Qihui Lyu
TomoGRAF: A Robust and Generalizable Reconstruction Network for Single-View Computed Tomography
Nov 12, 2024



Abstract:Computed tomography (CT) provides high spatial resolution visualization of 3D structures for scientific and clinical applications. Traditional analytical/iterative CT reconstruction algorithms require hundreds of angular data samplings, a condition that may not be met in practice due to physical and mechanical limitations. Sparse view CT reconstruction has been proposed using constrained optimization and machine learning methods with varying success, less so for ultra-sparse view CT reconstruction with one to two views. Neural radiance field (NeRF) is a powerful tool for reconstructing and rendering 3D natural scenes from sparse views, but its direct application to 3D medical image reconstruction has been minimally successful due to the differences between optical and X-ray photon transportation. Here, we develop a novel TomoGRAF framework incorporating the unique X-ray transportation physics to reconstruct high-quality 3D volumes using ultra-sparse projections without prior. TomoGRAF captures the CT imaging geometry, simulates the X-ray casting and tracing process, and penalizes the difference between simulated and ground truth CT sub-volume during training. We evaluated the performance of TomoGRAF on an unseen dataset of distinct imaging characteristics from the training data and demonstrated a vast leap in performance compared with state-of-the-art deep learning and NeRF methods. TomoGRAF provides the first generalizable solution for image-guided radiotherapy and interventional radiology applications, where only one or a few X-ray views are available, but 3D volumetric information is desired.
A Two-Step Framework for Multi-Material Decomposition of Dual Energy Computed Tomography from Projection Domain
Oct 31, 2023Abstract:Dual-energy computed tomography (DECT) utilizes separate X-ray energy spectra to improve multi-material decomposition (MMD) for various diagnostic applications. However accurate decomposing more than two types of material remains challenging using conventional methods. Deep learning (DL) methods have shown promise to improve the MMD performance, but typical approaches of conducing DL-MMD in the image domain fail to fully utilize projection information or under iterative setup are computationally inefficient in both training and prediction. In this work, we present a clinical-applicable MMD (>2) framework rFast-MMDNet, operating with raw projection data in non-recursive setup, for breast tissue differentiation. rFast-MMDNet is a two-stage algorithm, including stage-one SinoNet to perform dual energy projection decomposition on tissue sinograms and stage-two FBP-DenoiseNet to perform domain adaptation and image post-processing. rFast-MMDNet was tested on a 2022 DL-Spectral-Challenge breast phantom dataset. The two stages of rFast-MMDNet were evaluated separately and then compared with four noniterative reference methods including a direct inversion method (AA-MMD), an image domain DL method (ID-UNet), AA-MMD/ID-UNet + DenoiseNet and a sinogram domain DL method (Triple-CBCT). Our results show that models trained from information stored in DE transmission domain can yield high-fidelity decomposition of the adipose, calcification, and fibroglandular materials with averaged RMSE, MAE, negative PSNR, and SSIM of 0.004+/-~0, 0.001+/-~0, -45.027+/-~0.542, and 0.002+/-~0 benchmarking to the ground truth, respectively. Training of entire rFast-MMDNet on a 4xRTX A6000 GPU cluster took a day with inference time <1s. All DL methods generally led to more accurate MMD than AA-MMD. rFast-MMDNet outperformed Triple-CBCT, but both are superior to the image-domain based methods.
To what extent can Plug-and-Play methods outperform neural networks alone in low-dose CT reconstruction
Feb 15, 2022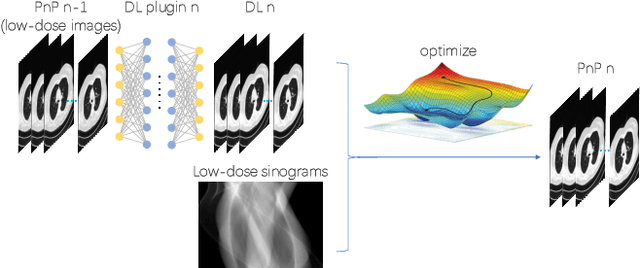
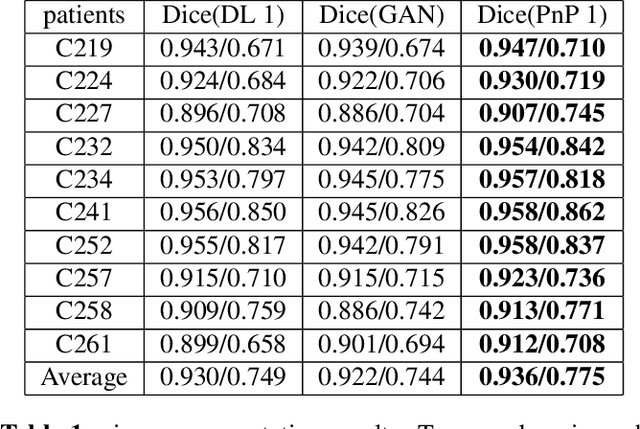
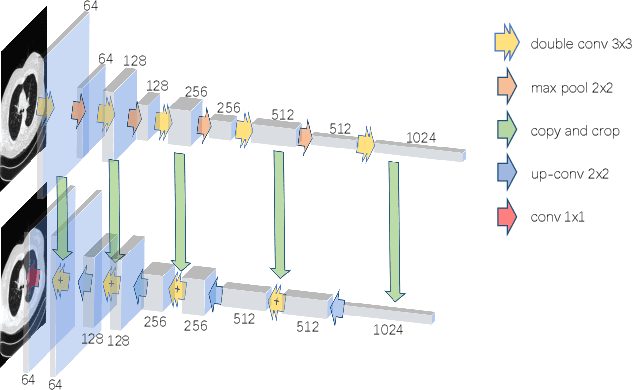
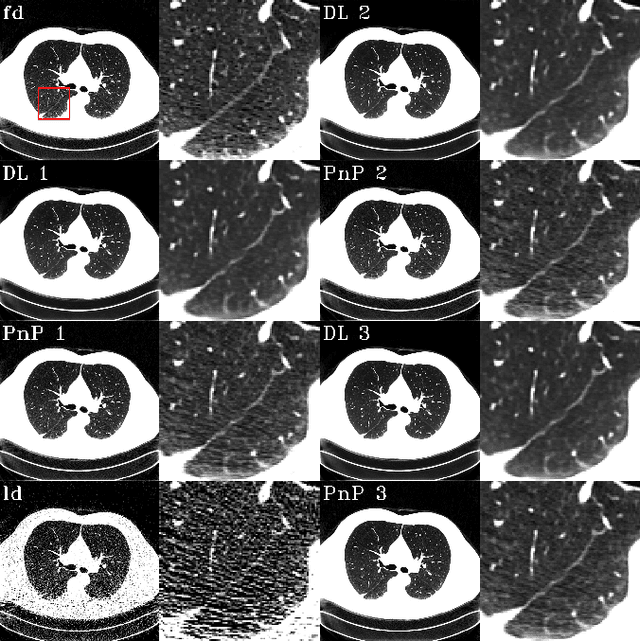
Abstract:The Plug-and-Play (PnP) framework was recently introduced for low-dose CT reconstruction to leverage the interpretability and the flexibility of model-based methods to incorporate various plugins, such as trained deep learning (DL) neural networks. However, the benefits of PnP vs. state-of-the-art DL methods have not been clearly demonstrated. In this work, we proposed an improved PnP framework to address the previous limitations and develop clinical-relevant segmentation metrics for quantitative result assessment. Compared with the DL alone methods, our proposed PnP framework was slightly inferior in MSE and PSNR. However, the power spectrum of the resulting images better matched that of full-dose images than that of DL denoised images. The resulting images supported higher accuracy in airway segmentation than DL denoised images for all the ten patients in the test set, more substantially on the airways with a cross-section smaller than 0.61cm$^2$, and outperformed the DL denoised images for 45 out of 50 lung lobes in lobar segmentation. Our PnP method proved to be significantly better at preserving the image texture, which translated to task-specific benefits in automated structure segmentation and detection.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge