Shreya Roy
Indian Institute of Science, Bangalore
Single Image Super-resolution with a Switch Guided Hybrid Network for Satellite Images
Nov 29, 2020
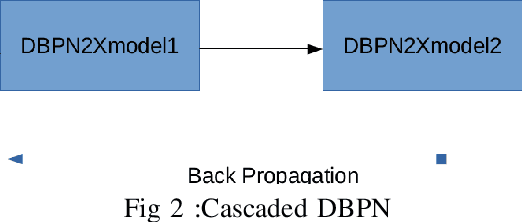
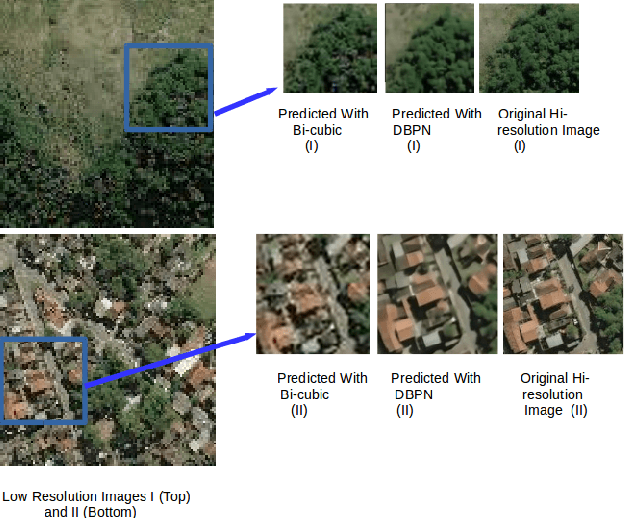
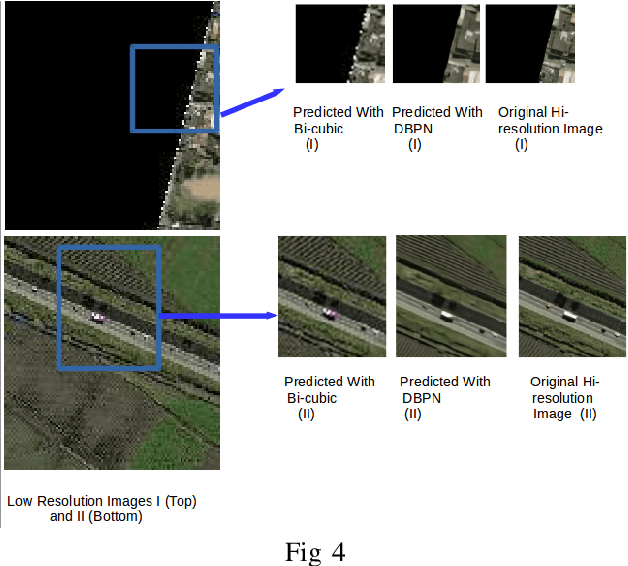
Abstract:The major drawbacks with Satellite Images are low resolution, Low resolution makes it difficult to identify the objects present in Satellite images. We have experimented with several deep models available for Single Image Superresolution on the SpaceNet dataset and have evaluated the performance of each of them on the satellite image data. We will dive into the recent evolution of the deep models in the context of SISR over the past few years and will present a comparative study between these models. The entire Satellite image of an area is divided into equal-sized patches. Each patch will be used independently for training. These patches will differ in nature. Say, for example, the patches over urban areas have non-homogeneous backgrounds because of different types of objects like vehicles, buildings, roads, etc. On the other hand, patches over jungles will be more homogeneous in nature. Hence, different deep models will fit on different kinds of patches. In this study, we will try to explore this further with the help of a Switching Convolution Network. The idea is to train a switch classifier that will automatically classify a patch into one category of models best suited for it.
Semantic Segmentation of highly class imbalanced fully labelled 3D volumetric biomedical images and unsupervised Domain Adaptation of the pre-trained Segmentation Network to segment another fully unlabelled Biomedical 3D Image stack
Mar 13, 2020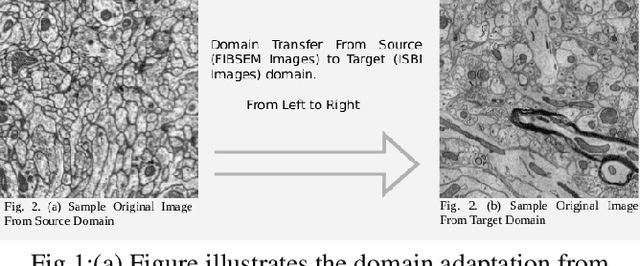
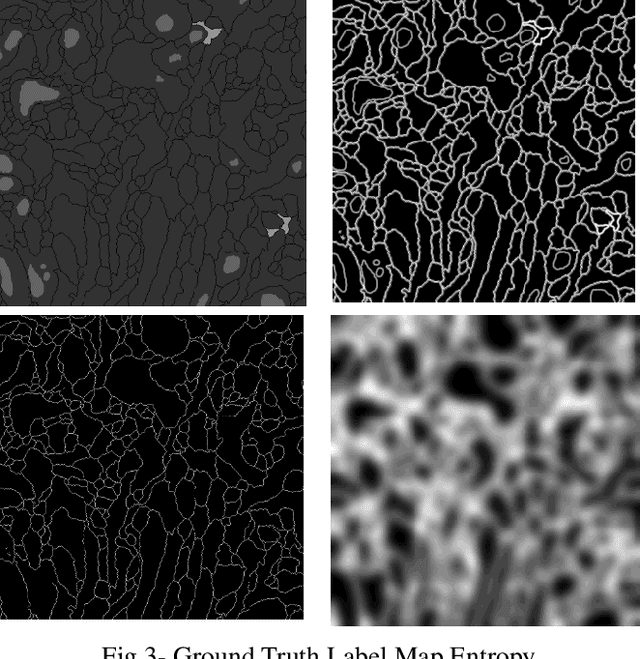
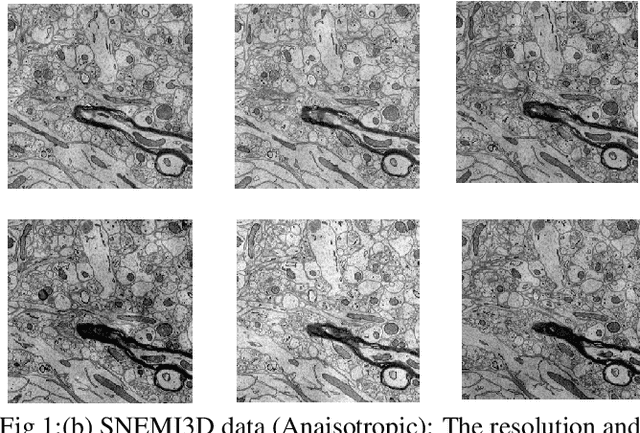
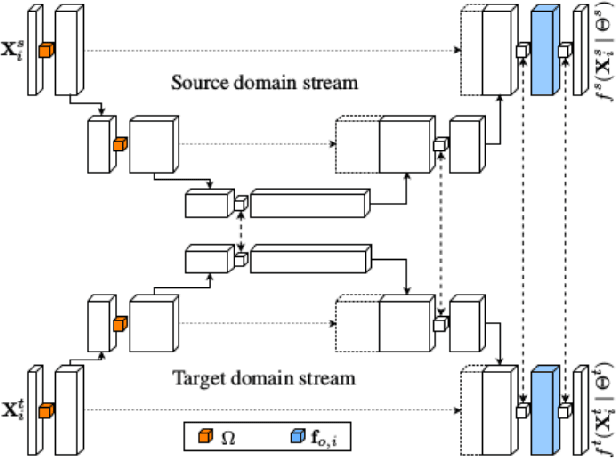

Abstract:The goal of our work is to perform pixel label semantic segmentation on 3D biomedical volumetric data. Manual annotation is always difficult for a large bio-medical dataset. So, we consider two cases where one dataset is fully labeled and the other dataset is assumed to be fully unlabelled. We first perform Semantic Segmentation on the fully labeled isotropic biomedical source data (FIBSEM) and try to incorporate the the trained model for segmenting the target unlabelled dataset(SNEMI3D)which shares some similarities with the source dataset in the context of different types of cellular bodies and other cellular components. Although, the cellular components vary in size and shape. So in this paper, we have proposed a novel approach in the context of unsupervised domain adaptation while classifying each pixel of the target volumetric data into cell boundary and cell body. Also, we have proposed a novel approach to giving non-uniform weights to different pixels in the training images while performing the pixel-level semantic segmentation in the presence of the corresponding pixel-wise label map along with the training original images in the source domain. We have used the Entropy Map or a Distance Transform matrix retrieved from the given ground truth label map which has helped to overcome the class imbalance problem in the medical image data where the cell boundaries are extremely thin and hence, extremely prone to be misclassified as non-boundary.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge