Patrick Ryan
Ontologizing Health Systems Data at Scale: Making Translational Discovery a Reality
Sep 10, 2022
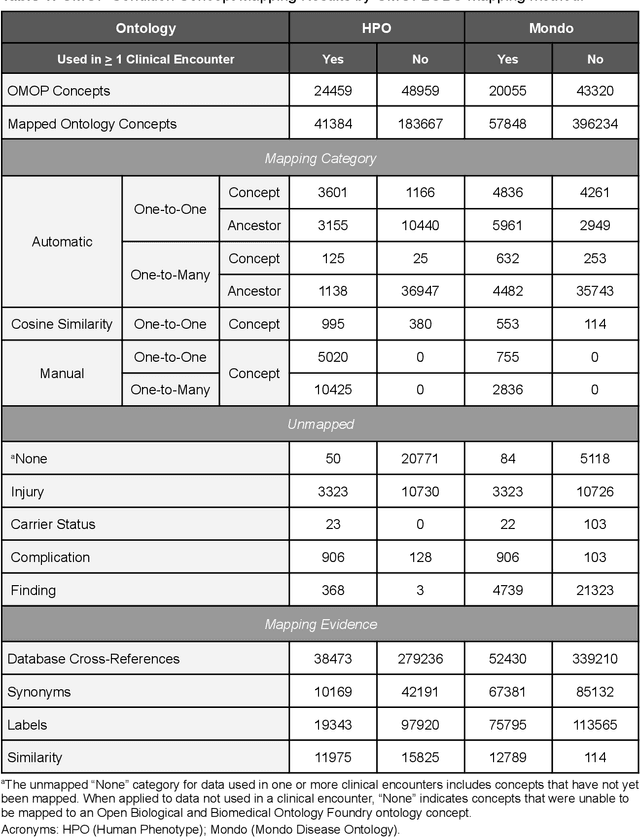
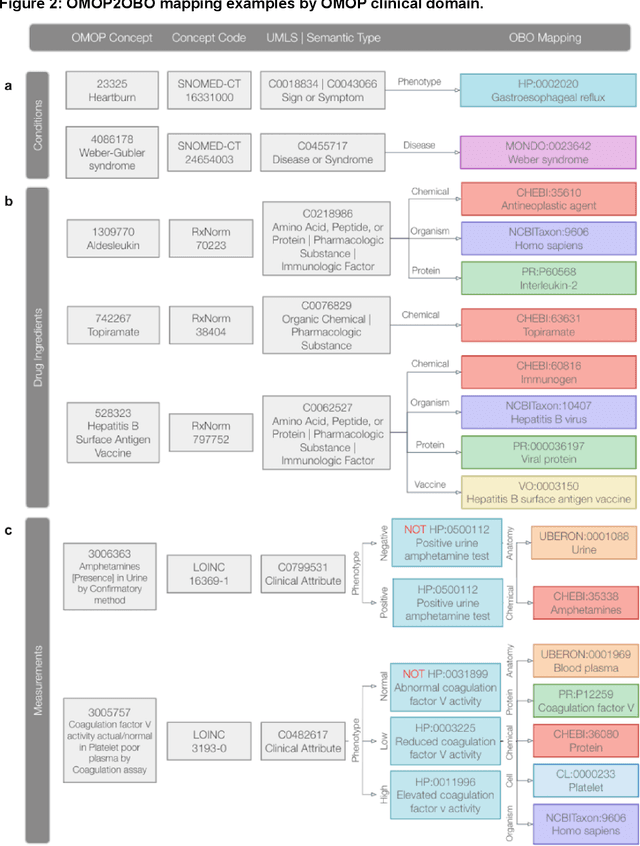
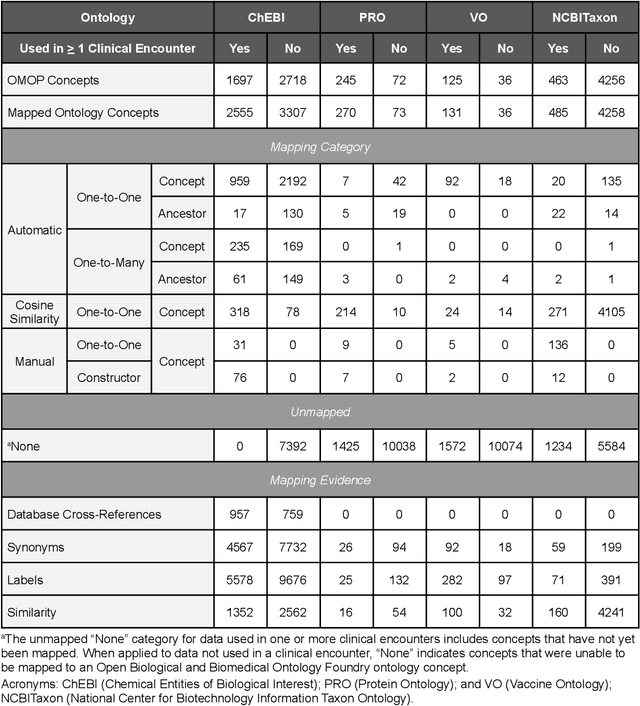
Abstract:Common data models solve many challenges of standardizing electronic health record (EHR) data, but are unable to semantically integrate the resources needed for deep phenotyping. Open Biological and Biomedical Ontology (OBO) Foundry ontologies provide semantically computable representations of biological knowledge and enable the integration of a variety of biomedical data. However, mapping EHR data to OBO Foundry ontologies requires significant manual curation and domain expertise. We introduce a framework for mapping Observational Medical Outcomes Partnership (OMOP) standard vocabularies to OBO Foundry ontologies. Using this framework, we produced mappings for 92,367 conditions, 8,615 drug ingredients, and 10,673 measurement results. Mapping accuracy was verified by domain experts and when examined across 24 hospitals, the mappings covered 99% of conditions and drug ingredients and 68% of measurements. Finally, we demonstrate that OMOP2OBO mappings can aid in the systematic identification of undiagnosed rare disease patients who might benefit from genetic testing.
A Comparison of Resampling and Recursive Partitioning Methods in Random Forest for Estimating the Asymptotic Variance Using the Infinitesimal Jackknife
Jan 29, 2018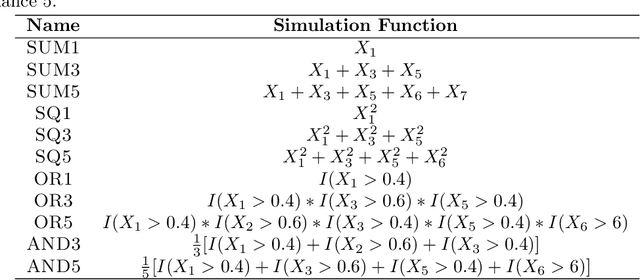
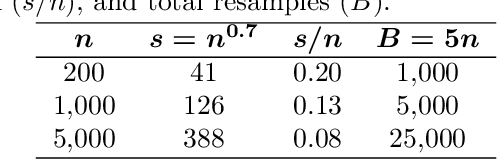
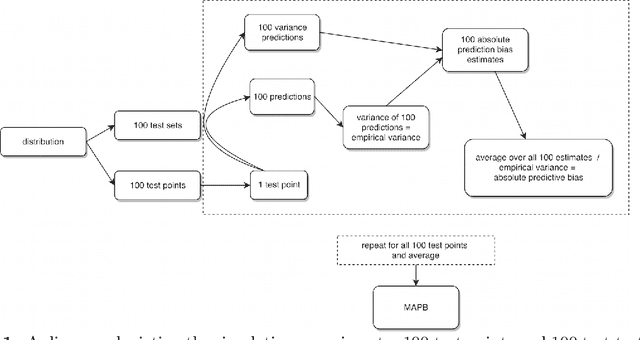

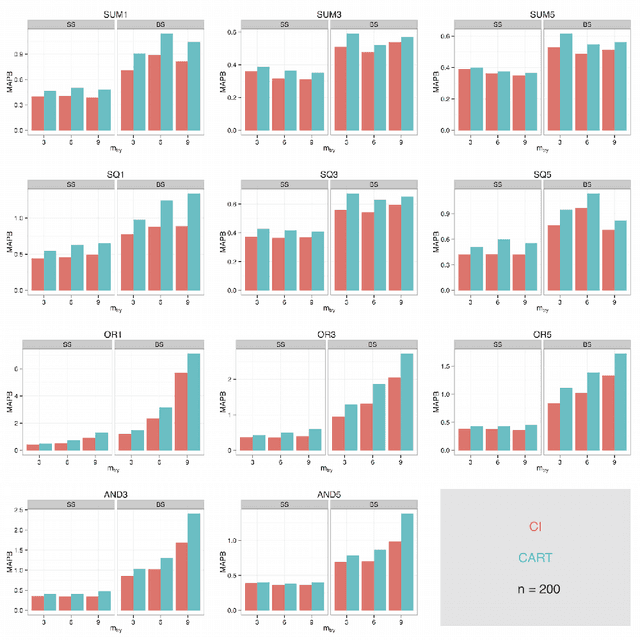
Abstract:The infinitesimal jackknife (IJ) has recently been applied to the random forest to estimate its prediction variance. These theorems were verified under a traditional random forest framework which uses classification and regression trees (CART) and bootstrap resampling. However, random forests using conditional inference (CI) trees and subsampling have been found to be not prone to variable selection bias. Here, we conduct simulation experiments using a novel approach to explore the applicability of the IJ to random forests using variations on the resampling method and base learner. Test data points were simulated and each trained using random forest on one hundred simulated training data sets using different combinations of resampling and base learners. Using CI trees instead of traditional CART trees as well as using subsampling instead of bootstrap sampling resulted in a much more accurate estimation of prediction variance when using the IJ. The random forest variations here have been incorporated into an open source software package for the R programming language.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge