Srinjoy Ganguly
Application of Quantum Pre-Processing Filter for Binary Image Classification with Small Samples
Aug 28, 2023



Abstract:Over the past few years, there has been significant interest in Quantum Machine Learning (QML) among researchers, as it has the potential to transform the field of machine learning. Several models that exploit the properties of quantum mechanics have been developed for practical applications. In this study, we investigated the application of our previously proposed quantum pre-processing filter (QPF) to binary image classification. We evaluated the QPF on four datasets: MNIST (handwritten digits), EMNIST (handwritten digits and alphabets), CIFAR-10 (photographic images) and GTSRB (real-life traffic sign images). Similar to our previous multi-class classification results, the application of QPF improved the binary image classification accuracy using neural network against MNIST, EMNIST, and CIFAR-10 from 98.9% to 99.2%, 97.8% to 98.3%, and 71.2% to 76.1%, respectively, but degraded it against GTSRB from 93.5% to 92.0%. We then applied QPF in cases using a smaller number of training and testing samples, i.e. 80 and 20 samples per class, respectively. In order to derive statistically stable results, we conducted the experiment with 100 trials choosing randomly different training and testing samples and averaging the results. The result showed that the application of QPF did not improve the image classification accuracy against MNIST and EMNIST but improved it against CIFAR-10 and GTSRB from 65.8% to 67.2% and 90.5% to 91.8%, respectively. Further research will be conducted as part of future work to investigate the potential of QPF to assess the scalability of the proposed approach to larger and complex datasets.
Development of a Novel Quantum Pre-processing Filter to Improve Image Classification Accuracy of Neural Network Models
Aug 22, 2023



Abstract:This paper proposes a novel quantum pre-processing filter (QPF) to improve the image classification accuracy of neural network (NN) models. A simple four qubit quantum circuit that uses Y rotation gates for encoding and two controlled NOT gates for creating correlation among the qubits is applied as a feature extraction filter prior to passing data into the fully connected NN architecture. By applying the QPF approach, the results show that the image classification accuracy based on the MNIST (handwritten 10 digits) and the EMNIST (handwritten 47 class digits and letters) datasets can be improved, from 92.5% to 95.4% and from 68.9% to 75.9%, respectively. These improvements were obtained without introducing extra model parameters or optimizations in the machine learning process. However, tests performed on the developed QPF approach against a relatively complex GTSRB dataset with 43 distinct class real-life traffic sign images showed a degradation in the classification accuracy. Considering this result, further research into the understanding and the design of a more suitable quantum circuit approach for image classification neural networks could be explored utilizing the baseline method proposed in this paper.
Quantum Natural Language Processing based Sentiment Analysis using lambeq Toolkit
May 30, 2023Abstract:Sentiment classification is one the best use case of classical natural language processing (NLP) where we can witness its power in various daily life domains such as banking, business and marketing industry. We already know how classical AI and machine learning can change and improve technology. Quantum natural language processing (QNLP) is a young and gradually emerging technology which has the potential to provide quantum advantage for NLP tasks. In this paper we show the first application of QNLP for sentiment analysis and achieve perfect test set accuracy for three different kinds of simulations and a decent accuracy for experiments ran on a noisy quantum device. We utilize the lambeq QNLP toolkit and $t|ket>$ by Cambridge Quantum (Quantinuum) to bring out the results.
* 6 pages, 9 figures
Optimal partition of feature using Bayesian classifier
Apr 27, 2023



Abstract:The Naive Bayesian classifier is a popular classification method employing the Bayesian paradigm. The concept of having conditional dependence among input variables sounds good in theory but can lead to a majority vote style behaviour. Achieving conditional independence is often difficult, and they introduce decision biases in the estimates. In Naive Bayes, certain features are called independent features as they have no conditional correlation or dependency when predicting a classification. In this paper, we focus on the optimal partition of features by proposing a novel technique called the Comonotone-Independence Classifier (CIBer) which is able to overcome the challenges posed by the Naive Bayes method. For different datasets, we clearly demonstrate the efficacy of our technique, where we achieve lower error rates and higher or equivalent accuracy compared to models such as Random Forests and XGBoost.
Hybrid Quantum Generative Adversarial Networks for Molecular Simulation and Drug Discovery
Dec 15, 2022



Abstract:In molecular research, simulation \& design of molecules are key areas with significant implications for drug development, material science, and other fields. Current classical computational power falls inadequate to simulate any more than small molecules, let alone protein chains on hundreds of peptide. Therefore these experiment are done physically in wet-lab, but it takes a lot of time \& not possible to examine every molecule due to the size of the search area, tens of billions of dollars are spent every year in these research experiments. Molecule simulation \& design has lately advanced significantly by machine learning models, A fresh perspective on the issue of chemical synthesis is provided by deep generative models for graph-structured data. By optimising differentiable models that produce molecular graphs directly, it is feasible to avoid costly search techniques in the discrete and huge space of chemical structures. But these models also suffer from computational limitations when dimensions become huge and consume huge amount of resources. Quantum Generative machine learning in recent years have shown some empirical results promising significant advantages over classical counterparts.
Variational Quantum Algorithms for Chemical Simulation and Drug Discovery
Nov 15, 2022



Abstract:Quantum computing has gained a lot of attention recently, and scientists have seen potential applications in this field using quantum computing for Cryptography and Communication to Machine Learning and Healthcare. Protein folding has been one of the most interesting areas to study, and it is also one of the biggest problems of biochemistry. Each protein folds distinctively, and the difficulty of finding its stable shape rapidly increases with an increase in the number of amino acids in the chain. A moderate protein has about 100 amino acids, and the number of combinations one needs to verify to find the stable structure is enormous. At some point, the number of these combinations will be so vast that classical computers cannot even attempt to solve them. In this paper, we examine how this problem can be solved with the help of quantum computing using two different algorithms, Variational Quantum Eigensolver (VQE) and Quantum Approximate Optimization Algorithm (QAOA), using Qiskit Nature. We compare the results of different quantum hardware and simulators and check how error mitigation affects the performance. Further, we make comparisons with SoTA algorithms and evaluate the reliability of the method.
Clustering using Vector Membership: An Extension of the Fuzzy C-Means Algorithm
Dec 14, 2013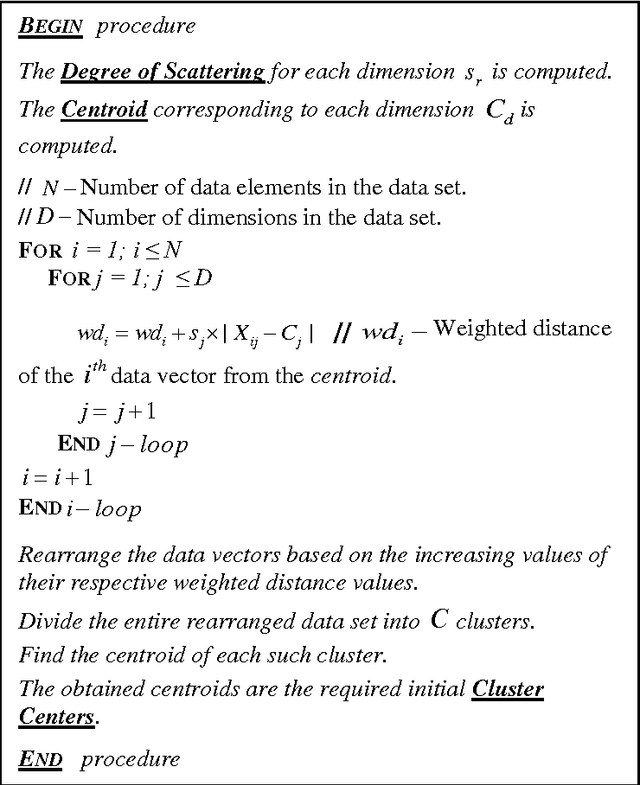
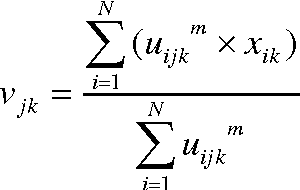
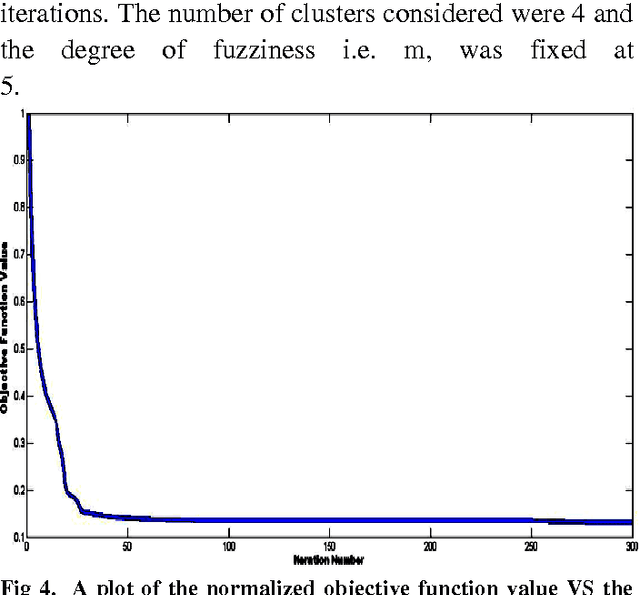
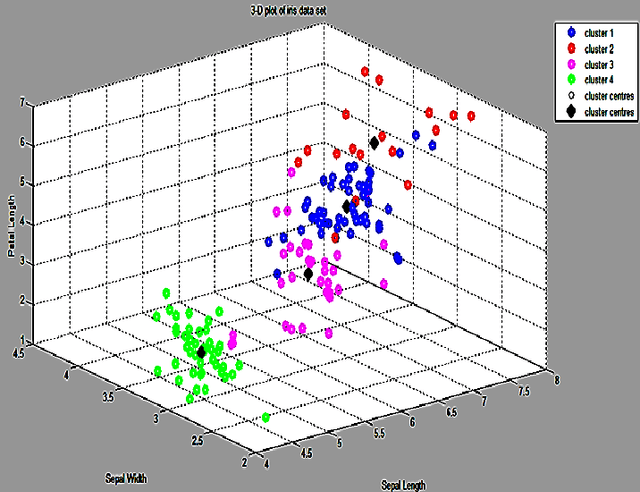
Abstract:Clustering is an important facet of explorative data mining and finds extensive use in several fields. In this paper, we propose an extension of the classical Fuzzy C-Means clustering algorithm. The proposed algorithm, abbreviated as VFC, adopts a multi-dimensional membership vector for each data point instead of the traditional, scalar membership value defined in the original algorithm. The membership vector for each point is obtained by considering each feature of that point separately and obtaining individual membership values for the same. We also propose an algorithm to efficiently allocate the initial cluster centers close to the actual centers, so as to facilitate rapid convergence. Further, we propose a scheme to achieve crisp clustering using the VFC algorithm. The proposed, novel clustering scheme has been tested on two standard data sets in order to analyze its performance. We also examine the efficacy of the proposed scheme by analyzing its performance on image segmentation examples and comparing it with the classical Fuzzy C-means clustering algorithm.
* 6 pages, 8 figures and 1 table (Conference Paper)
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge