Rustam Zhumagambetov
Generative clinical time series models trained on moderate amounts of patient data are privacy preserving
Feb 11, 2026Abstract:Sharing medical data for machine learning model training purposes is often impossible due to the risk of disclosing identifying information about individual patients. Synthetic data produced by generative artificial intelligence (genAI) models trained on real data is often seen as one possible solution to comply with privacy regulations. While powerful genAI models for heterogeneous hospital time series have recently been introduced, such modeling does not guarantee privacy protection, as the generated data may still reveal identifying information about individuals in the models' training cohort. Applying established privacy mechanisms to generative time series models, however, proves challenging as post-hoc data anonymization through k-anonymization or similar techniques is limited, while model-centered privacy mechanisms that implement differential privacy (DP) may lead to unstable training, compromising the utility of generated data. Given these known limitations, privacy audits for generative time series models are currently indispensable regardless of the concrete privacy mechanisms applied to models and/or data. In this work, we use a battery of established privacy attacks to audit state-of-the-art hospital time series models, trained on the public MIMIC-IV dataset, with respect to privacy preservation. Furthermore, the eICU dataset was used to mount a privacy attack against the synthetic data generator trained on the MIMIC-IV dataset. Results show that established privacy attacks are ineffective against generated multivariate clinical time series when synthetic data generators are trained on large enough training datasets. Furthermore, we discuss how the use of existing DP mechanisms for these synthetic data generators would not bring desired improvement in privacy, but only a decrease in utility for machine learning prediction tasks.
Explainable AI needs formal notions of explanation correctness
Sep 26, 2024Abstract:The use of machine learning (ML) in critical domains such as medicine poses risks and requires regulation. One requirement is that decisions of ML systems in high-risk applications should be human-understandable. The field of "explainable artificial intelligence" (XAI) seemingly addresses this need. However, in its current form, XAI is unfit to provide quality control for ML; it itself needs scrutiny. Popular XAI methods cannot reliably answer important questions about ML models, their training data, or a given test input. We recapitulate results demonstrating that popular XAI methods systematically attribute importance to input features that are independent of the prediction target. This limits their utility for purposes such as model and data (in)validation, model improvement, and scientific discovery. We argue that the fundamental reason for this limitation is that current XAI methods do not address well-defined problems and are not evaluated against objective criteria of explanation correctness. Researchers should formally define the problems they intend to solve first and then design methods accordingly. This will lead to notions of explanation correctness that can be theoretically verified and objective metrics of explanation performance that can be assessed using ground-truth data.
A biologically-inspired evaluation of molecular generative machine learning
Aug 20, 2022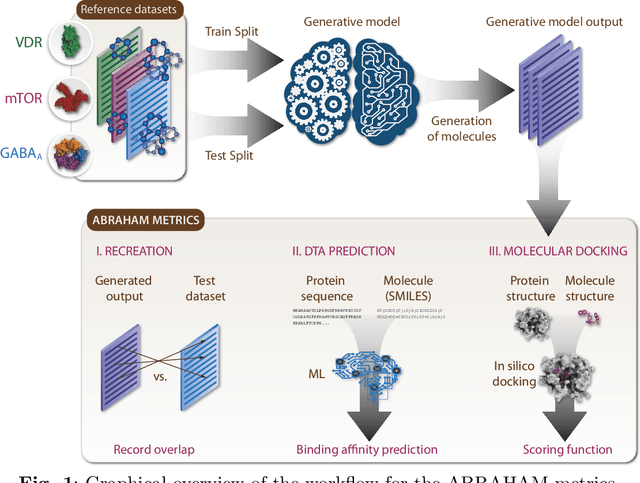
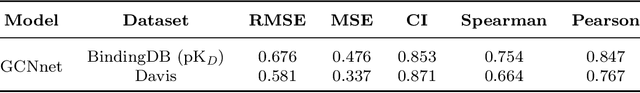
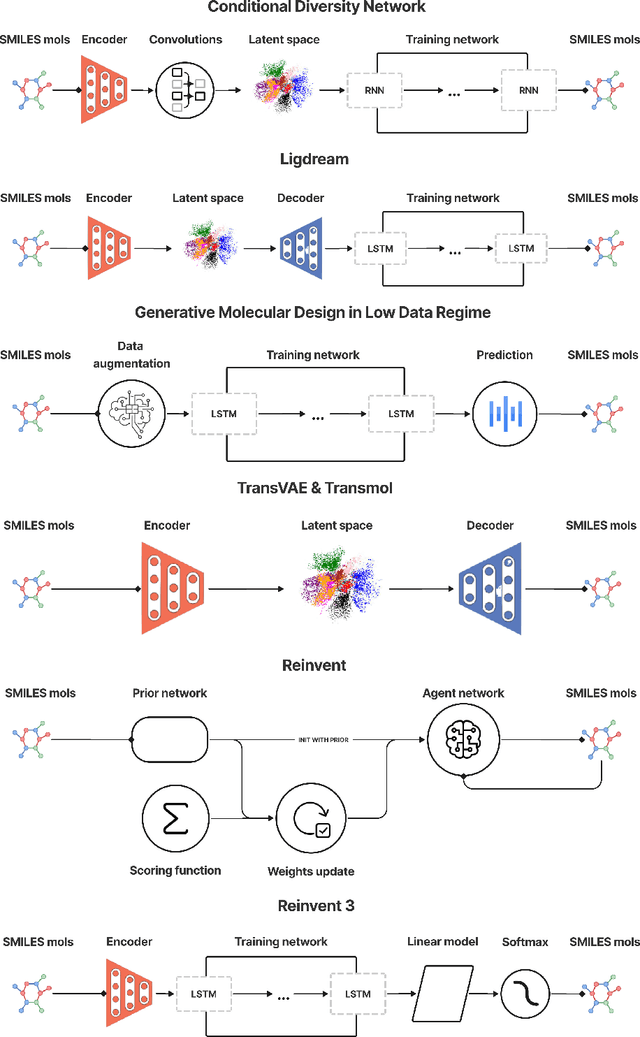
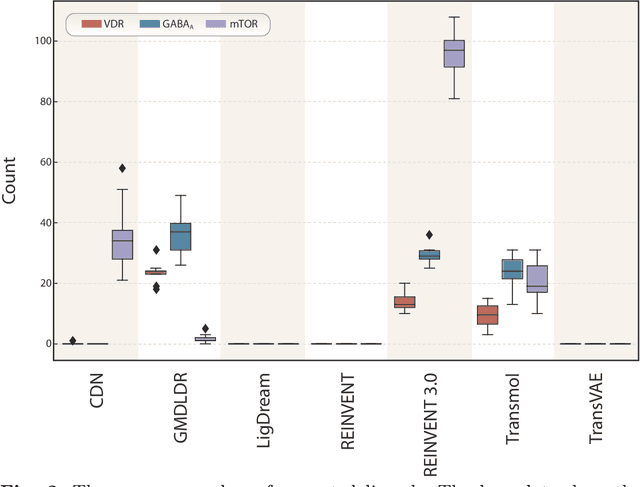
Abstract:While generative models have recently become ubiquitous in many scientific areas, less attention has been paid to their evaluation. For molecular generative models, the state-of-the-art examines their output in isolation or in relation to its input. However, their biological and functional properties, such as ligand-target interaction is not being addressed. In this study, a novel biologically-inspired benchmark for the evaluation of molecular generative models is proposed. Specifically, three diverse reference datasets are designed and a set of metrics are introduced which are directly relevant to the drug discovery process. In particular we propose a recreation metric, apply drug-target affinity prediction and molecular docking as complementary techniques for the evaluation of generative outputs. While all three metrics show consistent results across the tested generative models, a more detailed comparison of drug-target affinity binding and molecular docking scores revealed that unimodal predictiors can lead to erroneous conclusions about target binding on a molecular level and a multi-modal approach is thus preferrable. The key advantage of this framework is that it incorporates prior physico-chemical domain knowledge into the benchmarking process by focusing explicitly on ligand-target interactions and thus creating a highly efficient tool not only for evaluating molecular generative outputs in particular, but also for enriching the drug discovery process in general.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge