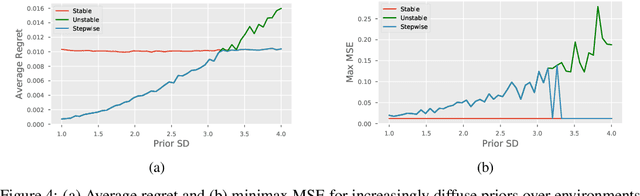
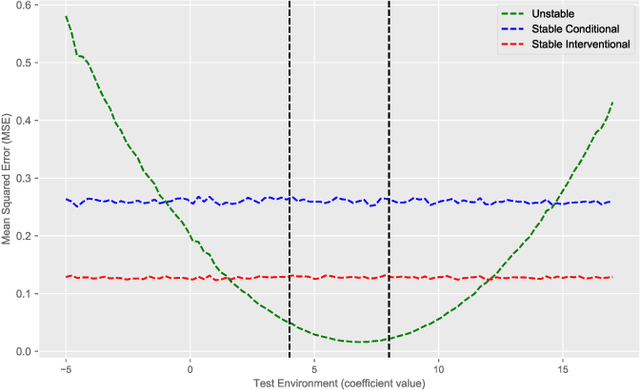
Abstract:Recent work addressing model reliability and generalization has resulted in a variety of methods that seek to proactively address differences between the training and unknown target environments. While most methods achieve this by finding distributions that will be invariant across environments, we will show they do not necessarily find the same distributions which has implications for performance. In this paper we unify existing work on prediction using stable distributions by relating environmental shifts to edges in the graph underlying a prediction problem, and characterize stable distributions as those which effectively remove these edges. We then quantify the effect of edge deletion on performance in the linear case and corroborate the findings in a simulated and real data experiment.
Tutorial: Safe and Reliable Machine Learning
Apr 15, 2019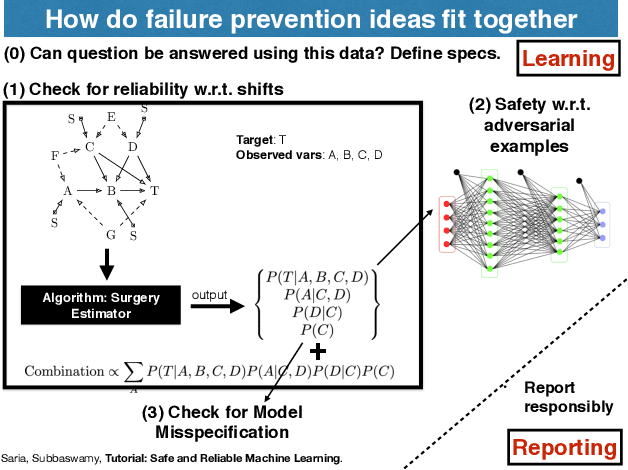
Abstract:This document serves as a brief overview of the "Safe and Reliable Machine Learning" tutorial given at the 2019 ACM Conference on Fairness, Accountability, and Transparency (FAT* 2019). The talk slides can be found here: https://bit.ly/2Gfsukp, while a video of the talk is available here: https://youtu.be/FGLOCkC4KmE, and a complete list of references for the tutorial here: https://bit.ly/2GdLPme.
Active Learning for Decision-Making from Imbalanced Observational Data
Apr 10, 2019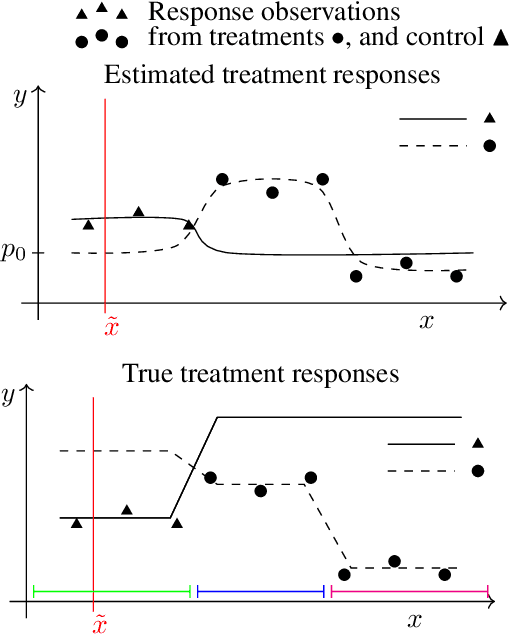
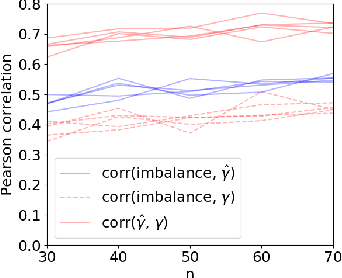
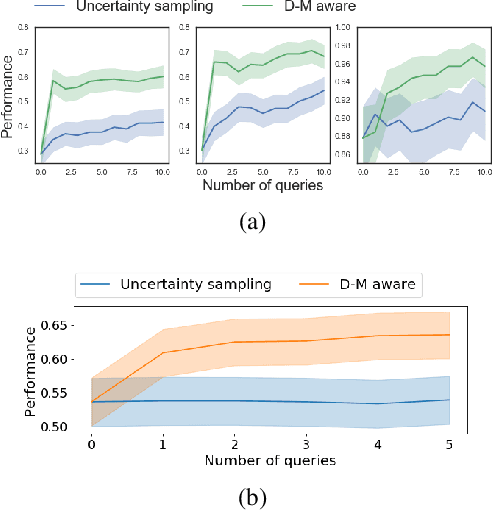
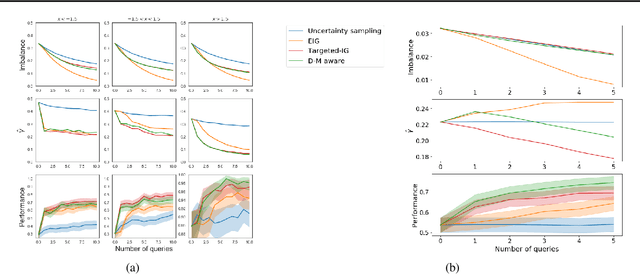
Abstract:Machine learning can help personalized decision support by learning models to predict individual treatment effects (ITE). This work studies the reliability of prediction-based decision-making in a task of deciding which action $a$ to take for a target unit after observing its covariates $\tilde{x}$ and predicted outcomes $\hat{p}(\tilde{y} \mid \tilde{x}, a)$. An example case is personalized medicine and the decision of which treatment to give to a patient. A common problem when learning these models from observational data is imbalance, that is, difference in treated/control covariate distributions, which is known to increase the upper bound of the expected ITE estimation error. We propose to assess the decision-making reliability by estimating the ITE model's Type S error rate, which is the probability of the model inferring the sign of the treatment effect wrong. Furthermore, we use the estimated reliability as a criterion for active learning, in order to collect new (possibly expensive) observations, instead of making a forced choice based on unreliable predictions. We demonstrate the effectiveness of this decision-making aware active learning in two decision-making tasks: in simulated data with binary outcomes and in a medical dataset with synthetic and continuous treatment outcomes.
Learning Models from Data with Measurement Error: Tackling Underreporting
Jan 25, 2019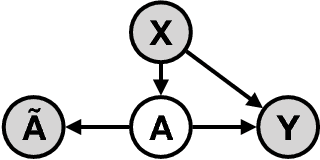
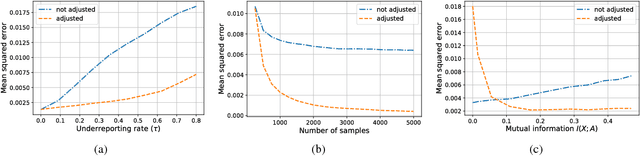
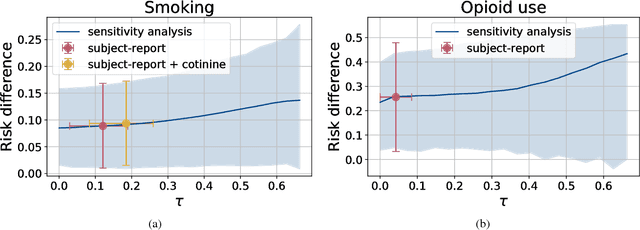
Abstract:Measurement error in observational datasets can lead to systematic bias in inferences based on these datasets. As studies based on observational data are increasingly used to inform decisions with real-world impact, it is critical that we develop a robust set of techniques for analyzing and adjusting for these biases. In this paper we present a method for estimating the distribution of an outcome given a binary exposure that is subject to underreporting. Our method is based on a missing data view of the measurement error problem, where the true exposure is treated as a latent variable that is marginalized out of a joint model. We prove three different conditions under which the outcome distribution can still be identified from data containing only error-prone observations of the exposure. We demonstrate this method on synthetic data and analyze its sensitivity to near violations of the identifiability conditions. Finally, we use this method to estimate the effects of maternal smoking and opioid use during pregnancy on childhood obesity, two import problems from public health. Using the proposed method, we estimate these effects using only subject-reported drug use data and substantially refine the range of estimates generated by a sensitivity analysis-based approach. Further, the estimates produced by our method are consistent with existing literature on both the effects of maternal smoking and the rate at which subjects underreport smoking.
Auditing Pointwise Reliability Subsequent to Training
Jan 02, 2019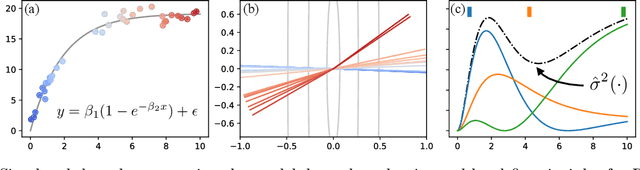
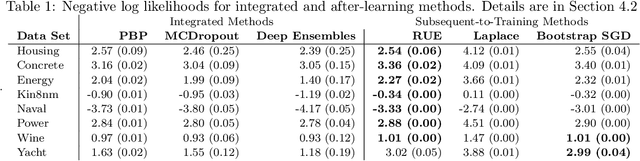
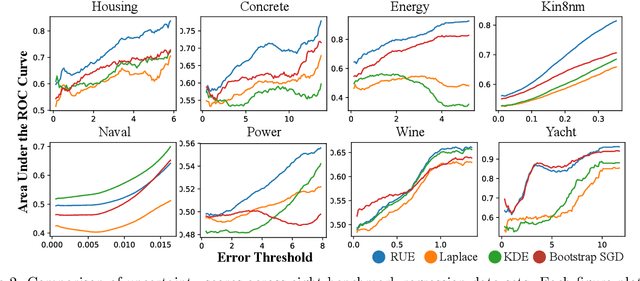
Abstract:To use machine learning in high stakes applications (e.g. medicine), we need tools for building confidence in the system and evaluating whether it is reliable. Methods to improve model reliability are often applied at train time (e.g. using Bayesian inference to obtain uncertainty estimates). An alternative is to audit a fixed model subsequent to training. In this paper, we describe resampling uncertainty estimation (RUE), an algorithm to audit the pointwise reliability of predictions. Intuitively, RUE estimates the amount that a single prediction would change if the model had been fit on different training data drawn from the same distribution by using the gradient and Hessian of the model's loss on training data. Experimentally, we show that RUE more effectively detects inaccurate predictions than existing tools for auditing reliability subsequent to training. We also show that RUE can create predictive distributions that are competitive with state-of-the-art methods like Monte Carlo dropout, probabilistic backpropagation, and deep ensembles, but does not depend on specific algorithms at train-time like these methods do.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge