Sakib Mostafa
Toward a universal foundation model for graph-structured data
Apr 07, 2026Abstract:Graphs are a central representation in biomedical research, capturing molecular interaction networks, gene regulatory circuits, cell--cell communication maps, and knowledge graphs. Despite their importance, currently there is not a broadly reusable foundation model available for graph analysis comparable to those that have transformed language and vision. Existing graph neural networks are typically trained on a single dataset and learn representations specific only to that graph's node features, topology, and label space, limiting their ability to transfer across domains. This lack of generalization is particularly problematic in biology and medicine, where networks vary substantially across cohorts, assays, and institutions. Here we introduce a graph foundation model designed to learn transferable structural representations that are not specific to specific node identities or feature schemes. Our approach leverages feature-agnostic graph properties, including degree statistics, centrality measures, community structure indicators, and diffusion-based signatures, and encodes them as structural prompts. These prompts are integrated with a message-passing backbone to embed diverse graphs into a shared representation space. The model is pretrained once on heterogeneous graphs and subsequently reused on unseen datasets with minimal adaptation. Across multiple benchmarks, our pretrained model matches or exceeds strong supervised baselines while demonstrating superior zero-shot and few-shot generalization on held-out graphs. On the SagePPI benchmark, supervised fine-tuning of the pretrained backbone achieves a mean ROC-AUC of 95.5%, a gain of 21.8% over the best supervised message-passing baseline. The proposed technique thus provides a unique approach toward reusable, foundation-scale models for graph-structured data in biomedical and network science applications.
Vision-based Deep Learning Analysis of Unordered Biomedical Tabular Datasets via Optimal Spatial Cartography
Mar 24, 2026Abstract:Tabular data are central to biomedical research, from liquid biopsy and bulk and single-cell transcriptomics to electronic health records and phenotypic profiling. Unlike images or sequences, however, tabular datasets lack intrinsic spatial organization: features are treated as unordered dimensions, and their relationships must be inferred implicitly by the model. This limits the ability of vision architectures to exploit local structure and higher-order feature interactions in non-spatial biomedical data. Here we introduce Dynamic Feature Mapping (Dynomap), an end-to-end deep learning framework that learns a task-optimized spatial topology of features directly from data. Dynomap jointly optimizes feature placement and prediction through a fully differentiable rendering mechanism, without relying on heuristics, predefined groupings, or external priors. By transforming high-dimensional tabular vectors into learned feature maps, Dynomap enables vision-based models to operate effectively on unordered biomedical inputs. Across multiple clinical and biological datasets, Dynomap consistently outperformed classical machine learning, modern deep tabular models, and existing vector-to-image approaches. In liquid biopsy data, Dynomap organized clinically relevant gene signatures into coherent spatial patterns and improved multiclass cancer subtype prediction accuracy by up to 18%. In a Parkinson disease voice dataset, it clustered disease-associated acoustic descriptors and improved accuracy by up to 8%. Similar gains and interpretable feature organization were observed in additional biomedical datasets. These results establish Dynomap as a general strategy for bridging tabular and vision-based deep learning and for uncovering structured, clinically relevant patterns in high-dimensional biomedical data.
On the Evolution of Neuron Communities in a Deep Learning Architecture
Jun 08, 2021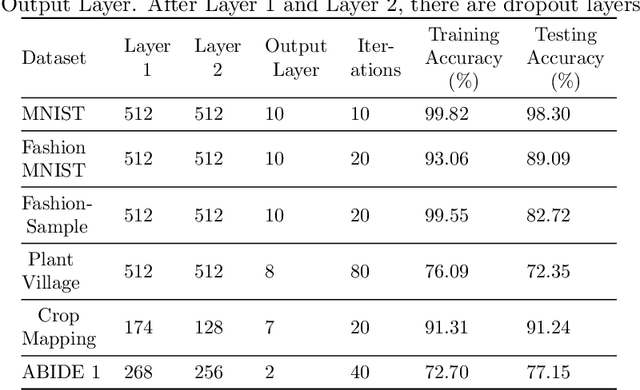
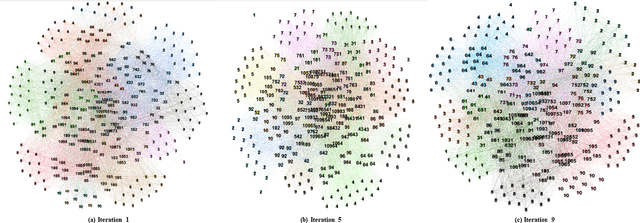
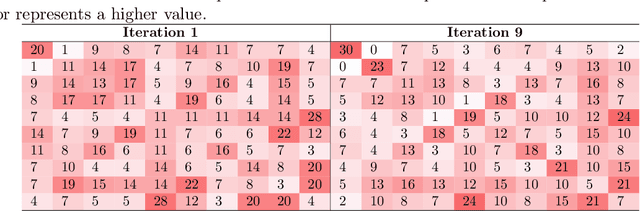
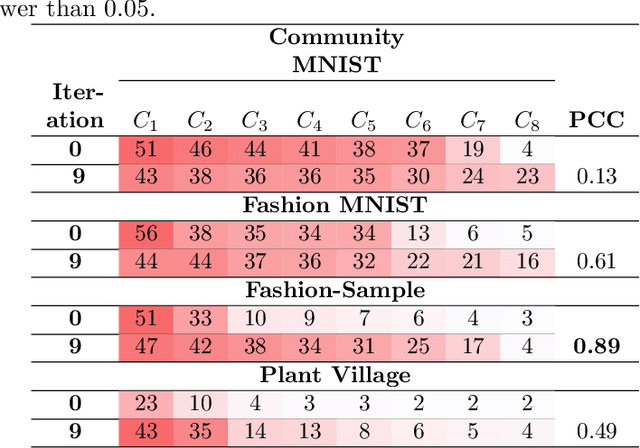
Abstract:Deep learning techniques are increasingly being adopted for classification tasks over the past decade, yet explaining how deep learning architectures can achieve state-of-the-art performance is still an elusive goal. While all the training information is embedded deeply in a trained model, we still do not understand much about its performance by only analyzing the model. This paper examines the neuron activation patterns of deep learning-based classification models and explores whether the models' performances can be explained through neurons' activation behavior. We propose two approaches: one that models neurons' activation behavior as a graph and examines whether the neurons form meaningful communities, and the other examines the predictability of neurons' behavior using entropy. Our comprehensive experimental study reveals that both the community quality (modularity) and entropy are closely related to the deep learning models' performances, thus paves a novel way of explaining deep learning models directly from the neurons' activation pattern.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge