Nathan Haut
The Effects of Population Size on the Performance of BEAGLE GPU-Based Genetic Programming Runs
Apr 27, 2026Abstract:The Beagle framework, through GPU-based Genetic Programming, enables population dynamics previously unattainable (within practical time frames) by CPU-constrained Genetic Programming systems. This work explores how GPU-enabled population sizes impact the success of training for symbolic regression problems. Specifically, when using constant population sizes, we see benefits of using very narrow and deep searches (as narrow as 1000 individuals) for some problems, while other problems benefit from very broad and shallow searches (as broad as 10 million individuals). We also explore stepped population sizes that start with large populations and drop to small populations to balance the breadth and depth of search.
EvoIQA - Explaining Image Distortions with Evolved White-Box Logic
Mar 16, 2026Abstract:Traditional Image Quality Assessment (IQA) metrics typically fall into one of two extremes: rigid, hand-crafted mathematical models or "black-box" deep learning architectures that completely lack interpretability. To bridge this gap, we propose EvoIQA, a fully explainable symbolic regression framework based on Genetic Programming that Evolves explicit, human-readable mathematical formulas for image quality assessment (IQA). Utilizing a rich terminal set from the VSI, VIF, FSIM, and HaarPSI metrics, our framework inherently maps structural, chromatic, and information-theoretic degradations into observable mathematical equations. Our results demonstrate that the evolved GP models consistently achieve strong alignment between the predictions and human visual preferences. Furthermore, they not only outperform traditional hand-crafted metrics but also achieve performance parity with complex, state-of-the-art deep learning models like DB-CNN, proving that we no longer have to sacrifice interpretability for state-of-the-art performance.
GPU-Accelerated Genetic Programming for Symbolic Regression with Beagle Framework
Mar 10, 2026Abstract:Beagle is a new software framework that enables execution of Genetic Programming tasks on the GPU. Currently available for symbolic regression, it processes individuals of the population and fitness cases for training in a way that maximizes throughput on extant GPU platforms. In this contribution, we report on the benchmarking of Beagle on the Feynman Symbolic Regression dataset and compare its performance with a fast CPU system called StackGP and the widely available PySR system under the same wall clock budget. We also report on the use of two different fitness functions, one a point-to-point error function, the other a correlation fitness function. The results demonstrate that the Beagle's GPU-aided Symbolic Regression significantly outperforms leading CPU-based frameworks.
Data-Informed Model Complexity Metric for Optimizing Symbolic Regression Models
Jan 29, 2025



Abstract:Choosing models from a well-fitted evolved population that generalizes beyond training data is difficult. We introduce a pragmatic method to estimate model complexity using Hessian rank for post-processing selection. Complexity is approximated by averaging the model output Hessian rank across a few points (N=3), offering efficient and accurate rank estimates. This method aligns model selection with input data complexity, calculated using intrinsic dimensionality (ID) estimators. Using the StackGP system, we develop symbolic regression models for the Penn Machine Learning Benchmark and employ twelve scikit-dimension library methods to estimate ID, aligning model expressiveness with dataset ID. Our data-informed complexity metric finds the ideal complexity window, balancing model expressiveness and accuracy, enhancing generalizability without bias common in methods reliant on user-defined parameters, such as parsimony pressure in weight selection.
Sharpness-Aware Minimization in Genetic Programming
May 17, 2024



Abstract:Sharpness-Aware Minimization (SAM) was recently introduced as a regularization procedure for training deep neural networks. It simultaneously minimizes the fitness (or loss) function and the so-called fitness sharpness. The latter serves as a measure of the nonlinear behavior of a solution and does so by finding solutions that lie in neighborhoods having uniformly similar loss values across all fitness cases. In this contribution, we adapt SAM for tree Genetic Programming (TGP) by exploring the semantic neighborhoods of solutions using two simple approaches. By capitalizing upon perturbing input and output of program trees, sharpness can be estimated and used as a second optimization criterion during the evolution. To better understand the impact of this variant of SAM on TGP, we collect numerous indicators of the evolutionary process, including generalization ability, complexity, diversity, and a recently proposed genotype-phenotype mapping to study the amount of redundancy in trees. The experimental results demonstrate that using any of the two proposed SAM adaptations in TGP allows (i) a significant reduction of tree sizes in the population and (ii) a decrease in redundancy of the trees. When assessed on real-world benchmarks, the generalization ability of the elite solutions does not deteriorate.
Active Learning in Genetic Programming: Guiding Efficient Data Collection for Symbolic Regression
Jul 31, 2023



Abstract:This paper examines various methods of computing uncertainty and diversity for active learning in genetic programming. We found that the model population in genetic programming can be exploited to select informative training data points by using a model ensemble combined with an uncertainty metric. We explored several uncertainty metrics and found that differential entropy performed the best. We also compared two data diversity metrics and found that correlation as a diversity metric performs better than minimum Euclidean distance, although there are some drawbacks that prevent correlation from being used on all problems. Finally, we combined uncertainty and diversity using a Pareto optimization approach to allow both to be considered in a balanced way to guide the selection of informative and unique data points for training.
Correlation versus RMSE Loss Functions in Symbolic Regression Tasks
May 31, 2022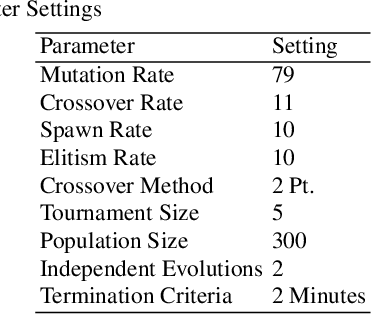
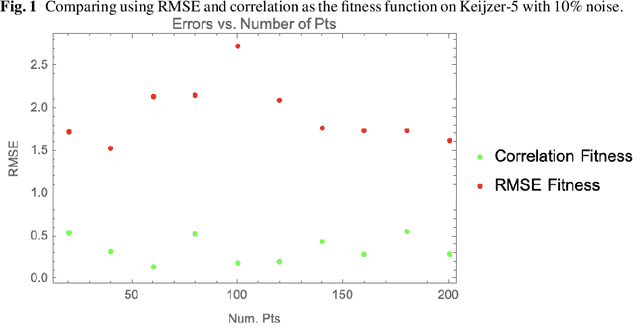
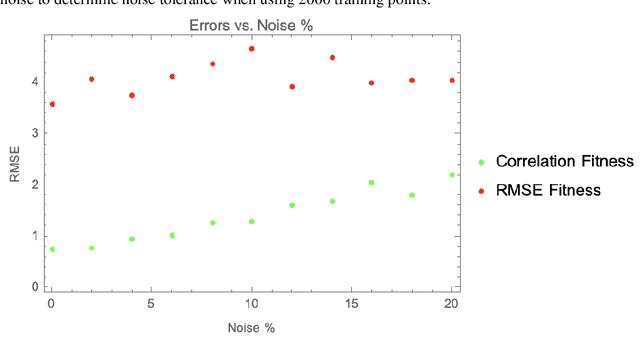
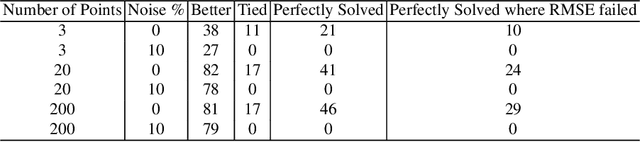
Abstract:The use of correlation as a fitness function is explored in symbolic regression tasks and the performance is compared against the typical RMSE fitness function. Using correlation with an alignment step to conclude the evolution led to significant performance gains over RMSE as a fitness function. Using correlation as a fitness function led to solutions being found in fewer generations compared to RMSE, as well it was found that fewer data points were needed in the training set to discover the correct equations. The Feynman Symbolic Regression Benchmark as well as several other old and recent GP benchmark problems were used to evaluate performance.
Active Learning Improves Performance on Symbolic RegressionTasks in StackGP
Feb 09, 2022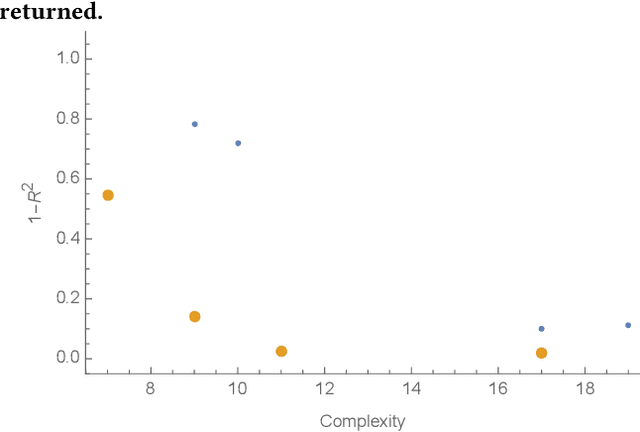
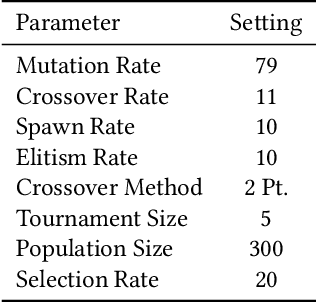

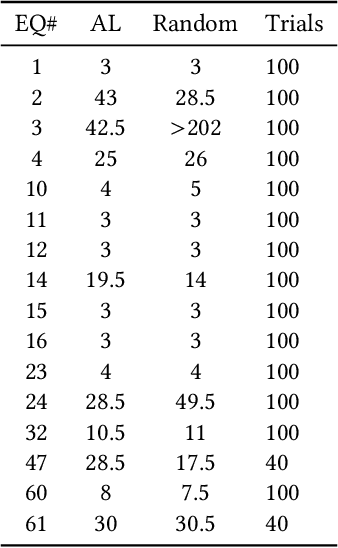
Abstract:In this paper we introduce an active learning method for symbolic regression using StackGP. The approach begins with a small number of data points for StackGP to model. To improve the model the system incrementally adds a data point such that the new point maximizes prediction uncertainty as measured by the model ensemble. Symbolic regression is re-run with the larger data set. This cycle continues until the system satisfies a termination criterion. We use the Feynman AI benchmark set of equations to examine the ability of our method to find appropriate models using fewer data points. The approach was found to successfully rediscover 72 of the 100 Feynman equations using as few data points as possible, and without use of domain expertise or data translation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge