Steven Johnson
Governance-Aware Vector Subscriptions for Multi-Agent Knowledge Ecosystems
Mar 21, 2026Abstract:As AI agent ecosystems grow, agents need mechanisms to monitor relevant knowledge in real time. Semantic publish-subscribe systems address this by matching new content against vector subscriptions. However, in multi-agent settings where agents operate under different data handling policies, unrestricted semantic subscriptions create policy violations: agents receive notifications about content they are not authorized to access. We introduce governance-aware vector subscriptions, a mechanism that composes semantic similarity matching with multi-dimensional policy predicates grounded in regulatory frameworks (EU DSM Directive, EU AI Act). The policy predicate operates over multiple independent dimensions (processing level, direct marketing restrictions, training opt-out, jurisdiction, and scientific usage) each with distinct legal bases. Agents subscribe to semantic regions of a curated knowledge base; notifications are dispatched only for validated content that passes both the similarity threshold and all applicable policy constraints. We formalize the mechanism, implement it within AIngram (an operational multi-agent knowledge base), and evaluate it using the PASA benchmark. We validate the mechanism on a synthetic corpus (1,000 chunks, 93 subscriptions, 5 domains): the governed mode correctly enforces all policy constraints while preserving delivery of authorized content. Ablation across five policy dimensions shows that no single dimension suffices for full compliance.
CancerLLM: A Large Language Model in Cancer Domain
Jun 15, 2024



Abstract:Medical Large Language Models (LLMs) such as ClinicalCamel 70B, Llama3-OpenBioLLM 70B have demonstrated impressive performance on a wide variety of medical NLP task.However, there still lacks a large language model (LLM) specifically designed for cancer domain. Moreover, these LLMs typically have billions of parameters, making them computationally expensive for healthcare systems.Thus, in this study, we propose CancerLLM, a model with 7 billion parameters and a Mistral-style architecture, pre-trained on 2,676,642 clinical notes and 515,524 pathology reports covering 17 cancer types, followed by fine-tuning on three cancer-relevant tasks, including cancer phenotypes extraction, cancer diagnosis generation, and cancer treatment plan generation. Our evaluation demonstrated that CancerLLM achieves state-of-the-art results compared to other existing LLMs, with an average F1 score improvement of 8.1\%. Additionally, CancerLLM outperforms other models on two proposed robustness testbeds. This illustrates that CancerLLM can be effectively applied to clinical AI systems, enhancing clinical research and healthcare delivery in the field of cancer.
Automated Assignment of Backbone NMR Data using Artificial Intelligence
Jun 18, 2015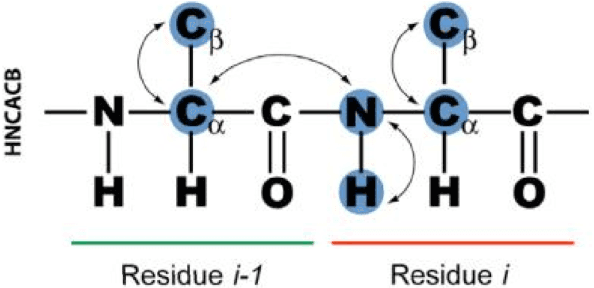
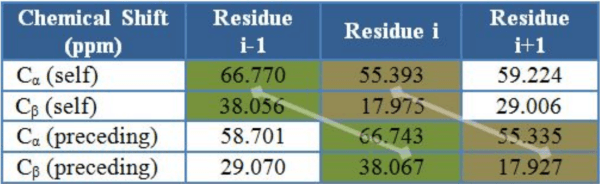
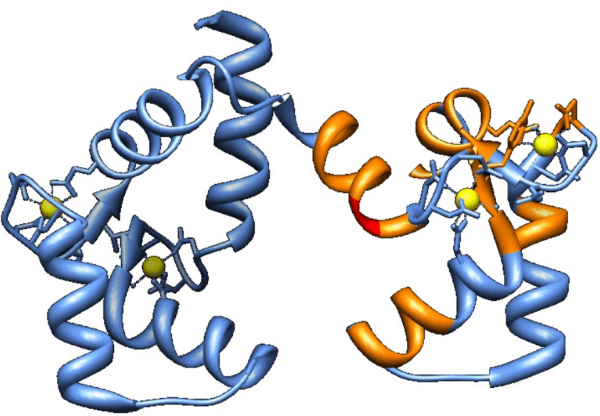
Abstract:Nuclear magnetic resonance (NMR) spectroscopy is a powerful method for the investigation of three-dimensional structures of biological molecules such as proteins. Determining a protein structure is essential for understanding its function and alterations in function which lead to disease. One of the major challenges of the post-genomic era is to obtain structural and functional information on the many unknown proteins encoded by thousands of newly identified genes. The goal of this research is to design an algorithm capable of automating the analysis of backbone protein NMR data by implementing AI strategies such as greedy and A* search.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge