Roger Ghanem
Transient anisotropic kernel for probabilistic learning on manifolds
Jul 31, 2024Abstract:PLoM (Probabilistic Learning on Manifolds) is a method introduced in 2016 for handling small training datasets by projecting an It\^o equation from a stochastic dissipative Hamiltonian dynamical system, acting as the MCMC generator, for which the KDE-estimated probability measure with the training dataset is the invariant measure. PLoM performs a projection on a reduced-order vector basis related to the training dataset, using the diffusion maps (DMAPS) basis constructed with a time-independent isotropic kernel. In this paper, we propose a new ISDE projection vector basis built from a transient anisotropic kernel, providing an alternative to the DMAPS basis to improve statistical surrogates for stochastic manifolds with heterogeneous data. The construction ensures that for times near the initial time, the DMAPS basis coincides with the transient basis. For larger times, the differences between the two bases are characterized by the angle of their spanned vector subspaces. The optimal instant yielding the optimal transient basis is determined using an estimation of mutual information from Information Theory, which is normalized by the entropy estimation to account for the effects of the number of realizations used in the estimations. Consequently, this new vector basis better represents statistical dependencies in the learned probability measure for any dimension. Three applications with varying levels of statistical complexity and data heterogeneity validate the proposed theory, showing that the transient anisotropic kernel improves the learned probability measure.
Data-based Discovery of Governing Equations
Dec 21, 2020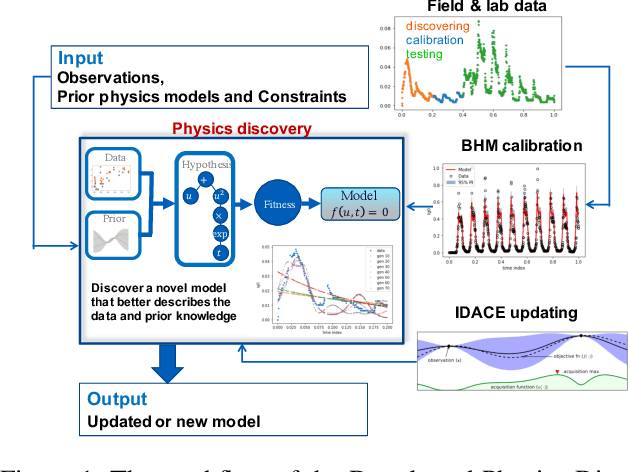
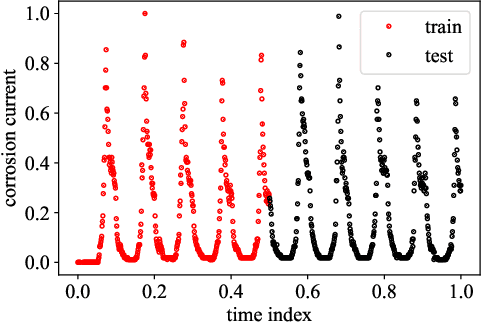
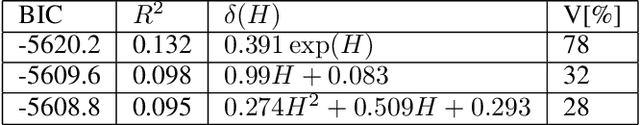
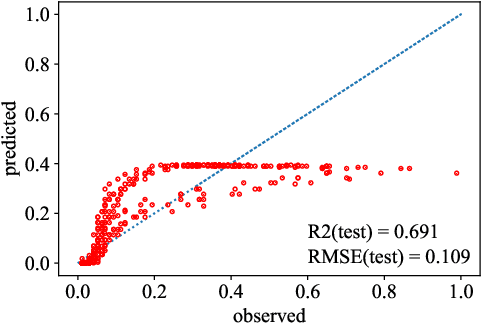
Abstract:Most common mechanistic models are traditionally presented in mathematical forms to explain a given physical phenomenon. Machine learning algorithms, on the other hand, provide a mechanism to map the input data to output without explicitly describing the underlying physical process that generated the data. We propose a Data-based Physics Discovery (DPD) framework for automatic discovery of governing equations from observed data. Without a prior definition of the model structure, first a free-form of the equation is discovered, and then calibrated and validated against the available data. In addition to the observed data, the DPD framework can utilize available prior physical models, and domain expert feedback. When prior models are available, the DPD framework can discover an additive or multiplicative correction term represented symbolically. The correction term can be a function of the existing input variable to the prior model, or a newly introduced variable. In case a prior model is not available, the DPD framework discovers a new data-based standalone model governing the observations. We demonstrate the performance of the proposed framework on a real-world application in the aerospace industry.
Probabilistic learning on manifolds constrained by nonlinear partial differential equations for small datasets
Oct 27, 2020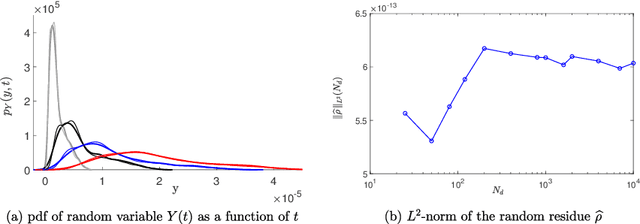
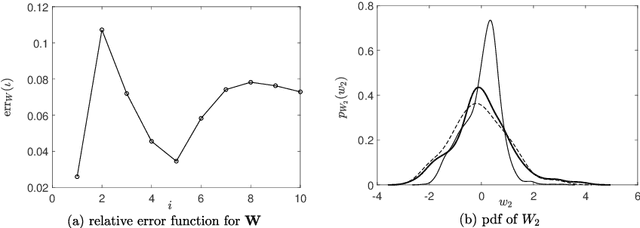
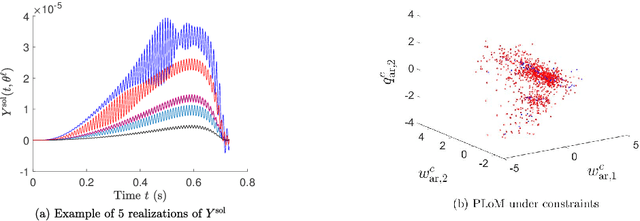
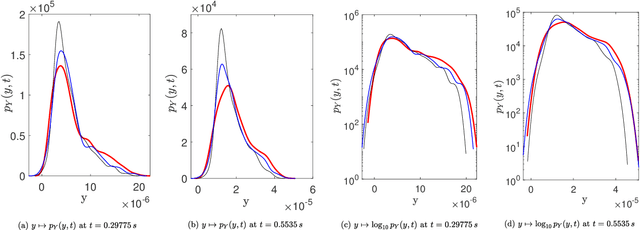
Abstract:A novel extension of the Probabilistic Learning on Manifolds (PLoM) is presented. It makes it possible to synthesize solutions to a wide range of nonlinear stochastic boundary value problems described by partial differential equations (PDEs) for which a stochastic computational model (SCM) is available and depends on a vector-valued random control parameter. The cost of a single numerical evaluation of this SCM is assumed to be such that only a limited number of points can be computed for constructing the training dataset (small data). Each point of the training dataset is made up realizations from a vector-valued stochastic process (the stochastic solution) and the associated random control parameter on which it depends. The presented PLoM constrained by PDE allows for generating a large number of learned realizations of the stochastic process and its corresponding random control parameter. These learned realizations are generated so as to minimize the vector-valued random residual of the PDE in the mean-square sense. Appropriate novel methods are developed to solve this challenging problem. Three applications are presented. The first one is a simple uncertain nonlinear dynamical system with a nonstationary stochastic excitation. The second one concerns the 2D nonlinear unsteady Navier-Stokes equations for incompressible flows in which the Reynolds number is the random control parameter. The last one deals with the nonlinear dynamics of a 3D elastic structure with uncertainties. The results obtained make it possible to validate the PLoM constrained by stochastic PDE but also provide further validation of the PLoM without constraint.
Normal-bundle Bootstrap
Jul 27, 2020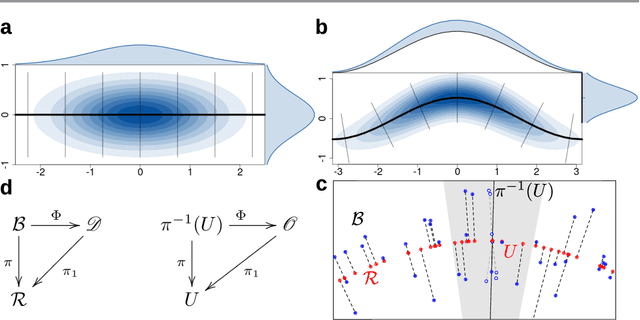
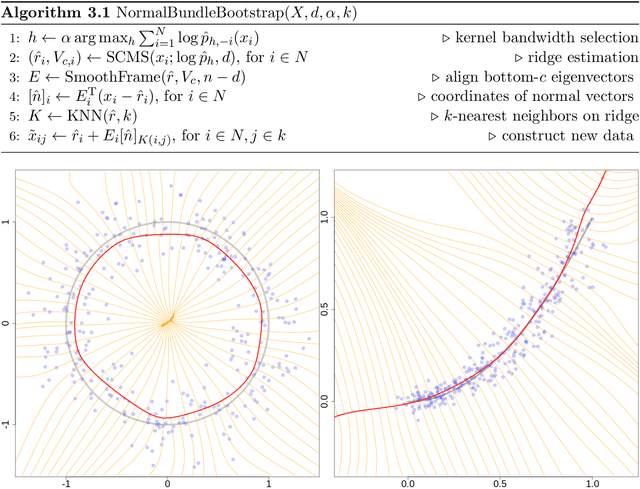
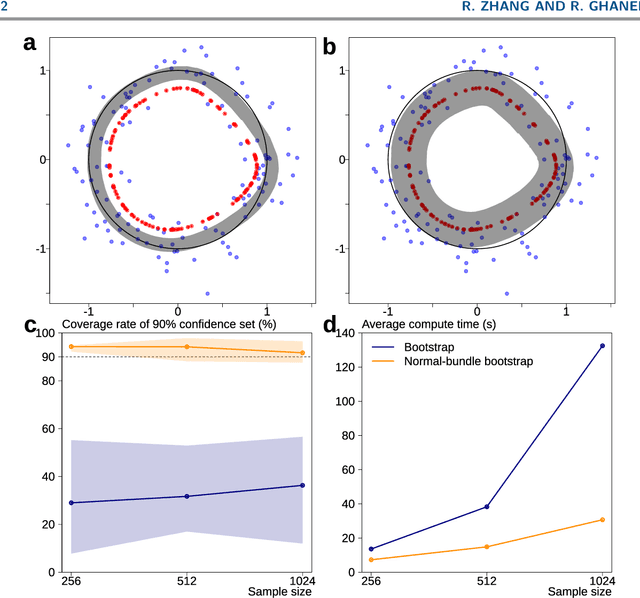
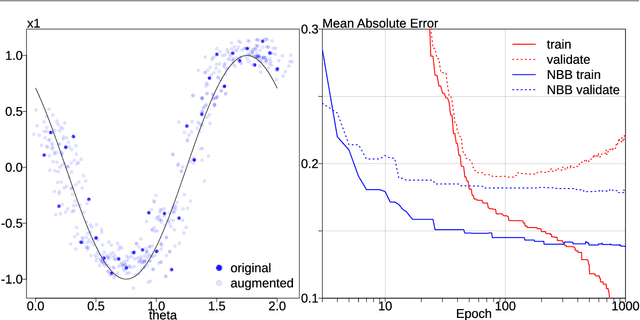
Abstract:Probabilistic models of data sets often exhibit salient geometric structure. Such a phenomenon is summed up in the manifold distribution hypothesis, and can be exploited in probabilistic learning. Here we present normal-bundle bootstrap (NBB), a method that generates new data which preserve the geometric structure of a given data set. Inspired by algorithms for manifold learning and concepts in differential geometry, our method decomposes the underlying probability measure into a marginalized measure on a learned data manifold and conditional measures on the normal spaces. The algorithm estimates the data manifold as a density ridge, and constructs new data by bootstrapping projection vectors and adding them to the ridge. We apply our method to the inference of density ridge and related statistics, and data augmentation to reduce overfitting.
Data-driven discovery of free-form governing differential equations
Nov 11, 2019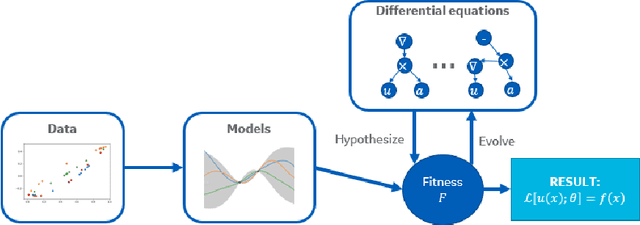
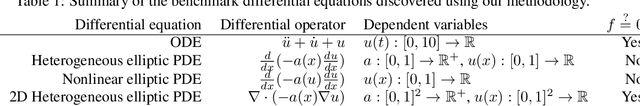

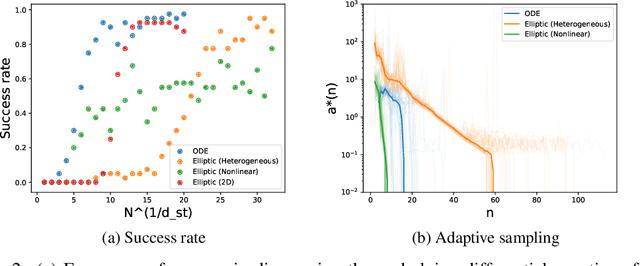
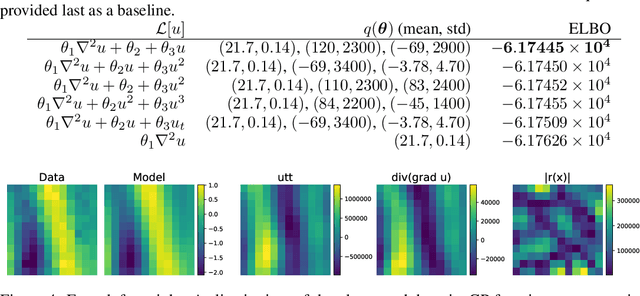
Abstract:We present a method of discovering governing differential equations from data without the need to specify a priori the terms to appear in the equation. The input to our method is a dataset (or ensemble of datasets) corresponding to a particular solution (or ensemble of particular solutions) of a differential equation. The output is a human-readable differential equation with parameters calibrated to the individual particular solutions provided. The key to our method is to learn differentiable models of the data that subsequently serve as inputs to a genetic programming algorithm in which graphs specify computation over arbitrary compositions of functions, parameters, and (potentially differential) operators on functions. Differential operators are composed and evaluated using recursive application of automatic differentiation, allowing our algorithm to explore arbitrary compositions of operators without the need for human intervention. We also demonstrate an active learning process to identify and remedy deficiencies in the proposed governing equations.
Sampling of Bayesian posteriors with a non-Gaussian probabilistic learning on manifolds from a small dataset
Oct 28, 2019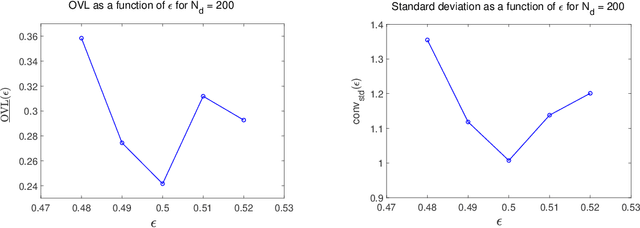
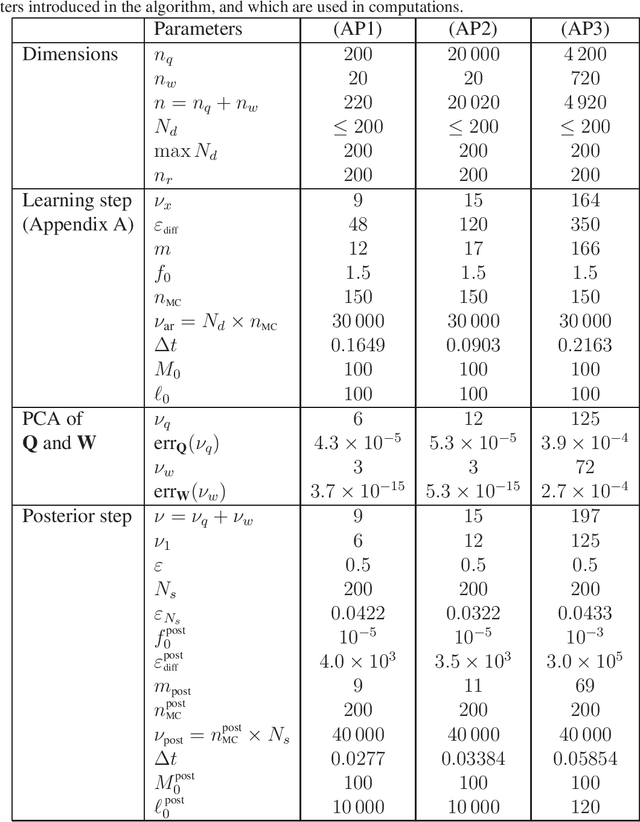
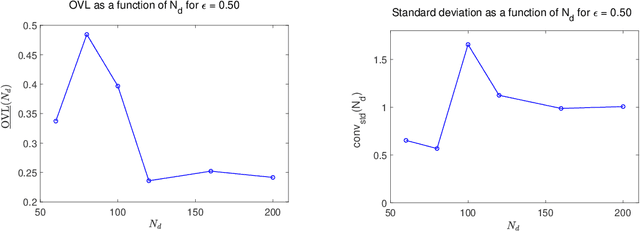
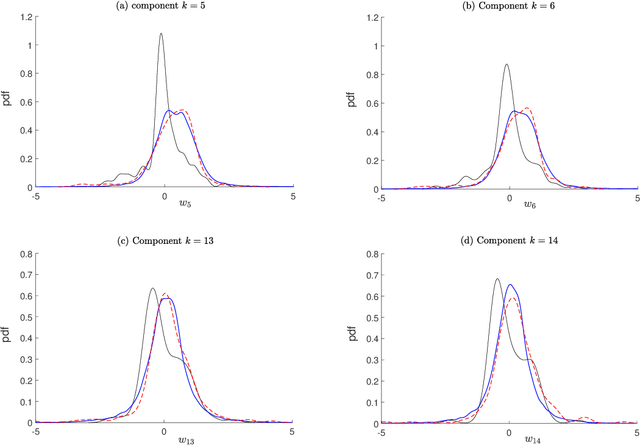
Abstract:This paper tackles the challenge presented by small-data to the task of Bayesian inference. A novel methodology, based on manifold learning and manifold sampling, is proposed for solving this computational statistics problem under the following assumptions: 1) neither the prior model nor the likelihood function are Gaussian and neither can be approximated by a Gaussian measure; 2) the number of functional input (system parameters) and functional output (quantity of interest) can be large; 3) the number of available realizations of the prior model is small, leading to the small-data challenge typically associated with expensive numerical simulations; the number of experimental realizations is also small; 4) the number of the posterior realizations required for decision is much larger than the available initial dataset. The method and its mathematical aspects are detailed. Three applications are presented for validation: The first two involve mathematical constructions aimed to develop intuition around the method and to explore its performance. The third example aims to demonstrate the operational value of the method using a more complex application related to the statistical inverse identification of the non-Gaussian matrix-valued random elasticity field of a damaged biological tissue (osteoporosis in a cortical bone) using ultrasonic waves.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge