Ofer Meshi
Train and Test Tightness of LP Relaxations in Structured Prediction
Apr 27, 2016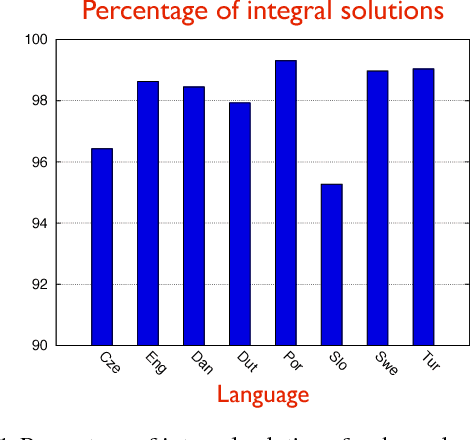
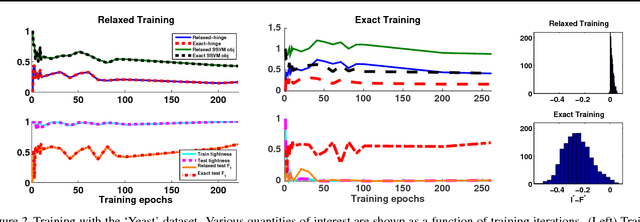
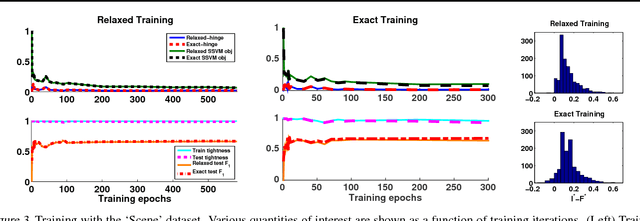
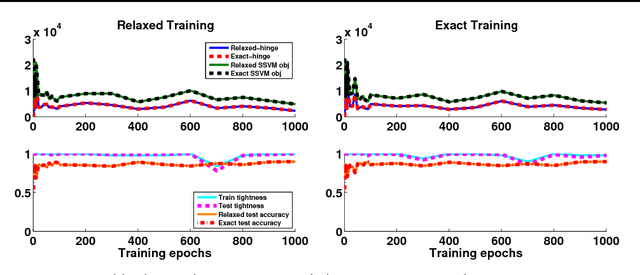
Abstract:Structured prediction is used in areas such as computer vision and natural language processing to predict structured outputs such as segmentations or parse trees. In these settings, prediction is performed by MAP inference or, equivalently, by solving an integer linear program. Because of the complex scoring functions required to obtain accurate predictions, both learning and inference typically require the use of approximate solvers. We propose a theoretical explanation to the striking observation that approximations based on linear programming (LP) relaxations are often tight on real-world instances. In particular, we show that learning with LP relaxed inference encourages integrality of training instances, and that tightness generalizes from train to test data.
Fast and Scalable Structural SVM with Slack Rescaling
Oct 27, 2015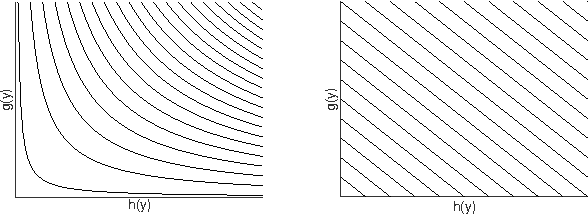
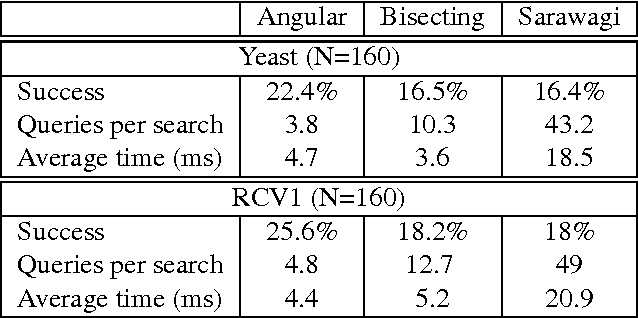
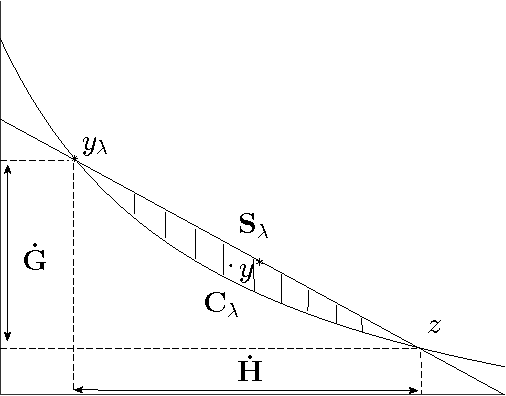
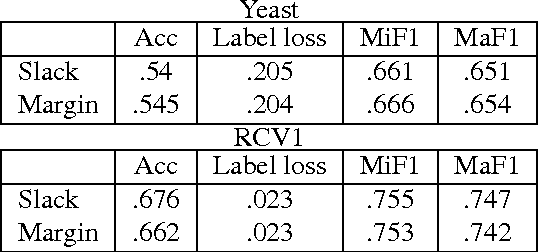
Abstract:We present an efficient method for training slack-rescaled structural SVM. Although finding the most violating label in a margin-rescaled formulation is often easy since the target function decomposes with respect to the structure, this is not the case for a slack-rescaled formulation, and finding the most violated label might be very difficult. Our core contribution is an efficient method for finding the most-violating-label in a slack-rescaled formulation, given an oracle that returns the most-violating-label in a (slightly modified) margin-rescaled formulation. We show that our method enables accurate and scalable training for slack-rescaled SVMs, reducing runtime by an order of magnitude compared to previous approaches to slack-rescaled SVMs.
Learning Max-Margin Tree Predictors
Sep 26, 2013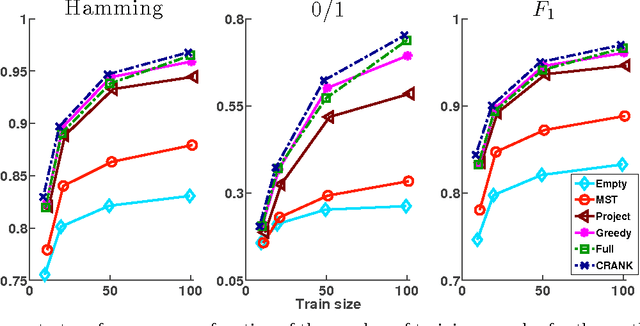
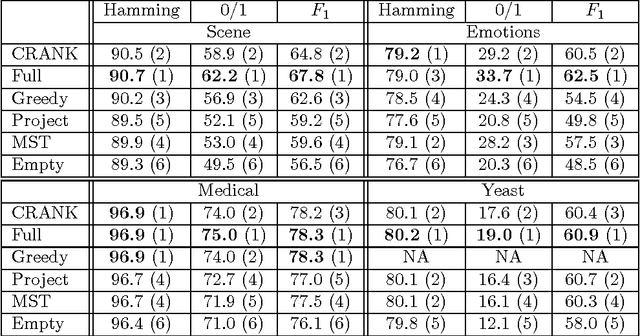
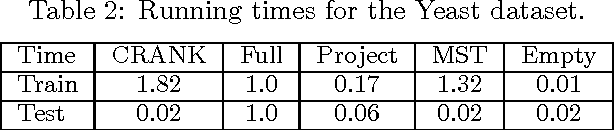
Abstract:Structured prediction is a powerful framework for coping with joint prediction of interacting outputs. A central difficulty in using this framework is that often the correct label dependence structure is unknown. At the same time, we would like to avoid an overly complex structure that will lead to intractable prediction. In this work we address the challenge of learning tree structured predictive models that achieve high accuracy while at the same time facilitate efficient (linear time) inference. We start by proving that this task is in general NP-hard, and then suggest an approximate alternative. Briefly, our CRANK approach relies on a novel Circuit-RANK regularizer that penalizes non-tree structures and that can be optimized using a CCCP procedure. We demonstrate the effectiveness of our approach on several domains and show that, despite the relative simplicity of the structure, prediction accuracy is competitive with a fully connected model that is computationally costly at prediction time.
Template Based Inference in Symmetric Relational Markov Random Fields
Jun 20, 2012
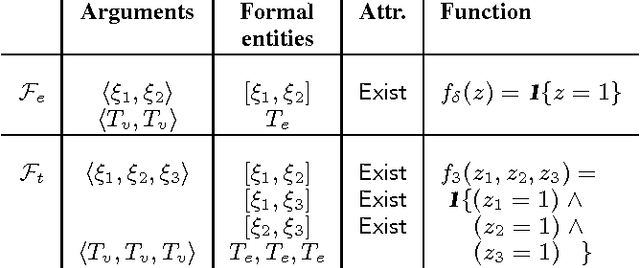

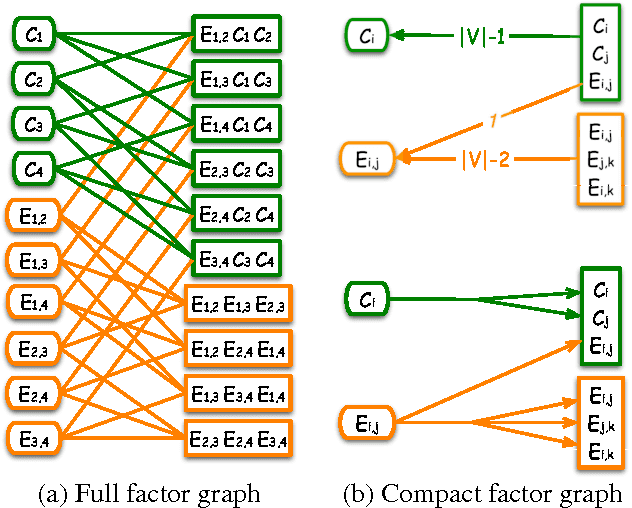
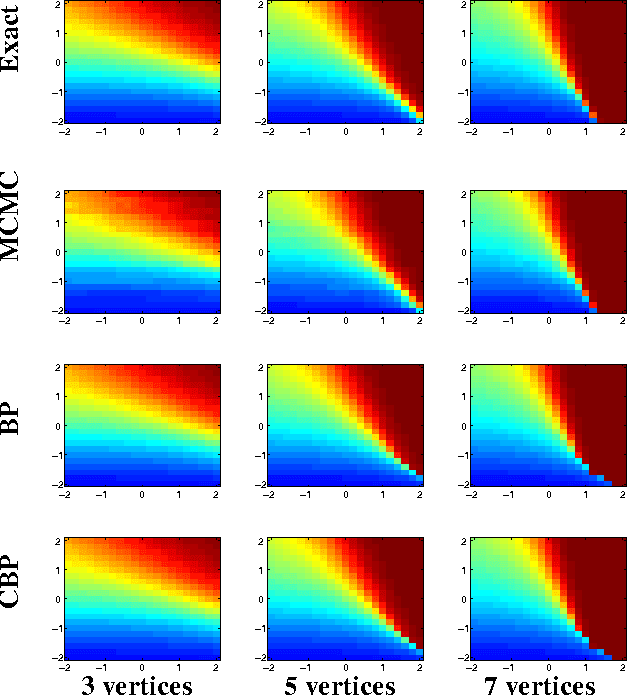
Abstract:Relational Markov Random Fields are a general and flexible framework for reasoning about the joint distribution over attributes of a large number of interacting entities. The main computational difficulty in learning such models is inference. Even when dealing with complete data, where one can summarize a large domain by sufficient statistics, learning requires one to compute the expectation of the sufficient statistics given different parameter choices. The typical solution to this problem is to resort to approximate inference procedures, such as loopy belief propagation. Although these procedures are quite efficient, they still require computation that is on the order of the number of interactions (or features) in the model. When learning a large relational model over a complex domain, even such approximations require unrealistic running time. In this paper we show that for a particular class of relational MRFs, which have inherent symmetry, we can perform the inference needed for learning procedures using a template-level belief propagation. This procedure's running time is proportional to the size of the relational model rather than the size of the domain. Moreover, we show that this computational procedure is equivalent to sychronous loopy belief propagation. This enables a dramatic speedup in inference and learning time. We use this procedure to learn relational MRFs for capturing the joint distribution of large protein-protein interaction networks.
Convexifying the Bethe Free Energy
May 09, 2012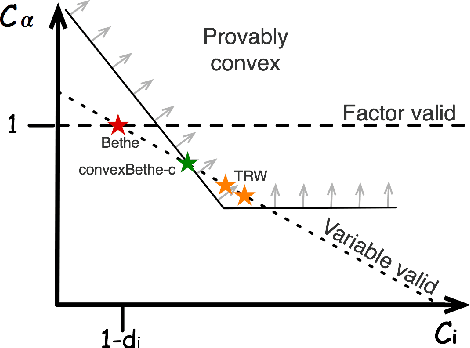
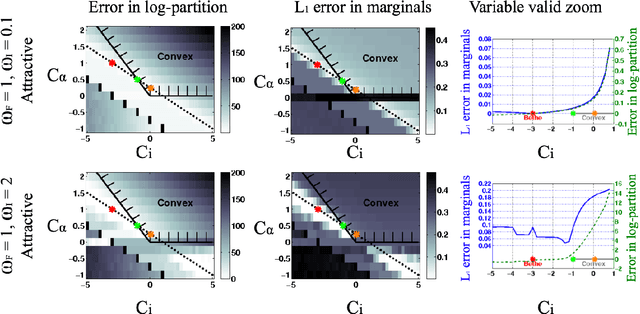
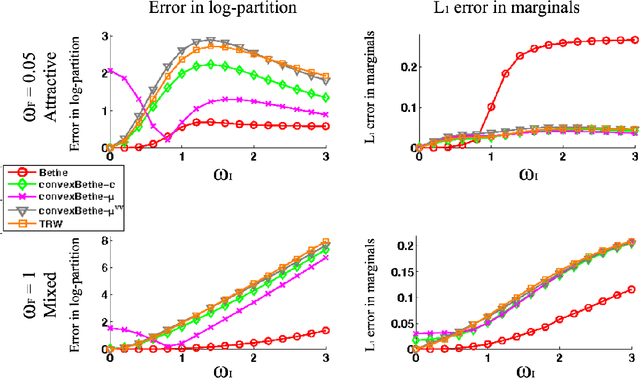
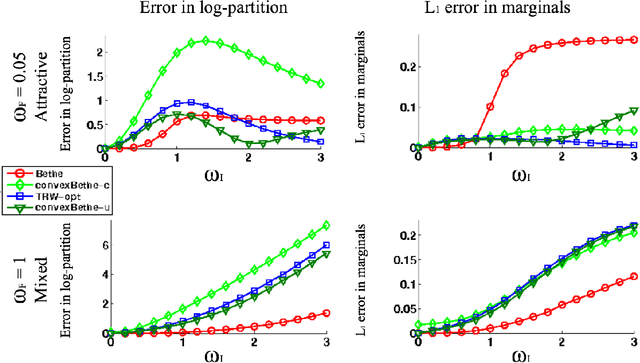
Abstract:The introduction of loopy belief propagation (LBP) revitalized the application of graphical models in many domains. Many recent works present improvements on the basic LBP algorithm in an attempt to overcome convergence and local optima problems. Notable among these are convexified free energy approximations that lead to inference procedures with provable convergence and quality properties. However, empirically LBP still outperforms most of its convex variants in a variety of settings, as we also demonstrate here. Motivated by this fact we seek convexified free energies that directly approximate the Bethe free energy. We show that the proposed approximations compare favorably with state-of-the art convex free energy approximations.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge