Peter Szolovits
N3C Natural Language Processing
Clinically Accurate Chest X-Ray Report Generation
Apr 04, 2019
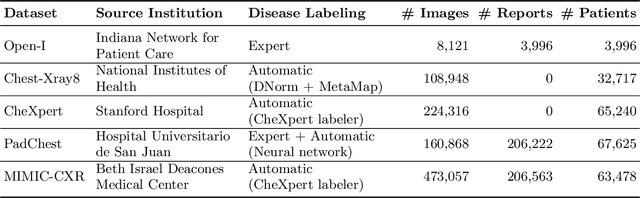
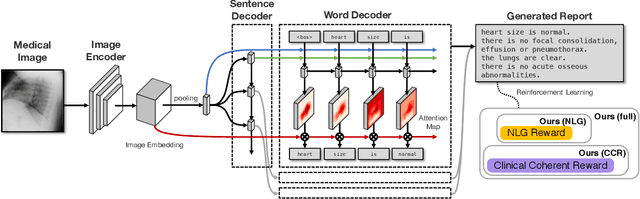
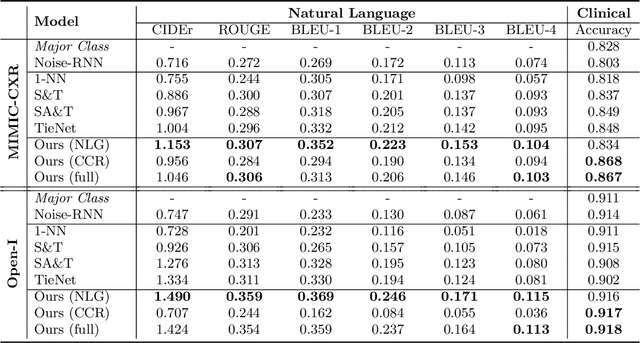
Abstract:The automatic generation of radiology reports given medical radiographs has significant potential to operationally and clinically improve patient care. A number of prior works have focused on this problem, employing advanced methods from computer vision and natural language generation to produce readable reports. However, these works often fail to account for the particular nuances of the radiology domain, and, in particular, the critical importance of clinical accuracy in the resulting generated reports. In this work, we present a domain-aware automatic chest X-Ray radiology report generation system which first predicts what topics will be discussed in the report, then conditionally generates sentences corresponding to these topics. The resulting system is fine-tuned using reinforcement learning, considering both readability and clinical accuracy, as assessed by the proposed Clinically Coherent Reward. We verify this system on two datasets, Open-I and MIMIC-CXR, and demonstrate that our model offers marked improvements on both language generation metrics and CheXpert assessed accuracy over a variety of competitive baselines.
Unsupervised Clinical Language Translation
Feb 04, 2019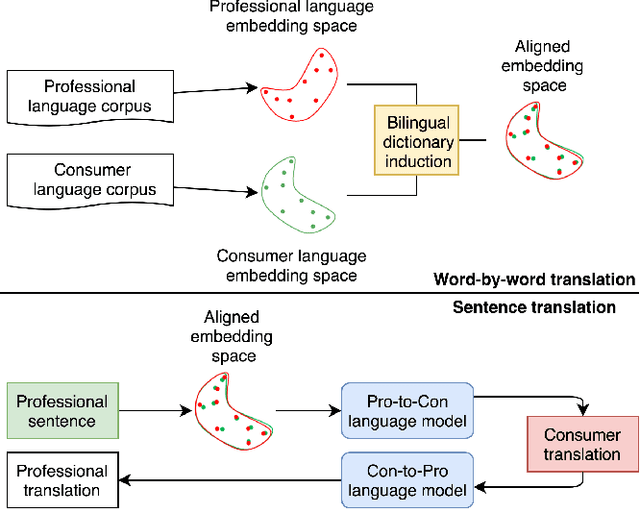
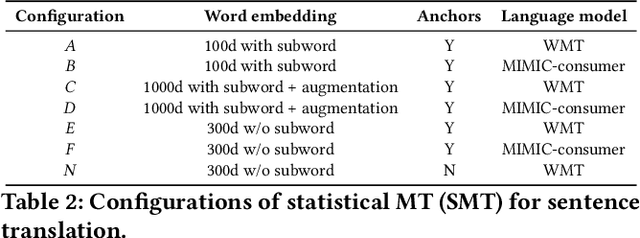
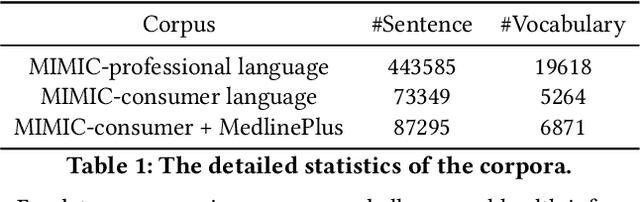

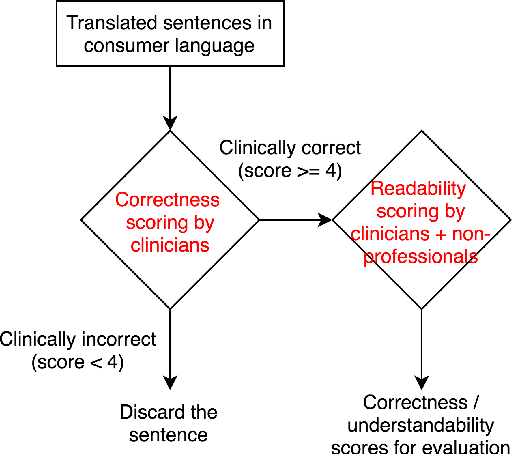
Abstract:As patients' access to their doctors' clinical notes becomes common, translating professional, clinical jargon to layperson-understandable language is essential to improve patient-clinician communication. Such translation yields better clinical outcomes by enhancing patients' understanding of their own health conditions, and thus improving patients' involvement in their own care. Existing research has used dictionary-based word replacement or definition insertion to approach the need. However, these methods are limited by expert curation, which is hard to scale and has trouble generalizing to unseen datasets that do not share an overlapping vocabulary. In contrast, we approach the clinical word and sentence translation problem in a completely unsupervised manner. We show that a framework using representation learning, bilingual dictionary induction and statistical machine translation yields the best precision at 10 of 0.827 on professional-to-consumer word translation, and mean opinion scores of 4.10 and 4.28 out of 5 for clinical correctness and layperson readability, respectively, on sentence translation. Our fully-unsupervised strategy overcomes the curation problem, and the clinically meaningful evaluation reduces biases from inappropriate evaluators, which are critical in clinical machine learning.
Predicting Blood Pressure Response to Fluid Bolus Therapy Using Attention-Based Neural Networks for Clinical Interpretability
Dec 03, 2018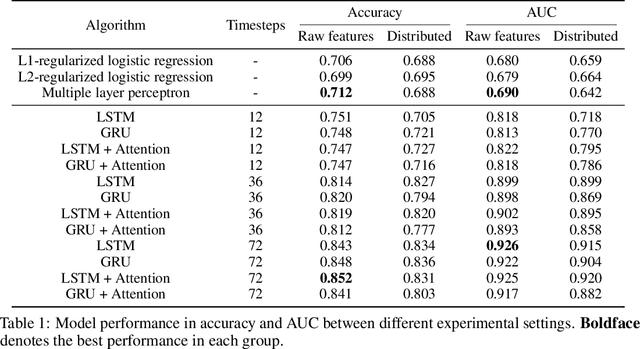
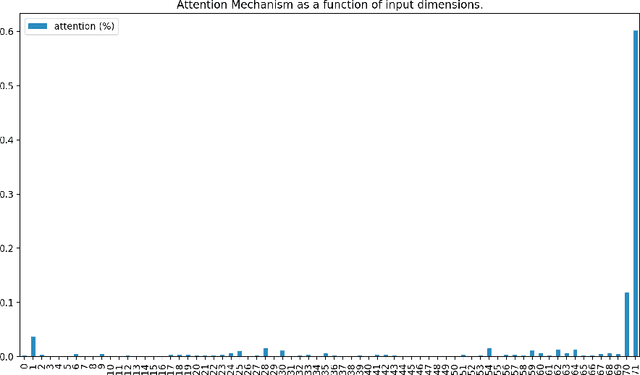
Abstract:Determining whether hypotensive patients in intensive care units (ICUs) should receive fluid bolus therapy (FBT) has been an extremely challenging task for intensive care physicians as the corresponding increase in blood pressure has been hard to predict. Our study utilized regression models and attention-based recurrent neural network (RNN) algorithms and a multi-clinical information system large-scale database to build models that can predict the successful response to FBT among hypotensive patients in ICUs. We investigated both time-aggregated modeling using logistic regression algorithms with regularization and time-series modeling using the long short term memory network (LSTM) and the gated recurrent units network (GRU) with the attention mechanism for clinical interpretability. Among all modeling strategies, the stacked LSTM with the attention mechanism yielded the most predictable model with the highest accuracy of 0.852 and area under the curve (AUC) value of 0.925. The study results may help identify hypotensive patients in ICUs who will have sufficient blood pressure recovery after FBT.
Unsupervised Multimodal Representation Learning across Medical Images and Reports
Nov 21, 2018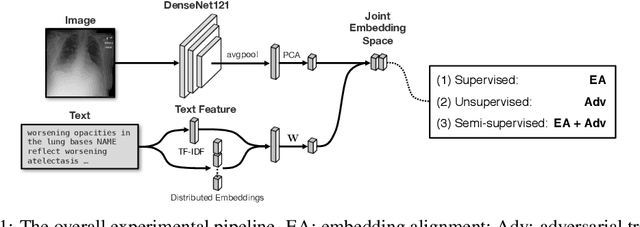
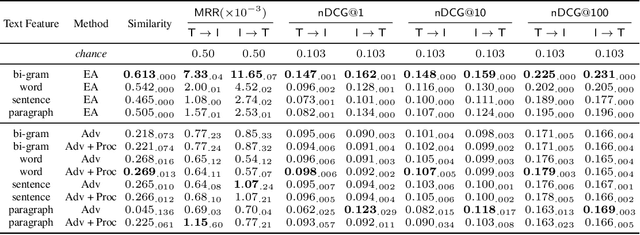
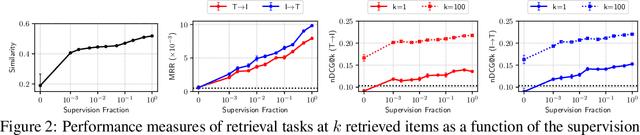
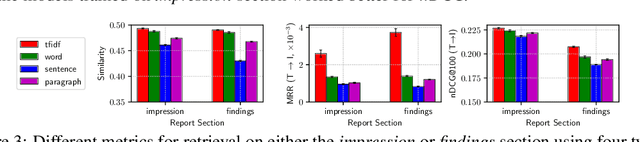
Abstract:Joint embeddings between medical imaging modalities and associated radiology reports have the potential to offer significant benefits to the clinical community, ranging from cross-domain retrieval to conditional generation of reports to the broader goals of multimodal representation learning. In this work, we establish baseline joint embedding results measured via both local and global retrieval methods on the soon to be released MIMIC-CXR dataset consisting of both chest X-ray images and the associated radiology reports. We examine both supervised and unsupervised methods on this task and show that for document retrieval tasks with the learned representations, only a limited amount of supervision is needed to yield results comparable to those of fully-supervised methods.
Implementing a Portable Clinical NLP System with a Common Data Model - a Lisp Perspective
Nov 15, 2018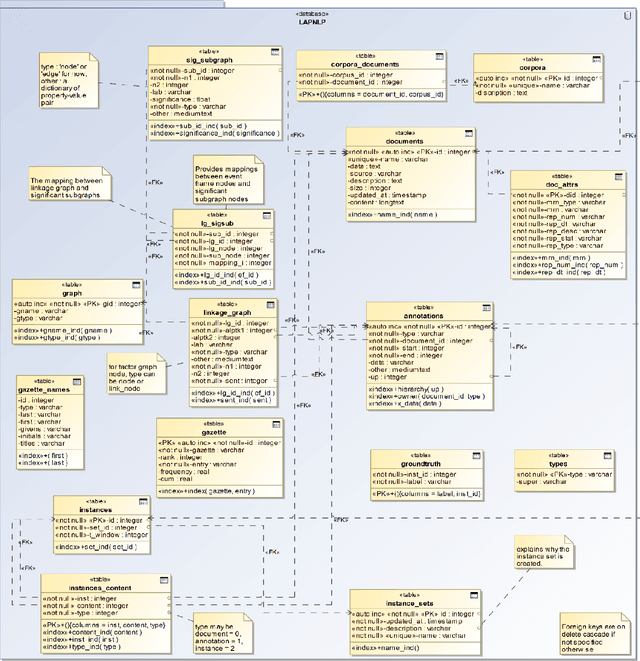
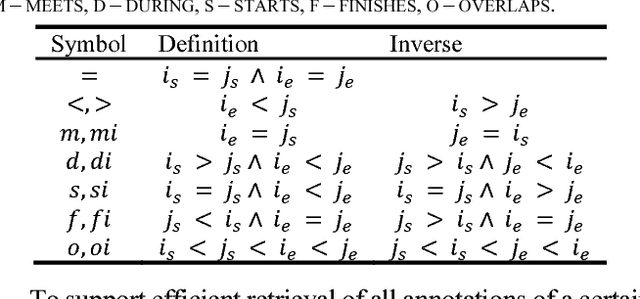

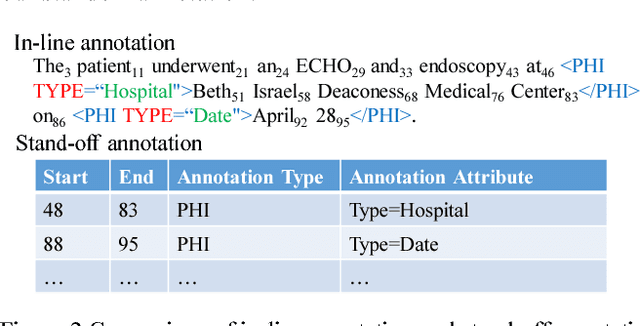
Abstract:This paper presents a Lisp architecture for a portable NLP system, termed LAPNLP, for processing clinical notes. LAPNLP integrates multiple standard, customized and in-house developed NLP tools. Our system facilitates portability across different institutions and data systems by incorporating an enriched Common Data Model (CDM) to standardize necessary data elements. It utilizes UMLS to perform domain adaptation when integrating generic domain NLP tools. It also features stand-off annotations that are specified by positional reference to the original document. We built an interval tree based search engine to efficiently query and retrieve the stand-off annotations by specifying positional requirements. We also developed a utility to convert an inline annotation format to stand-off annotations to enable the reuse of clinical text datasets with inline annotations. We experimented with our system on several NLP facilitated tasks including computational phenotyping for lymphoma patients and semantic relation extraction for clinical notes. These experiments showcased the broader applicability and utility of LAPNLP.
Advancing PICO Element Detection in Medical Text via Deep Neural Networks
Oct 30, 2018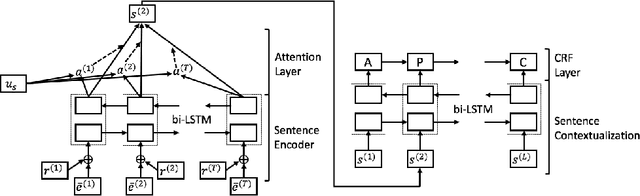
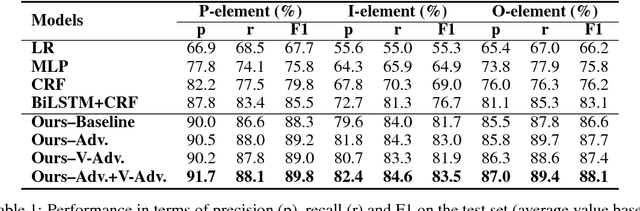
Abstract:In evidence-based medicine (EBM), structured medical questions are always favored for efficient search of the best available evidence for treatments. PICO element detection is widely used to help structurize the clinical studies and question by identifying the sentences in a given medical text that belong to one of the four components: Participants (P), Intervention (I), Comparison (C), and Outcome (O). In this work, we propose a hierarchical deep neural network (DNN) architecture that contains dual bi-directional long short-term memory (bi-LSTM) layers to automatically detect the PICO element in medical texts. Within the model, the lower layer of bi-LSTM is for sentence encoding while the upper one is to contextualize the encoded sentence representation vector. In addition, we adopt adversarial and virtual adversarial training to regularize the model. Overall, we advance the PICO element detection to new state-of-the-art performance, outperforming the previous works by at least 4\% in F1 score for all P/I/O categories.
Hierarchical Neural Networks for Sequential Sentence Classification in Medical Scientific Abstracts
Aug 19, 2018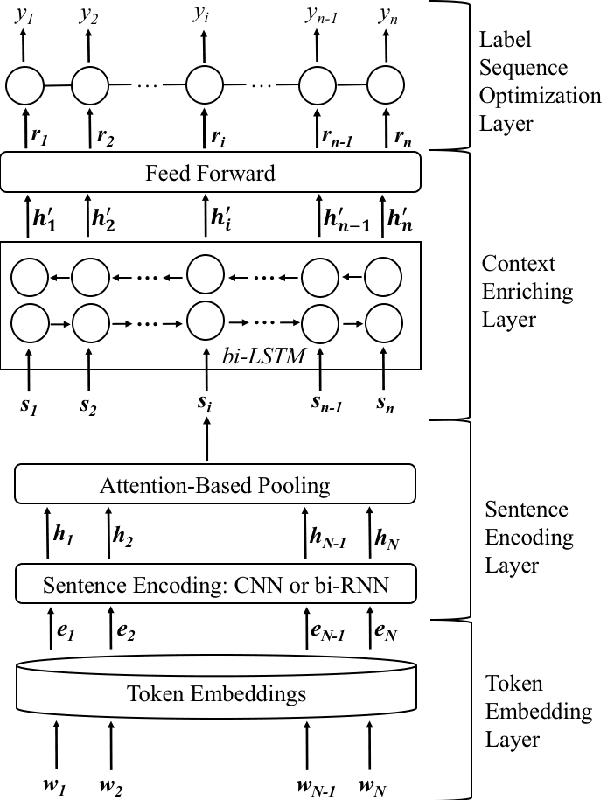
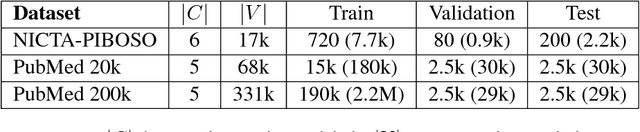
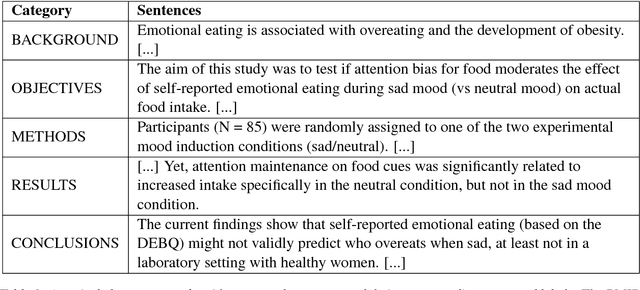
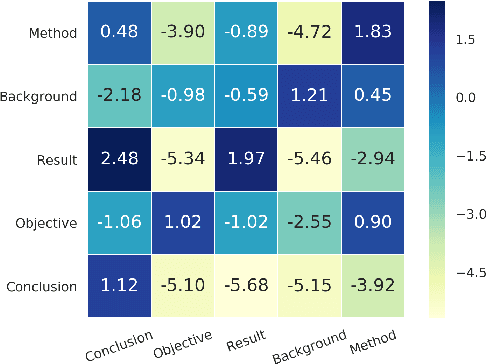
Abstract:Prevalent models based on artificial neural network (ANN) for sentence classification often classify sentences in isolation without considering the context in which sentences appear. This hampers the traditional sentence classification approaches to the problem of sequential sentence classification, where structured prediction is needed for better overall classification performance. In this work, we present a hierarchical sequential labeling network to make use of the contextual information within surrounding sentences to help classify the current sentence. Our model outperforms the state-of-the-art results by 2%-3% on two benchmarking datasets for sequential sentence classification in medical scientific abstracts.
Modeling Mistrust in End-of-Life Care
Jun 30, 2018
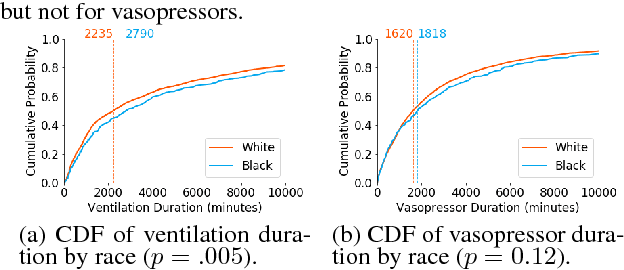
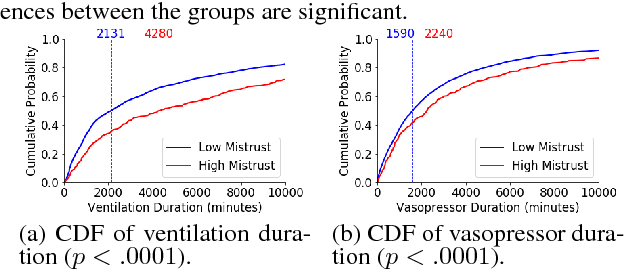
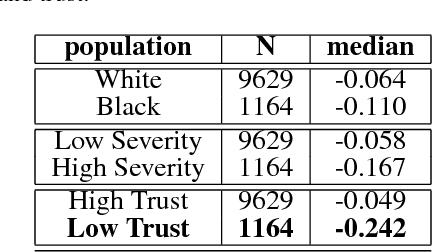
Abstract:In this work, we characterize the doctor-patient relationship using a machine learning-derived trust score. We show that this score has statistically significant racial associations, and that by modeling trust directly we find stronger disparities in care than by stratifying on race. We further demonstrate that mistrust is indicative of worse outcomes, but is only weakly associated with physiologically-created severity scores. Finally, we describe sentiment analysis experiments indicating patients with higher levels of mistrust have worse experiences and interactions with their caregivers. This work is a step towards measuring fairer machine learning in the healthcare domain.
Mapping Unparalleled Clinical Professional and Consumer Languages with Embedding Alignment
Jun 25, 2018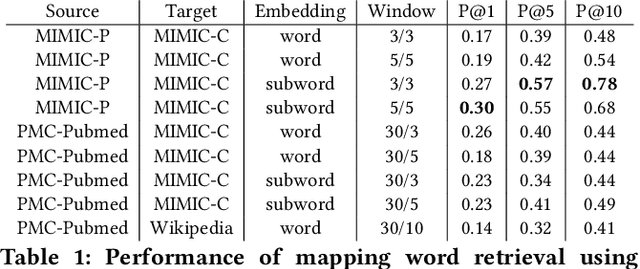
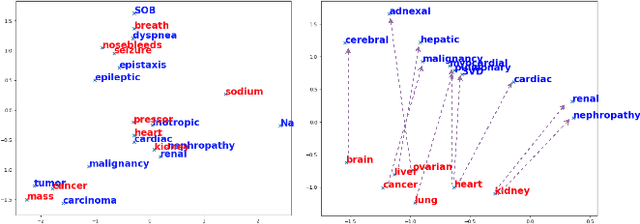
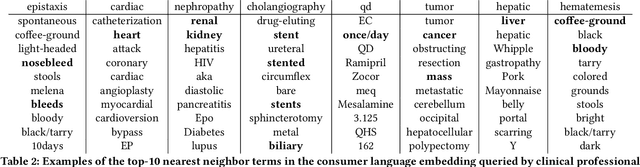
Abstract:Mapping and translating professional but arcane clinical jargons to consumer language is essential to improve the patient-clinician communication. Researchers have used the existing biomedical ontologies and consumer health vocabulary dictionary to translate between the languages. However, such approaches are limited by expert efforts to manually build the dictionary, which is hard to be generalized and scalable. In this work, we utilized the embeddings alignment method for the word mapping between unparalleled clinical professional and consumer language embeddings. To map semantically similar words in two different word embeddings, we first independently trained word embeddings on both the corpus with abundant clinical professional terms and the other with mainly healthcare consumer terms. Then, we aligned the embeddings by the Procrustes algorithm. We also investigated the approach with the adversarial training with refinement. We evaluated the quality of the alignment through the similar words retrieval both by computing the model precision and as well as judging qualitatively by human. We show that the Procrustes algorithm can be performant for the professional consumer language embeddings alignment, whereas adversarial training with refinement may find some relations between two languages.
Towards the Creation of a Large Corpus of Synthetically-Identified Clinical Notes
Mar 07, 2018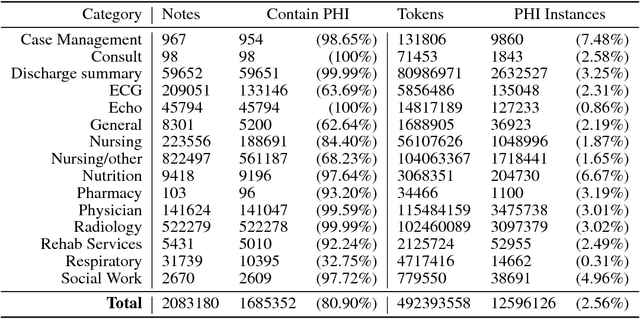
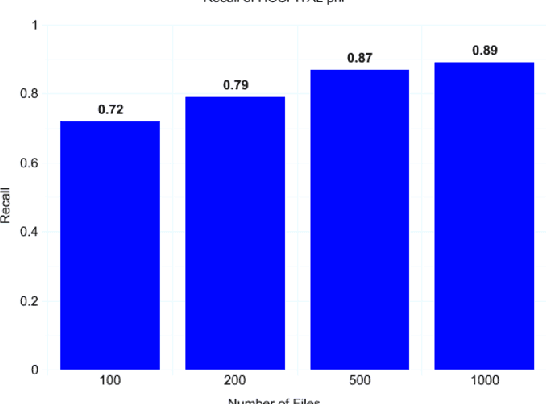
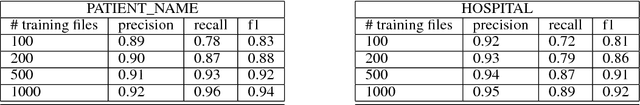
Abstract:Clinical notes often describe the most important aspects of a patient's physiology and are therefore critical to medical research. However, these notes are typically inaccessible to researchers without prior removal of sensitive protected health information (PHI), a natural language processing (NLP) task referred to as deidentification. Tools to automatically de-identify clinical notes are needed but are difficult to create without access to those very same notes containing PHI. This work presents a first step toward creating a large synthetically-identified corpus of clinical notes and corresponding PHI annotations in order to facilitate the development de-identification tools. Further, one such tool is evaluated against this corpus in order to understand the advantages and shortcomings of this approach.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge